Abstract
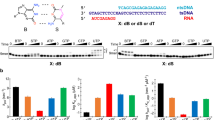
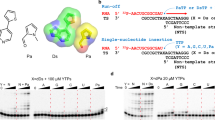
The development of unnatural base pairs (UBPs) has greatly increased the information storage capacity of DNA, allowing for transcription of unnatural RNA by the heterologously expressed T7 RNA polymerase (RNAP) in Escherichia coli. However, little is known about how UBPs are transcribed by cellular RNA polymerases. Here, we investigated how synthetic unnatural nucleotides, NaM and TPT3, are recognized by eukaryotic RNA polymerase II (Pol II) and found that Pol II is able to selectively recognize UBPs with high fidelity when dTPT3 is in the template strand and rNaMTP acts as the nucleotide substrate. Our structural analysis and molecular dynamics simulation provide structural insights into transcriptional processing of UBPs in a stepwise manner. Intriguingly, we identified a novel 3′-RNA binding site after rNaM addition, termed the swing state. These results may pave the way for future studies in the design of transcription and translation strategies in higher organisms with expanded genetic codes.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Crystal structure coordinates of apo dTPT3, dTPT3–rNaMTP and dTPT3–rNaM Pol II complexes are deposited in the Protein Databank database (PDB, https://www.rcsb.org) with accession nos. 7KED, 7KEE and 7KEF, respectively. Source data are provided with this paper.
References
Malyshev, D. A. & Romesberg, F. E. The expanded genetic alphabet. Angew. Chem. Int. Ed. Engl. 54, 11930–11944 (2015).
Seo, Y. J., Matsuda, S. & Romesberg, F. E. Transcription of an expanded genetic alphabet. J. Am. Chem. Soc. 131, 5046–5047 (2009).
Feldman, A. W. et al. Optimization of replication, transcription, and translation in a semi-synthetic organism. J. Am. Chem. Soc. 141, 10644–10653 (2019).
Piccirilli, J. A., Krauch, T., Moroney, S. E. & Benner, S. A. Enzymatic incorporation of a new base pair into DNA and RNA extends the genetic alphabet. Nature 343, 33–37 (1990).
Ohtsuki, T. et al. Unnatural base pairs for specific transcription. Proc. Natl Acad. Sci. USA 98, 4922–4925 (2001).
Betz, K. et al. KlenTaq polymerase replicates unnatural base pairs by inducing a Watson–Crick geometry. Nat. Chem. Biol. 8, 612–614 (2012).
Malyshev, D. A. et al. A semi-synthetic organism with an expanded genetic alphabet. Nature 509, 385–388 (2014).
Zhang, Y. et al. A semisynthetic organism engineered for the stable expansion of the genetic alphabet. Proc. Natl Acad. Sci. USA 114, 1317–1322 (2017).
Zhang, Y. et al. A semi-synthetic organism that stores and retrieves increased genetic information. Nature 551, 644–647 (2017).
Matsuda, S. et al. Efforts toward expansion of the genetic alphabet: structure and replication of unnatural base pairs. J. Am. Chem. Soc. 129, 10466–10473 (2007).
Malyshev, D. A. et al. Solution structure, mechanism of replication, and optimization of an unnatural base pair. Chemistry 16, 12650–12659 (2010).
Betz, K. et al. Structural insights into DNA replication without hydrogen bonds. J. Am. Chem. Soc. 135, 18637–18643 (2013).
Li, L. et al. Natural-like replication of an unnatural base pair for the expansion of the genetic alphabet and biotechnology applications. J. Am. Chem. Soc. 136, 826–829 (2014).
Zhou, A. X., Dong, X. & Romesberg, F. E. Transcription and reverse transcription of an expanded genetic alphabet in vitro and in a semisynthetic organism. J. Am. Chem. Soc. 142, 19029–19032 (2020).
Seo, Y. J., Malyshev, D. A., Lavergne, T., Ordoukhanian, P. & Romesberg, F. E. Site-specific labeling of DNA and RNA using an efficiently replicated and transcribed class of unnatural base pairs. J. Am. Chem. Soc. 133, 19878–19888 (2011).
Liu, X., Bushnell, D. A. & Kornberg, R. D. RNA polymerase II transcription: structure and mechanism. Biochim. Biophys. Acta 1829, 2–8 (2013).
Werner, F. & Grohmann, D. Evolution of multisubunit RNA polymerases in the three domains of life. Nat. Rev. Microbiol. 9, 85–98 (2011).
Gout, J. F. et al. The landscape of transcription errors in eukaryotic cells. Sci. Adv. 3, e1701484 (2017).
Brueckner, F., Hennecke, U., Carell, T. & Cramer, P. CPD damage recognition by transcribing RNA polymerase II. Science 315, 859–862 (2007).
Damsma, G. E., Alt, A., Brueckner, F., Carell, T. & Cramer, P. Mechanism of transcriptional stalling at cisplatin-damaged DNA. Nat. Struct. Mol. Biol. 14, 1127–1133 (2007).
Walmacq, C. et al. Mechanism of translesion transcription by RNA polymerase II and its role in cellular resistance to DNA damage. Mol. Cell 46, 18–29 (2012).
Walmacq, C. et al. Mechanism of RNA polymerase II bypass of oxidative cyclopurine DNA lesions. Proc. Natl Acad. Sci. USA 112, E410–E419 (2015).
Wang, W., Walmacq, C., Chong, J., Kashlev, M. & Wang, D. Structural basis of transcriptional stalling and bypass of abasic DNA lesion by RNA polymerase II. Proc. Natl Acad. Sci. USA 115, E2538–E2545 (2018).
Oh, J. et al. RNA polymerase II stalls on oxidative DNA damage via a torsion-latch mechanism involving lone pair-pi and CH-pi interactions. Proc. Natl Acad. Sci. USA 117, 9338–9348 (2020).
Sydow, J. F. et al. Structural basis of transcription: mismatch-specific fidelity mechanisms and paused RNA polymerase II with frayed RNA. Mol. Cell 34, 710–721 (2009).
Wang, D. et al. Structural basis of transcription: backtracked RNA polymerase II at 3.4 angstrom resolution. Science 324, 1203–1206 (2009).
Gnatt, A. L., Cramer, P., Fu, J., Bushnell, D. A. & Kornberg, R. D. Structural basis of transcription: an RNA polymerase II elongation complex at 3.3 A resolution. Science 292, 1876–1882 (2001).
Oh, J., Xu, J., Chong, J. & Wang, D. Structural and biochemical analysis of DNA lesion-induced RNA polymerase II arrest. Methods 159–160, 29–34 (2019).
Batada, N. N., Westover, K. D., Bushnell, D. A., Levitt, M. & Kornberg, R. D. Diffusion of nucleoside triphosphates and role of the entry site to the RNA polymerase II active center. Proc. Natl Acad. Sci. USA 101, 17361–17364 (2004).
Westover, K. D., Bushnell, D. A. & Kornberg, R. D. Structural basis of transcription: nucleotide selection by rotation in the RNA polymerase II active center. Cell 119, 481–489 (2004).
Wang, D., Bushnell, D. A., Westover, K. D., Kaplan, C. D. & Kornberg, R. D. Structural basis of transcription: role of the trigger loop in substrate specificity and catalysis. Cell 127, 941–954 (2006).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1, 19–25 (2015).
Lindorff-Larsen, K. et al. Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins 78, 1950–1958 (2010).
Case, D. A. et al. Amber 13 (Univ. of California, 2012).
Carvalho, A. T., Fernandes, P. A. & Ramos, M. J. The catalytic mechanism of RNA polymerase II. J. Chem. Theory Comput. 7, 1177–1188 (2011).
Wang, B., Opron, K., Burton, Z. F., Cukier, R. I. & Feig, M. Five checkpoints maintaining the fidelity of transcription by RNA polymerases in structural and energetic details. Nucleic Acids Res. 43, 1133–1146 (2015).
Svetlov, V. & Nudler, E. Basic mechanism of transcription by RNA polymerase II. Biochim. Biophys. Acta 1829, 20–28 (2013).
Yang, W., Lee, J. Y. & Nowotny, M. Making and breaking nucleic acids: two-Mg2+-ion catalysis and substrate specificity. Mol. Cell 22, 5–13 (2006).
Da, L. T., Wang, D. & Huang, X. Dynamics of pyrophosphate ion release and its coupled trigger loop motion from closed to open state in RNA polymerase II. J. Am. Chem. Soc. 134, 2399–2406 (2012).
Cheung, A. C. & Cramer, P. Structural basis of RNA polymerase II backtracking, arrest and reactivation. Nature 471, 249–253 (2011).
Yin, Y. W. & Steitz, T. A. The structural mechanism of translocation and helicase activity in T7 RNA polymerase. Cell 116, 393–404 (2004).
Temiakov, D. et al. Structural basis for substrate selection by t7 RNA polymerase. Cell 116, 381–391 (2004).
Huang, J., Brieba, L. G. & Sousa, R. Misincorporation by wild-type and mutant T7 RNA polymerases: identification of interactions that reduce misincorporation rates by stabilizing the catalytically incompetent open conformation. Biochemistry 39, 11571–11580 (2000).
Xu, L. et al. RNA polymerase II transcriptional fidelity control and its functional interplay with DNA modifications. Crit. Rev. Biochem. Mol. Biol. 50, 503–519 (2015).
Silva, D. A. et al. Millisecond dynamics of RNA polymerase II translocation at atomic resolution. Proc. Natl Acad. Sci. USA 111, 7665–7670 (2014).
Huang, X. et al. RNA polymerase II trigger loop residues stabilize and position the incoming nucleotide triphosphate in transcription. Proc. Natl Acad. Sci. USA 107, 15745–15750 (2010).
Zhou, A. X., Sheng, K., Feldman, A. W. & Romesberg, F. E. Progress toward eukaryotic semisynthetic organisms: translation of unnatural codons. J. Am. Chem. Soc. 141, 20166–20170 (2019).
Battye, T. G., Kontogiannis, L., Johnson, O., Powell, H. R. & Leslie, A. G. iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr. D. Biol. Crystallogr. 67, 271–281 (2011).
Kabsch, W. Xds. Acta Crystallogr. D. Biol. Crystallogr. 66, 125–132 (2010).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. D. Biol. Crystallogr. 69, 1204–1214 (2013).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Meagher, K. L., Redman, L. T. & Carlson, H. A. Development of polyphosphate parameters for use with the AMBER force field. J. Comput. Chem. 24, 1016–1025 (2003).
M. J. Frisch et al. Gaussian 09, Revision A.02 (Gaussian, Inc., 2016).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Berendsen, H. J. C., Postma, J. P. M., Vangunsteren, W. F., Dinola, A. & Haak, J. R. Molecular-dynamics with coupling to an external bath. J. Chem. Phys. 81, 3684–3690 (1984).
Parrinello, M. & Rahman, A. Polymorphic transitions in single-crystals – a new molecular-dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Acknowledgements
This work was supported by the National Institutes of Health (nos. R01 GM102362 to D.W., GM118178 to F.E.R. and GM128376 to R.J.K.). R.K. acknowledges supported from NASA Exobiology (no. NNX14AP59G). F.E.R. acknowledges support from Synthorx, a Sanofi company. X.H. acknowledges support from the Padma Harilela Endowment fund.
Author information
Authors and Affiliations
Contributions
F.E.R. and D.W. conceived the project. J.O. and W.W. performed structural analysis. I.C.U. and X.H. performed MD simulation. J.O., J.S., J.X., J.C. and L.X. performed biochemistry experiments. A.W.F., R.J.K., R.K. and F.E.R. prepared unnatural DNA templates and nucleotide triphosphate. J.O. and D.W. performed data analysis. D.W. supervised different aspects of the work. J.O., D.W. and F.E.R. wrote the manuscript, with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Chemical Biology thanks Seth Darst, Xianyang Fang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Electron density map of rNaMTP or rNaM.
a, Unbiased 2Fo-Fc omit electron density map of rNaMTP is contoured at 1.2 σ. b, Unbiased 2Fo-Fc omit electron density map of rNaM is contoured at 1.2 σ.
Extended Data Fig. 2 The relative stability of NTPs to maintain good activation geometry is plotted as a function of total simulation time.
The relative stability is defined as the stability of NTPs in comparison with rNaMTP to maintain good activation geometry, -ln(pNTP/prNaMTP), in energy unit (RT). pNTP and prNaMTP are the percentage of frames with good activation in NTP and rNaMTP simulations, respectively. The criteria for good activation geometry are 3.0 Å ≤ distance between O3′ - Pα ≤ 3.5 Å and 7.0 Å ≤ base pair distance (rNaMTP, ATP or GTP) ≤ 9.0 Å, 6.0 Å ≤ base pair distance (CTP, UTP) ≤ 8.0 Å. The plot shows that the simulation has converged as the order of stability among NTPs remains the same regardless of the simulation time. Importantly, rNaMTP indeed is the most stable substrate when dTPT3 is the template DNA. The data are shown as mean values ± standard deviation, which were calculated by bootstrapping of N independent production MD simulations (N=4, 8, 12, 16 for data at the time point of 200, 400, 600, 800 ns in the x-axis, respectively).
Extended Data Fig. 3 MD simulation of ATP and rTPT3TP at A site across dNaM (with both Mg2+ ion A & Mg2+ ion B).
MD simulation of ATP and rTPT3TP at A site across dNaM (with both Mg2+ ion A & Mg2+ ion b). (a and b) Left panel: two dimensional heatmap plot of the base pairing geometry. Base pair distance is the distance between center of mass of dNaM and NTPs. We observed strong localization of simulation frames in the dNaM-rTPT3TP pair, while ATP was highly dispersed both in distance and angle. Right panel: Distance of nucleophilic attack. Distribution of simulation frames sorted by the distance between Pα of incoming NTP and O3´of terminal RNA is plotted. Good activation geometry (3.0 Å ≤ distance between O3´- Pα ≤ 3.5 Å and 6.0 Å ≤ base pair distance ≤ 8.0 Å) is indicated with red dotted lines. Percentage of simulation frames with catalytically active conformation was shown as mean values ± standard deviation, which were calculated by bootstrapping of N independent production MD simulations (N=16).
Extended Data Fig. 4 UBP structure from DNA polymerase (dNaM-d5SICSTP, PDB 3SV3) shows co-planar edge-to-edge configuration.
UBP structure from DNA polymerase (dNaM-d5SICSTP, PDB 3SV3) shows co-planar edge-to-edge configuration.
Supplementary information
Supplementary Information
Supplementary Tables 1–3.
Supplementary Data 1
wwPDB validation report for 7KEE.
Supplementary Data 2
wwPDB validation report for 7KEF.
Supplementary Data 3
wwPDB validation report for 7KED.
Source data
Source Data Fig. 1
Unprocessed gels for Fig. 1c,e.
Rights and permissions
About this article
Cite this article
Oh, J., Shin, J., Unarta, I.C. et al. Transcriptional processing of an unnatural base pair by eukaryotic RNA polymerase II. Nat Chem Biol 17, 906–914 (2021). https://doi.org/10.1038/s41589-021-00817-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-021-00817-3
This article is cited by
-
A unified Watson-Crick geometry drives transcription of six-letter expanded DNA alphabets by E. coli RNA polymerase
Nature Communications (2023)
-
Structural basis of transcription recognition of a hydrophobic unnatural base pair by T7 RNA polymerase
Nature Communications (2023)