Abstract
The RNA-guided CRISPR-associated (Cas) nucleases are versatile tools for genome editing in various organisms. The large sizes of the commonly used Cas9 and Cas12a nucleases restrict their flexibility in therapeutic applications that use the cargo-size-limited adeno-associated virus delivery vehicle. More compact systems would thus offer more therapeutic options and functionality for this field. Here, we report a miniature class 2 type V-F CRISPR-Cas genome-editing system from Acidibacillus sulfuroxidans (AsCas12f1, 422 amino acids). AsCas12f1 is an RNA-guided endonuclease that recognizes 5′ T-rich protospacer adjacent motifs and creates staggered double-stranded breaks to target DNA. We show that AsCas12f1 functions as an effective genome-editing tool in both bacteria and human cells using various delivery methods, including plasmid, ribonucleoprotein and adeno-associated virus. The small size of AsCas12f1 offers advantages for cellular delivery, and characterizations of AsCas12f1 may facilitate engineering more compact genome-manipulation technologies.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
NGS data are available at the NCBI Sequence Read Archive (SRA) under accession codes SRX9277304, SRX9277305, SRX9277306, SRX10828545, SRX10828546, SRX10828547 and SRX10839449. Plasmids can be accessed in Addgene under accession codes 171609, 171610, 171611, 171612, 171613 and 171614. Source data are provided with this paper.
Code availability
The custom Python scripts for calculating sequence frequencies in the PAM depletion assay are available at GitHub (https://github.com/atlasbioinfo/midseq). The computational pipeline of CIRCLE-seq to identify genome-wide off-target sites for AsCas12f1 is available at GitHub (https://github.com/tsailabSJ/circleseq).
References
Makarova, K. S. et al. Evolutionary classification of CRISPR-Cas systems: a burst of class 2 and derived variants. Nat. Rev. Microbiol. 18, 67–83 (2020).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163, 759–771 (2015).
Shan, Q. et al. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat. Biotechnol. 31, 686–688 (2013).
Wang, H., La Russa, M. & Qi, L. S. CRISPR/Cas9 in genome editing and beyond. Annu. Rev. Biochem. 85, 227–264 (2016).
Ran, F. A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Anzalone, A. V., Koblan, L. W. & Liu, D. R. Genome editing with CRISPR-Cas nucleases, base editors, transposases and prime editors. Nat. Biotechnol. 38, 824–844 (2020).
Amoasii, L. et al. Gene editing restores dystrophin expression in a canine model of Duchenne muscular dystrophy. Science 362, 86–91 (2018).
Pickar-Oliver, A. & Gersbach, C. A. The next generation of CRISPR-Cas technologies and applications. Nat. Rev. Mol. Cell Biol. 20, 490–507 (2019).
Nelson, C. E. et al. Long-term evaluation of AAV-CRISPR genome editing for Duchenne muscular dystrophy. Nat. Methods 25, 427–432 (2019).
Wang, D., Tai, P. W. L. & Gao, G. Adeno-associated virus vector as a platform for gene therapy delivery. Nat. Rev. Drug Discov. 18, 358–378 (2019).
Liu, J. J. et al. CasX enzymes comprise a distinct family of RNA-guided genome editors. Nature 566, 218–223 (2019).
Pausch, P. et al. CRISPR-CasΦ from huge phages is a hypercompact genome editor. Science 369, 333–337 (2020).
Harrington, L. B. et al. Programmed DNA destruction by miniature CRISPR-Cas14 enzymes. Science 362, 839–842 (2018).
Karvelis, T. et al. PAM recognition by miniature CRISPR-Cas12f nucleases triggers programmable double-stranded DNA target cleavage. Nucleic Acids Res. 48, 5016–5023 (2020).
Strecker, J. et al. Engineering of CRISPR-Cas12b for human genome editing. Nat. Commun. 10, 212 (2019).
Takeda, S. N. et al. Structure of the miniature type V-F CRISPR-Cas effector enzyme. Mol. Cell 81, 558–570 (2021).
Xiao, R., Li, Z., Wang, S., Han, R. & Chang, L. Structural basis for substrate recognition and cleavage by the dimerization-dependent CRISPR-Cas12f nuclease. Nucleic Acids Res. 49, 4120–4128 (2021).
Lei, C. et al. The CCTL (Cpf1-assisted Cutting and Taq DNA ligase-assisted Ligation) method for efficient editing of large DNA constructs in vitro. Nucleic Acids Res. 45, e74 (2017).
Chen, J. S. et al. CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science 360, 436–439 (2018).
Teng, F. et al. CDetection: CRISPR-Cas12b-based DNA detection with sub-attomolar sensitivity and single-base specificity. Genome Biol. 20, 132 (2019).
Harrington, L. B. et al. A scoutRNA is required for some type V CRISPR-Cas systems. Mol. Cell 79, 416–424 (2020).
Yan, W. X. et al. Functionally diverse type V CRISPR-Cas systems. Science 363, 88–91 (2019).
Wu, Z. et al. Strategies for developing CRISPR-based gene editing methods in bacteria. Small Methods 4, 1900560 (2020).
Perrenoud, A. & Sauer, U. Impact of global transcriptional regulation by ArcA, ArcB, Cra, Crp, Cya, Fnr and Mlc on glucose catabolism in Escherichia coli. J. Bacteriol. 187, 3171–3179 (2005).
Wyres, K. L. & Holt, K. E. Klebsiella pneumoniae as a key trafficker of drug resistance genes from environmental to clinically important bacteria. Curr. Opin. Microbiol. 45, 131–139 (2018).
Hu, Z. et al. A compact Cas9 ortholog from Staphylococcus auricularis (SauriCas9) expands the DNA targeting scope. PLoS Biol. 18, e3000686 (2020).
Grimm, D. et al. In vitro and in vivo gene therapy vector evolution via multispecies interbreeding and retargeting of adeno-associated viruses. J. Virol. 82, 5887–5911 (2008).
Tsai, S. Q. et al. CIRCLE-seq: a highly sensitive in vitro screen for genome-wide CRISPR-Cas9 nuclease off-targets. Nat. Methods 14, 607–614 (2017).
Karvelis, T. et al. Rapid characterization of CRISPR-Cas9 protospacer adjacent motif sequence elements. Genome Biol. 16, 253 (2015).
Quan, J. & Tian, J. Circular polymerase extension cloning for high-throughput cloning of complex and combinatorial DNA libraries. Nat. Protoc. 6, 242–251 (2011).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Zhang, Y. et al. Catalytic-state structure and engineering of Streptococcus thermophilus Cas9. Nat. Catal. 3, 813–823 (2020).
Chen, W. et al. CRISPR/Cas9-based genome editing in Pseudomonas aeruginosa and cytidine deaminase-mediated base editing in Pseudomonas species. Iscience 6, 222–231 (2018).
Wang, Y. et al. CRISPR-Cas9 and CRISPR-assisted cytidine deaminase enable precise and efficient genome editing in Klebsiella pneumoniae. Appl. Environ. Microbiol. 84, e01834 (2018).
Park, J., Lim, K., Kim, J. S. & Bae, S. Cas-analyzer: an online tool for assessing genome editing results using NGS data. Bioinformatics 33, 286–288 (2017).
Lazzarotto, C. R. et al. Defining CRISPR-Cas9 genome-wide nuclease activities with CIRCLE-seq. Nat. Protoc. 13, 2615–2642 (2018).
Acknowledgements
This work was supported by grants 21922705 (Q.J.), 91753127 (Q.J.) and 2207783 (Q.J.) from the National Natural Science Foundation of China; 19QA1406000 (Q.J.) from the Shanghai Committee of Science and Technology, China; 2017YFA0506800 (Q.J.) from the National Key R&D Program of China; EKPG21-18 (Q.J.) from the Emergency Key Program of Guangzhou Laboratory; and 2019M651627 (Z.W.) from the China Postdoctoral Science Foundation. We thank C. He at The University of Chicago for helpful discussions. We thank R. Liu, X. Huang and J. Li at ShanghaiTech University for kindly providing HEK293 cells and valuable advice regarding cell culture experiments. We thank G. Luo and Z. Ren at Sun Yat-sen University for helpful advice in RNA-seq experimental design. We thank J. Li, Y. Xiong and X. Ren from the Molecular and Cell Biology Core Facility (MCBCF) at the School of Life Science and Technology, ShanghaiTech University, for providing technical support.
Author information
Authors and Affiliations
Contributions
Z.W., Y.Z. and Q.J. conceived the initial study. Z.W. and Yujue Wang performed the PAM depletion assay. Z.W. and H.Y. conducted the small RNA-seq and computational analyses. Z.W., Y.Z., Yannan Wang, F.L., D.P. and W.C. performed plasmid construction, protein purification and biochemical experiments. Z.W., Yujue Wang and Yannan Wang performed the genome editing in bacteria. Z.W., H.Y., D.P. and Yujue Wang performed the genome editing in human cells and NGS data analyses. Z.W., H.N. and D.P. designed the AAV vector and performed the AAV-based genome editing. Z.W. and C.L. analyzed the sequence homology among Cas12 nucleases and built the structure model. Z.W., H.Y. and D.P. performed the CIRCLE-seq. Z.W. and Q.J. prepared the figures and wrote the manuscript. The manuscript was reviewed and approved by all authors.
Corresponding author
Ethics declarations
Competing interests
Q.J., Z.W., Y.Z., Yujue Wang and Yannan Wang have filed a patent application (PCT/CN2020/132759) related to this work through ShanghaiTech University. The remaining authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Optimization of the sgRNA for AsCas12f1.
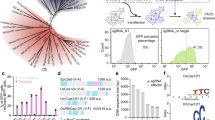
a, The predicted secondary structure of sgRNA_v1 and the truncation information. The gray shaded tri-guanosine is added to the beginning of sgRNA to maximize the RNA production. b, in vitro dsDNA cleavage assay for different truncations of sgRNA. c, Bacterial genome targeting using different truncations of sgRNA. d, The predicted secondary structure of sgRNA_v2.
Extended Data Fig. 2 Sequence and structural analysis of AsCas12f1 and its mutations in the RuvC active site.
a, Multiple sequence alignment of Cas12 nucleases. b, Structure model of AsCas12f1 was built using the cryo-EM structure of Cas14a1/sgRNA/dsDNA (PDB ID: 7C7L) as a template using SWISS-MODEL web server with its default parameters. c, D225A and E324A completely abolished the nuclease activity of AsCas12f1, whereas R383A and D401A retained minor NTS nickase activity at 18-nt downstream the PAM.
Extended Data Fig. 3 AsCas12f1 cleaves ssDNA in a PAM-independent manner.
a, Cleavage pattern for complementary-PAM (cPAM)-containing ssDNA. b, Cleavage pattern for non-cPAM-containing ssDNA. c, Cleavage pattern for PAM-containing staggered substrate. d, Cleavage pattern for non-PAM-containing staggered substrate. e, Non-targeting sgRNA cannot mediate targeted cleavage on short ssDNA substrate. f, AsCas12f1 cannot non-specifically cleave short ssDNA substrate.
Extended Data Fig. 4 Flow cytometry gating strategies.
a, b, Gates used for the analysis of flow cytometric data shown in Fig. 3c. Gate 1 is used to remove the debris from the cell population, gate 2 specifies single cells from cell aggregates, and gate 3 identifies the cells with GFP fluorescence. Gates were set on the control sample (a) and was then kept constant for the experimental group (b).
Supplementary information
Supplementary Information
Supplementary Figs. 1–8 and Tables 1–3.
Supplementary Data 1
Primers and substrates.
Supplementary Data 2
Predicted and experimental off-targets
Source data
Source Data Fig. 1
Unmodified gels.
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Unmodified gel.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 1
Unmodified gel.
Source Data Extended Data Fig. 2
Unmodified gels.
Source Data Extended Data Fig. 3
Unmodified gels.
Rights and permissions
About this article
Cite this article
Wu, Z., Zhang, Y., Yu, H. et al. Programmed genome editing by a miniature CRISPR-Cas12f nuclease. Nat Chem Biol 17, 1132–1138 (2021). https://doi.org/10.1038/s41589-021-00868-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-021-00868-6
This article is cited by
-
Recent advances in CRISPR-based functional genomics for the study of disease-associated genetic variants
Experimental & Molecular Medicine (2024)
-
Discovery and structural mechanism of DNA endonucleases guided by RAGATH-18-derived RNAs
Cell Research (2024)
-
Engineering a transposon-associated TnpB-ωRNA system for efficient gene editing and phenotypic correction of a tyrosinaemia mouse model
Nature Communications (2024)
-
Molecular basis and engineering of miniature Cas12f with C-rich PAM specificity
Nature Chemical Biology (2024)
-
Targeted Gene Insertion: The Cutting Edge of CRISPR Drug Development with Hemophilia as a Highlight
BioDrugs (2024)