Abstract
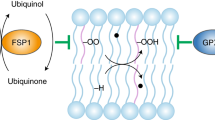
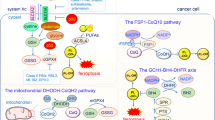
Ferroptosis, an iron-dependent form of cell death driven by lipid peroxidation, provides a potential treatment avenue for drug-resistant cancers and may play a role in the pathology of some degenerative diseases. Identifying the subcellular membranes essential for ferroptosis and the sequence of their peroxidation will illuminate drug discovery strategies and ferroptosis-relevant disease mechanisms. In this study, we employed fluorescence and stimulated Raman scattering imaging to examine the structure–activity–distribution relationship of ferroptosis-modulating compounds. We found that, although lipid peroxidation in various subcellular membranes can induce ferroptosis, the endoplasmic reticulum (ER) membrane is a key site of lipid peroxidation. Our results suggest an ordered progression model of membrane peroxidation during ferroptosis that accumulates initially in the ER membrane and later in the plasma membrane. Thus, the design of ER-targeted inhibitors and inducers of ferroptosis may be used to optimally control the dynamics of lipid peroxidation in cells undergoing ferroptosis.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Lipidomics data are available at Academic Commons at https://doi.org/10.7916/rphv-v394. All other data, including statistical data and blots, are available as Source Data and at https://doi.org/10.7916/hggm-7r90. Source data are provided with this paper.
References
Stockwell, B. R. Ferroptosis turns 10: emerging mechanisms, physiological functions, and therapeutic applications. Cell 185, 2401–2421 (2022).
Stockwell, B. R., Jiang, X. & Gu, W. Emerging mechanisms and disease relevance of ferroptosis. Trends Cell Biol. 30, 478–490 (2020).
Su, Y. et al. Ferroptosis, a novel pharmacological mechanism of anti-cancer drugs. Cancer Lett. 483, 127–136 (2020).
Reichert, C. O. et al. Ferroptosis mechanisms involved in neurodegenerative diseases. Int. J. Mol. Sci. 21, 8765 (2020).
Ye, L. F. et al. Radiation-Induced lipid peroxidation triggers ferroptosis and synergizes with ferroptosis inducers. ACS Chem. Biol. 15, 469–484 (2020).
Zhang, Y. et al. Imidazole ketone erastin induces ferroptosis and slows tumor growth in a mouse lymphoma model. Cell Chem. Biol. 26, 623–633 (2019).
Badgley, M. A. et al. Cysteine depletion induces pancreatic tumor ferroptosis in mice. Science 368, 85–89 (2020).
Louandre, C. et al. Iron-dependent cell death of hepatocellular carcinoma cells exposed to sorafenib. Int. J. Cancer 133, 1732–1742 (2013).
Lachaier, E. et al. Sorafenib induces ferroptosis in human cancer cell lines originating from different solid tumors. Anticancer Res. 34, 6417–6422 (2014).
Wang, Q. et al. GSTZ1 sensitizes hepatocellular carcinoma cells to sorafenib-induced ferroptosis via inhibition of NRF2/GPX4 axis. Cell Death Dis. 12, 426 (2021).
Eling, N., Reuter, L., Hazin, J., Hamacher-Brady, A. & Brady, N. R. Identification of artesunate as a specific activator of ferroptosis in pancreatic cancer cells. Oncoscience 2, 517–532 (2015).
Friedmann Angeli, J. P. et al. Inactivation of the ferroptosis regulator Gpx4 triggers acute renal failure in mice. Nat. Cell Biol. 16, 1180–1191 (2014).
Li, Q. et al. Inhibition of neuronal ferroptosis protects hemorrhagic brain. JCI Insight 2, e90777 (2017).
Liu, P. et al. Ferrostatin-1 alleviates lipopolysaccharide-induced acute lung injury via inhibiting ferroptosis. Cell. Mol. Biol. Lett. 25, 10 (2020).
Feng, Y., Madungwe, N. B., Imam Aliagan, A. D., Tombo, N. & Bopassa, J. C. Liproxstatin-1 protects the mouse myocardium against ischemia/reperfusion injury by decreasing VDAC1 levels and restoring GPX4 levels. Biochem. Biophys. Res. Commun. 520, 606–611 (2019).
Cao, Y. et al. Selective ferroptosis inhibitor liproxstatin-1 attenuates neurological deficits and neuroinflammation after subarachnoid hemorrhage. Neurosci. Bull. 37, 535–549 (2021).
Hatami, A. et al. Deuterium-reinforced linoleic acid lowers lipid peroxidation and mitigates cognitive impairment in the Q140 knock in mouse model of Huntington’s disease. FEBS J. 285, 3002–3012 (2018).
Elharram, A. et al. Deuterium-reinforced polyunsaturated fatty acids improve cognition in a mouse model of sporadic Alzheimer’s disease. FEBS J. 284, 4083–4095 (2017).
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012).
Kagan, V. E. et al. Oxidized arachidonic and adrenic PEs navigate cells to ferroptosis. Nat. Chem. Biol. 13, 81–90 (2017).
Yang, W. S. et al. Peroxidation of polyunsaturated fatty acids by lipoxygenases drives ferroptosis. Proc. Natl Acad. Sci. USA 113, E4966–E4975 (2016).
Yang, W. S. et al. Regulation of ferroptotic cancer cell death by GPX4. Cell 156, 317–331 (2014).
Shimada, K. et al. Global survey of cell death mechanisms reveals metabolic regulation of ferroptosis. Nat. Chem. Biol. 12, 497–503 (2016).
Gaschler, M. M. et al. FINO2 initiates ferroptosis through GPX4 inactivation and iron oxidation. Nat. Chem. Biol. 14, 507–515 (2018).
Wenzel, S. E. et al. PEBP1 wardens ferroptosis by enabling lipoxygenase generation of lipid death signals. Cell 171, 628–641 (2017).
Doll, S. et al. ACSL4 dictates ferroptosis sensitivity by shaping cellular lipid composition. Nat. Chem. Biol. 13, 91–98 (2017).
Gaschler, M. M. et al. Determination of the subcellular localization and mechanism of action of ferrostatins in suppressing ferroptosis. ACS Chem. Biol. 13, 1013–1020 (2018).
Gao, M. et al. Role of mitochondria in ferroptosis. Mol. Cell 73, 354–363 (2019).
Mao, C. et al. DHODH-mediated ferroptosis defence is a targetable vulnerability in cancer. Nature 593, 586–590 (2021).
Magtanong, L. et al. Exogenous monounsaturated fatty acids promote a ferroptosis-resistant cell state. Cell Chem. Biol. 26, 420–432 (2019).
Bersuker, K. et al. The CoQ oxidoreductase FSP1 acts parallel to GPX4 to inhibit ferroptosis. Nature 575, 688–692 (2019).
Doll, S. et al. FSP1 is a glutathione-independent ferroptosis suppressor. Nature 575, 693–698 (2019).
Shen, Y., Hu, F. & Min, W. Raman imaging of small biomolecules. Annu. Rev. Biophys. 48, 347–369 (2019).
Hu, F., Shi, L. & Min, W. Biological imaging of chemical bonds by stimulated Raman scattering microscopy. Nat. Methods 16, 830–842 (2019).
Hill, S. et al. Small amounts of isotope-reinforced polyunsaturated fatty acids suppress lipid autoxidation. Free Radic. Biol. Med. 53, 893–906 (2012).
Firsov, A. M. et al. Threshold protective effect of deuterated polyunsaturated fatty acids on peroxidation of lipid bilayers. FEBS J. 286, 2099–2117 (2019).
Shah, R., Shchepinov, M. S. & Pratt, D. A. Resolving the role of lipoxygenases in the initiation and execution of ferroptosis. ACS Cent. Sci. 4, 387–396 (2018).
Raefsky, S. M. et al. Deuterated polyunsaturated fatty acids reduce brain lipid peroxidation and hippocampal amyloid β-peptide levels, without discernable behavioral effects in an APP/PS1 mutant transgenic mouse model of Alzheimer’s disease. Neurobiol. Aging 66, 165–176 (2018).
Zou, Y. et al. Plasticity of ether lipids promotes ferroptosis susceptibility and evasion. Nature 585, 603–608 (2020).
Cui, W., Liu, D., Gu, W. & Chu, B. Peroxisome-driven ether-linked phospholipids biosynthesis is essential for ferroptosis. Cell Death Differ. 28, 2536–2551 (2021).
Gijón, M. A., Riekhof, W. R., Zarini, S., Murphy, R. C. & Voelker, D. R. Lysophospholipid acyltransferases and arachidonate recycling in human neutrophils. J. Biol. Chem. 283, 30235–30245 (2008).
Jiménez-López, J. M., Ríos-Marco, P., Marco, C., Segovia, J. L. & Carrasco, M. P. Alterations in the homeostasis of phospholipids and cholesterol by antitumor alkylphospholipids. Lipids Health Dis. 9, 33 (2010).
Neve, E. P. A., Boyer, C. S. & Moldéus, P. N-ethyl maleimide stimulates arachidonic acid release through activation of the signal-responsive phospholipase A2 in endothelial cells. Biochem. Pharmacol. 49, 57–63 (1995).
Loi, M., Fregno, I., Guerra, C. & Molinari, M. Eat it right: ER-phagy and recovER-phagy. Biochem. Soc. Trans. 46, 699–706 (2018).
Morishita, H. et al. Organelle degradation in the lens by PLAAT phospholipases. Nature 592, 634–638 (2021).
Liu, L. et al. Mitochondrial outer-membrane protein FUNDC1 mediates hypoxia-induced mitophagy in mammalian cells. Nat. Cell Biol. 14, 177–185 (2012).
Dierge, E. et al. Peroxidation of n-3 and n-6 polyunsaturated fatty acids in the acidic tumor environment leads to ferroptosis-mediated anticancer effects. Cell Metab. 33, 1701–1715 (2021).
Dixon, S. J. et al. Pharmacological inhibition of cystine-glutamate exchange induces endoplasmic reticulum stress and ferroptosis. eLife 3, e02523 (2014).
Lee, Y. S. et al. Ferroptotic agent-induced endoplasmic reticulum stress response plays a pivotal role in the autophagic process outcome. J. Cell. Physiol. 235, 6767–6778 (2020).
Riegman, M. et al. Ferroptosis occurs through an osmotic mechanism and propagates independently of cell rupture. Nat. Cell Biol. 22, 1042–1048 (2020).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
McQuin, C. et al. CellProfiler 3.0: next-generation image processing for biology. PLoS Biol. 16, 1–17 (2018).
Kraft, V. A. N. et al. GTP cyclohydrolase 1/tetrahydrobiopterin counteract ferroptosis through lipid remodeling. ACS Cent. Sci. 6, 41–53 (2020).
Smith, C. A., Want, E. J., O’Maille, G., Abagyan, R. & Siuzdak, G. XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 78, 779–787 (2006).
R Core Team. R: a language and environment for statistical computing (2014).
Stewart, S. A. et al. Lentivirus-delivered stable gene silencing by RNAi in primary cells. RNA 9, 493–501 (2003).
Acknowledgements
A.N.K. was funded by National Institutes of Health National Research Service Award F30AG066272. The research of B.R.S. was supported by National Cancer Institute grants P01CA87497 and R35CA209896. The research of K.A.W. was supported by the National Institute of General Medical Sciences of the National Institutes of Health (R01GM118730). The research of W.M. was supported by the National Institute of Biomedical Imaging and Bioengineering of the National Institutes of Health (R01 EB029523).
Author information
Authors and Affiliations
Contributions
A.N.K. performed all biochemical experiments with D-PUFAs. A.N.K. prepared D-PUFA samples for imaging, and F.H. and N.Q. performed SRS imaging. Relative quantification of D-PUFAs in membranes by SRS was done by N.Q. High-resolution SRS imaging samples for mitochondrial evaluation were prepared by T.H. and imaged by N.Q. Quantification of PUFA incorporation into knockdown cell lines by SRS imaging was done by F.H. A.N.K. and F.Z. designed the lipidomics experiment and prepared samples, and F.Z. performed liquid chromatography–mass spectrometry and analysis. R.N.R. and V.M.E. synthesized the FINO2 analogs. A.N.K. performed all biochemical and confocal fluorescence experiments of FINO2 analogs. SRS imaging samples of FINO2-2 were prepared by A.N.K. and imaged by F.H. and N.Q. C11 BODIPY imaging and quantification was performed by A.N.K. and B.Q. Membrane fractionation and relative PUFA quantification by mass spectrometry was performed by E.R. and N.S. Immunofluorescence staining of ACSL4 was performed by B.Q., and western blot quantification of ACSL4 in membrane fractions was done by E.R. and N.S. BQR viability and C11 BODIPY experiments were performed by B.Q. Knockdowns of lipid processing genes and experiments with lipid synthesis inhibitors were performed by A.N.K. Plasmid design was performed by A.N.K., and cloning was done by A.N.K. and M.D. ER-phagy experiments, including knockdowns, overexpressions and evaluation, were performed by A.N.K. and M.D. Experimental design and execution was overseen by W.M., K.A.W. and B.R.S. D-PUFAs and consultation were provided by M.S.S. A.N.K. drafted the manuscript, with contributions and revisions by B.R.S., W.M., K.A.W., R.N.R., F.H., F.Z., N.Q. and M.S.S.
Corresponding authors
Ethics declarations
Competing interests
B.R.S. is an inventor on patents and patent applications involving ferroptosis; co-founded and serves as a consultant to ProJenX, Inc. and Exarta Therapeutics; holds equity in Sonata Therapeutics; serves as a consultant to Weatherwax Biotechnologies Corporation and Akin Gump Strauss Hauer & Feld LLP; and receives sponsored research support from Sumitomo Dainippon Pharma Oncology. M.S.S. is the Chief Scientific Officer of Retrotrope, Inc. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Dose-response of D-PUFAs, peroxisome and mitochondrial staining of D-PUFA-treated cells, and mitochondrial/ER quantification of D-PUFAs.
a. Dose-response curves of HT-1080 cells pre-treated with varying concentrations of D-PUFAs and then treated with FINs. Data are plotted as mean ± SEM, n=3 biologically independent samples. b. HT-1080 cells treated for 24 hours with 20 µM ARA-d6 and CellLight Peroxisome-GFP, and imaged by fluorescence and SRS imaging. c. HT-1080 cells treated for 24 hours with 20 µM DHA-d10, and then stained with MitoTracker Red CMXRos and imaged by fluorescence and SRS imaging. d. High resolution SRS imaging of HT-1080 cells treated for 24 hours with 20 µM DHA-d10 with DGAT inhibitors PF-06424439 (1 µM) and A922500 (1 µM), then stained with MitoTracker Red CMXRos and ERTracker Green and imaged by fluorescence and SRS imaging. e. Quantification of arachidonic acid-d11 in mitochondrial and ER fractions isolated from HT-1080 cells treated at 20 µM for 24 hours. Values determined by high resolution mass spectrometry and plotted as normalized to internal standard. Data are plotted as mean of three biological replicates ± SEM. f. Western blotting of mitochondrial and ER fractions stained for PDI as ER marker and Cytochrome C as mitochondrial marker, representative of four experiments. Results indicate no mitochondrial contamination of ER fraction, but indicate ER contamination of mitochondrial fraction.
Extended Data Fig. 2 Knockdown of PUFA-related genes shows some impact on D-PUFA potency, but no observable decrease in incorporation.
a. Dose-response curves of stable non-targeting (NT) or ACSL knockdown HT-1080 cells pretreated with EtOH or D-PUFAs and then treated with RSL3. Data are represented as mean ± SEM, n=3. b. Dose-response curves of stable nontargeting (NT) or AGPAT3 knockdown HT-1080 cells pretreated with EtOH or D-PUFAs and then treated with RSL3 or IKE. Data are represented as mean ± SEM, n=3. c. qPCR data of stable shRNA knockdowns. Data are represented as mean of three technical replicates ± upper and lower limit of 95% confidence interval. d. SRS images of shNT, shACSL5, and shAGPAT3 HT-1080 cells treated with DHA-d10 (10 µM) (left), and quantification of their signal intensity (right), Data are plotted as mean ± SEM, n=3.
Extended Data Fig. 3 Thimerosal, Miltefosine, and N-Ethyl Maleimide do not impact anti-ferroptotic potency of D-PUFAs, and knockdown of ER-phagy related genes SEC62, RTN3, and FAM134B did not result in apparently altered ER area.
a. Dose-response curves of HT-1080 cells treated with vehicle (water) or 400 nM thimerosal, and subsequently treated with vehicle (EtOH) or PUFAs 4 hours later, followed by varying concentrations of IKE and RSL3 24 hours later. Data are represented as mean ± SEM, n=3. b. Dose-response curves of HT-1080 cells treated with vehicle (water) or 7.5 µM miltefosine, and subsequently treated with vehicle (EtOH) or PUFAs 4 hours later, followed by varying concentrations of RSL3 24 hours later. Data are represented as mean ± SEM, n=3. c. Dose-response curves of HT-1080 cells treated with D-PUFAs and either pretreated, cotreated, or post-treated with vehicle (EtOH) or 4.5 µM NME, and subsequently treated with varying concentrations IKE and RSL3. In the post-treatment experiments, media containing PUFAs was removed before NME was added. Data are represented as mean ± SEM, n=3. d. Dose-response curves of stable shNT, shSEC62, shRTN3, and shFAM134B HT-1080 cells treated with varying doses of IKE, RSL3, FIN56, and FINO2. Data are represented as mean ± SEM, n=3. e. ER area of ER-Phagy knockdown cell lines as compared to control. Cells were stained with ERTracker Blue-White, imaged with confocal microscopy, and their ER areas were measured using the CellProfiler software. Individual ER areas are shown for each cell, as well as the mean ± SD. Sample sizes (number of cells) are as follows: shNT n=123, shFAM134B n=98, shRTN3 n=60, shSEC62 n=26. f. qPCR analysis of knockdowns of ER-phagy genes in HT-1080 cells. Data are represented as mean of three technical replicates ± upper and lower limit of 95% confidence interval.
Extended Data Fig. 4 Overexpression and targeting of organelle-phagy related genes did not result in apparently altered ER area.
a. Confocal fluorescence images of HT-1080 cells overexpressing GFP, GFP-PLA2G16, GFP-FUNDC1 (as well as the cytoplasmic N-terminal FUNDC1 sequence), with and without C terminal-cytochrome b5 ER targeting signals, stained with ERTracker Red. GFP channels show protein distribution as all overexpressed proteins are tagged with GFP. Representative images of at least six images per sample are shown. b. Dose-response curves of overexpression cell lines with varying doses of RSL3. Data are represented as mean ± SEM, n=3 biologically independent samples. c. ER area overexpression cell lines as compared to control. Cells were stained with ERTracker Red, imaged with confocal microscopy, and their ER areas were measured using the CellProfiler software. Violin quartile plots are shown. Sample sizes (number of cells) are as follows: GFP n=358, GFP-cyb5 n=478, PLA2G16 n=353, PLA2G16-cyb5 n=292, FUNDC1 n=354, FUNDC1-cyb5 n=212, Nterm-FUNDC1-cyb5 n=399.
Extended Data Fig. 5 Subcellular localization and effect on ferroptosis of myristic acid and cholesterol, overexpression and distribution of ACSL4.
a. Structure and SRS image of myristic acid-d27 (20 µM) in HT-1080 cells. b. Structure and SRS image of cholesterol-d6 (20 µM) in HT-1080 cells. c. Fluorescence and confocal imaging of cholesterol-d6 to evaluate its subcellular localization. d. Solution Raman spectra of the FAs and cholesterol used in these experiments. e. Dose-response curves of HT-1080 cells pretreated with either MA-d27 or cholesterol-d6 (20 µM) followed by varying concentrations of RSL3. Data are represented as mean ± SEM, n=3. f. Western blot of HT-1080 cells overexpressing GFP (control) or GFP-ACSL4. ACSL4 antibody was used, with actin as a control. g. Immunofluorescence staining of HT-1080 cells labeled for ACSL4 (anti-ACSL4 antibody), ER (anti-calnexin antibody), and nucleus (DAPI). Individual channels and overlay is shown. h. Western blotting of mitochondrial and ER HT-1080 fractions stained for ACSL4 PDI (ER), and Cytochrome C (mitochondria), indicating presence in both fractions, representative of two experiments.
Extended Data Fig. 6 FINO2-0 and FINO2-2 accumulate in the ER, and FINO2-1/FINO2-3/FINO2-4 do not accumulate in the plasma membrane.
a. Structure of FINO2-0. b. Dose-response curve of HT-1080 cells treated with FINO2-0 ± fer-1. Data are represented as mean ± SEM, n=3. c. Confocal fluorescence images of HT-1080 cells treated for 3 hours with 3 µM FINO2-1 alone or with 3 µM fer-1. d. Confocal fluorescence image of HT-1080 cells treated with 3 µM FINO2-0. f. Confocal fluorescence imaging of HT-1080 cells treated with FINO2-1 (3 µM) and fer-1 (3 µM) for 3 hours, co-stained with BODIPY TR ceramide. g. Structure of FINO2-2. h. Dose-response curve of HT-1080 cells treated with FINO2-2 ± fer-1. Data are represented as mean ± SEM, n=3. i. SRS and fluorescence imaging of HT-1080 cells treated with 20 µM FINO2-2 and 2 µM fer-1, and stained with Nile Red and Lysotracker green. j. Confocal fluorescence image of HT-1080 cells treated with 3 µM FINO2-1 and 3 µM fer-1, and costained with CellMask Deep Red. k. Confocal fluorescence image of HT-1080 cells treated with 3 µM FINO2-3 or 3 µM FINO2-4, and 3 µM fer-1, and co-stained with CellMask Deep Red.
Extended Data Fig. 7 Analogs of FINO2 redistribute throughout the cell.
a. Structures of analogs of FINO2. b. Confocal fluorescence images of HT-1080 cells treated with FINO2-5 (3 µM) or FINO2-6 (3 µM) and fer-1 (3 µM), and co-stained with LysoTracker Red. c. Dose-response curves of HT-1080 cells treated with fixed concentrations of FINO2 analogs (10 µM) and varying concentrations of ferroptosis inhibitors. Lysosome-directed ferrostatin previously published by Gaschler et al.27. Data are represented as mean ± SEM, n=3. d. Confocal fluorescence image of HT-1080 cells treated with FINO2-7 (3 µM) and fer-1 (3 µM). e. Dose-response curves of HT-1080 cells comparing treatment with FINO2-1 and FINO2-3 or FINO2-4. Data are represented as mean ± SEM, n=3.
Extended Data Fig. 8 Ferroptosis induced by DHODH inhibition is amplified by ER peroxidation, ferroptosis induced by IKE results in ER membrane peroxidation followed by PM peroxidation, early perinuclear lipid peroxidation occurs in the ER.
a. Brequinar (BQR) induces ferroptosis in HT-1080 cells as rescued by fer-1, and cotreatment with BQR (500 µM) increases sensitivity to RSL3 in HT-1080 cells. Data are represented as mean ± SEM, n=3. b. Dose-response curve of HT-1080 cells cotreated with 1 µM of FINO2-1, FINO2-3, or FINO2-4. Data are represented as mean ± SEM, n=3. c. HT-1080 cells treated with DMSO or IKE (10 µM) for 6 hours and stained with C11 BODIPY (oxidized and reduced overlay), CellMask Deep Red, and ERTracker Blue-White. d. HT-1080 cells treated with DMSO or IKE (10 µM) for 10 hours and stained with C11 BODIPY (oxidized and reduced overlay), CellMask Deep Red, and ERTracker Blue-White. e. Correlation (Manders coefficient) of C11 BODIPY oxidized signal with ERTracker Blue-White or MitoTracker Deep Red in HT-1080 cells treated with either RSL3, IKE, FINO2, or FIN56 at designated timepoints. Data are represented as mean ± SEM, each individual point represents an image of multiple cells. Number of images for each condition are RSL3 2 hour (n=3), 5 hour (n=2); IKE 2 hour (n=7), 5 hour (n=5), FINO2 2.5 hour (n=6), 4.5 hour (n=9), FIN56 2 hour (n=6), 4 hour (n=5), 5 hour (n=2). Ordinary one way ANOVA with Tukey’s test for multiple comparisons was used with p values of: RSL3 (0.0016, 0.6120), IKE (0.0023, 0.1279), FINO2 (<0.0001, <0.0001), FIN56 (0.0034, 0.0005, >0.9999). f. Correlation (Manders coefficient) of C11 BODIPY oxidized signal with ERTracker Blue-White or MitoTracker Deep Red in HT-1080 cells treated with either RSL3 (1 µM), BQR (1 mM) or both at designated timepoints. Data are represented as mean ± SEM, each individual point represents an image of multiple cells. Number of images for each condition are RSL3 1.5 hour (n=4), 2 hour (n=2); BQR 2 hour (n=3), 2.5 hour (n=3); RSL3 + BQR 1.5 hour (n=2), 2 hour (n=2). Two-sided unpaired t test was used with p values of: RSL3 (<0.0001, 0.0178), BQR (0.0006, 0.1011), RSL3 + BQR (0.0117, 0.2027). For all panels, GraphPad Prism (GP) P value style of 0.1234 (ns), 0.0332 (*), 0.0021 (**), 0.0002 (***), <0.0001 (****).
Supplementary information
Supplementary Information
Supplementary Tables 1–3 and Supplementary Note containing synthetic methods
Source data
Source Data Fig. 1
Statistical source data
Source Data Fig. 2
Statistical source data
Source Data Fig. 3
Statistical source data
Source Data Fig. 4
Statistical source data
Source Data Fig. 5
Statistical source data
Source Data Fig. 6
Statistical source data
Source Data Extended Data Fig. 1
Statistical source data
Source Data Extended Data Fig. 1
Unprocessed western blot
Source Data Extended Data Fig. 2
Statistical source data
Source Data Extended Data Fig. 3
Statistical source data
Source Data Extended Data Fig. 4
Statistical source data
Source Data Extended Data Fig. 5
Statistical source data
Source Data Extended Data Fig. 5
Unprocessed western blot
Source Data Extended Data Fig. 6
Statistical source data
Source Data Extended Data Fig. 7
Statistical source data
Source Data Extended Data Fig. 8
Statistical source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
von Krusenstiern, A.N., Robson, R.N., Qian, N. et al. Identification of essential sites of lipid peroxidation in ferroptosis. Nat Chem Biol 19, 719–730 (2023). https://doi.org/10.1038/s41589-022-01249-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-022-01249-3
This article is cited by
-
Ferroptotic therapy in cancer: benefits, side effects, and risks
Molecular Cancer (2024)
-
Tumor-repopulating cells evade ferroptosis via PCK2-dependent phospholipid remodeling
Nature Chemical Biology (2024)
-
Unlocking ferroptosis in prostate cancer — the road to novel therapies and imaging markers
Nature Reviews Urology (2024)
-
The cell biology of ferroptosis
Nature Reviews Molecular Cell Biology (2024)
-
Photoswitchable polyynes for multiplexed stimulated Raman scattering microscopy with reversible light control
Nature Communications (2024)