Abstract
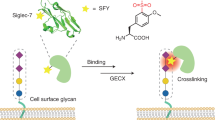
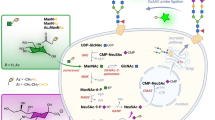
Cell surfaces are glycosylated in various ways with high heterogeneity, which usually leads to ambiguous conclusions about glycan-involved biological functions. Here, we describe a two-step chemoenzymatic approach for N-glycan-subtype-selective editing on the surface of living cells that consists of a first ‘delete’ step to remove heterogeneous N-glycoforms of a certain subclass and a second ‘insert’ step to assemble a well-defined N-glycan back onto the pretreated glyco-sites. Such glyco-edited cells, carrying more homogeneous oligosaccharide structures, could enable precise understanding of carbohydrate-mediated functions. In particular, N-glycan-subtype-selective remodeling and imaging with different monosaccharide motifs at the non-reducing end were successfully achieved. Using a combination of the expression system of the Lec4 CHO cell line and this two-step glycan-editing approach, opioid receptor delta 1 (OPRD1) was investigated to correlate its glycostructures with the biological functions of receptor dimerization, agonist-induced signaling and internalization.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All of the data are available in the paper and its Supplementary Information. Synthetic procedures and the characterization data of the new N-glycan substrates are also provided. Supporting data and processing protocols of cell imaging, FRET, FACS and western blot gels are available in the Supplementary files. The sequence data for the OPRD1 plasmid have been deposited to GenBank (accession code MN922300).
References
Varki, A. & Kornfeld, S. in Essentials of Glycobiology (eds. Cummings, R. D. et al.) 1–18 (Cold Spring Harbor, 2015).
Moremen, K. W., Tiemeyer, M. & Nairn, A. V. Vertebrate protein glycosylation: diversity, synthesis and function. Nat. Rev. Mol. Cell Biol. 13, 448–462 (2012).
Pinho, S. S. & Reis, C. A. Glycosylation in cancer: mechanisms and clinical implications. Nat. Rev. Cancer 15, 540–555 (2015).
Stanley, P. Chinese hamster ovary cell mutants with multiple glycosylation defects for production of glycoproteins with minimal carbohydrate heterogeneity. Mol. Cell Biol. 9, 377–383 (1989).
Weinstein, J. et al. A point mutation causes mistargeting of Golgi GlcNAc-TV in the Lec4A Chinese hamster ovary glycosylation mutant. J. Biol. Chem. 271, 27462–27469 (1996).
Chen, W. & Stanley, P. Five Lec1 CHO cell mutants have distinct Mgat1 gene mutations that encode truncated N-acetylglucosaminyltransferase I. Glycobiology 13, 43–50 (2003).
Yang, Z. et al. Engineered CHO cells for production of diverse, homogeneous glycoproteins. Nat. Biotechnol. 33, 842–844 (2015).
Yamane-Ohnuki, N. et al. Establishment of FUT8 knockout Chinese hamster ovary cells: an ideal host cell line for producing completely defucosylated antibodies with enhanced antibody-dependent cellular cytotoxicity. Biotechnol. Bioeng. 87, 614–622 (2004).
Li, H. et al. Optimization of humanized IgGs in glycoengineered Pichia pastoris. Nat. Biotechnol. 24, 210–215 (2006).
North, S. J. et al. Glycomics profiling of Chinese hamster ovary cell glycosylation mutants reveals N-glycans of a novel size and complexity. J. Biol. Chem. 285, 5759–5775 (2010).
Mahal, L. K., Yarema, K. J. & Bertozzi, C. R. Engineering chemical reactivity on cell surfaces through oligosaccharide biosynthesis. Science 276, 1125–1128 (1997).
Prescher, J. A., Dube, D. H. & Bertozzi, C. R. Chemical remodelling of cell surfaces in living animals. Nature 430, 873–877 (2004).
Vocadlo, D. J., Hang, H. C., Kim, E.-J., Hanover, J. A. & Bertozzi, C. R. A chemical approach for identifying O-GlcNAc-modified proteins in cells. Proc. Natl Acad. Sci. USA 100, 9116–9121 (2003).
Hang, H. C., Yu, C., Kato, D. L. & Bertozzi, C. R. A metabolic labeling approach toward proteomic analysis of mucin-type O-linked glycosylation. Proc. Natl Acad. Sci. USA 100, 14846–14851 (2003).
Kayser, H. et al. Biosynthesis of a nonphysiological sialic acid in different rat organs, using N-propanoyl-d-hexosamines as precursors. J. Biol. Chem. 267, 16934–16938 (1992).
Zeng, Y., Ramya, T. N. C., Dirksen, A., Dawson, P. E. & Paulson, J. C. High-efficiency labeling of sialylated glycoproteins on living cells. Nat. Methods 6, 207–209 (2009).
Hui, J. et al. Localized chemical remodeling for live cell imaging of protein-specific glycoform. Angew. Chem. Int. Ed. 56, 8139–8143 (2017).
Mbua, N. E. et al. Selective exo-enzymatic labeling of N-glycans on the surface of living cells by recombinant ST6Gal I. Angew. Chem. Int. Ed. 52, 13012–13015 (2013).
Sun, T. et al. One-step selective exoenzymatic labeling (SEEL) strategy for the biotinylation and identification of glycoproteins of living cells. J. Am. Chem. Soc. 138, 11575–11582 (2016).
Wen, L. et al. Two-step chemoenzymatic detection of N-acetylneuraminic acid-α(2-3)-galactose glycans. J. Am. Chem. Soc. 138, 11473–11476 (2016).
Wang, W. et al. Chemoenzymatic synthesis of GDP-l-fucose and the Lewis X glycan derivatives. Proc. Natl Acad. Sci. USA 106, 16096–16101 (2009).
Briard, J. G., Jiang, H., Moremen, K. W., Macauley, M. S. & Wu, P. Cell-based glycan arrays for probing glycan–glycan binding protein interactions. Nat. Commun. 9, 880 (2018).
Rabuka, D., Hubbard, S. C., Laughlin, S. T., Argade, S. P. & Bertozzi, C. R. A chemical reporter strategy to probe glycoprotein fucosylation. J. Am. Chem. Soc. 128, 12078–12079 (2006).
Zheng, T. et al. Tracking N-acetyllactosamine on cell surface glycans in vivo. Angew. Chem. Int. Ed. 50, 4113–4118 (2011).
Li, Q., Li, Z., Duan, X. & Yi, W. A tandem enzymatic approach for detecting and imaging tumor-associated Thomsen–Friedenreich antigen disaccharide. J. Am. Chem. Soc. 136, 12536–12539 (2014).
Wang, L. X. & Huang, W. Enzymatic transglycosylation for glycoconjugate synthesis. Curr. Opin. Chem. Biol. 13, 592–600 (2009).
Fairbanks, A. J. The ENGases: versatile biocatalysts for the production of homogeneous N-linked glycopeptides and glycoproteins. Chem. Soc. Rev. 46, 5128–5146 (2017).
Huang, W., Yang, Q., Umekawa, M., Yamamoto, K. & Wang, L. X. Arthrobacter endo-β-N-acetylglucosaminidase shows transglycosylation activity on complex-type N-glycan oxazolines: one-pot conversion of ribonuclease B to sialylated ribonuclease C. Chembiochem 11, 1350–1355 (2010).
Huang, W., Giddens, J., Fan, S. Q., Toonstra, C. & Wang, L. X. Chemoenzymatic glycoengineering of intact IgG antibodies for gain of functions. J. Am. Chem. Soc. 134, 12308–12318 (2012).
Giddens, J. P., Lomino, J. V., Amin, M. N. & Wang, L. X. Endo-F3 glycosynthase mutants enable chemoenzymatic synthesis of core-fucosylated triantennary complex type glycopeptides and glycoproteins. J. Biol. Chem. 291, 9356–9370 (2016).
Tang, F., Wang, L. X. & Huang, W. Chemoenzymatic synthesis of glycoengineered IgG antibodies and glycosite-specific antibody-drug conjugates. Nat. Protoc. 12, 1702–1721 (2017).
Tang, F. et al. One-pot N-glycosylation remodeling of IgG with non-natural sialylglycopeptides enables glycosite-specific and dual-payload antibody-drug conjugates. Org. Biomol. Chem. 14, 9501–9518 (2016).
Tang, Y. et al. Real-time analysis on drug–antibody ratio of antibody–drug conjugates for synthesis, process optimization and quality control. Sci. Rep. 7, 7763 (2017).
Yang, Q. et al. Glycan remodeling of human erythropoietin (EPO) through combined mammalian cell engineering and chemoenzymatic transglycosylation. ACS Chem. Biol. 12, 1665–1673 (2017).
Takegawa, K., Nakoshi, M., Iwahara, S., Yamamoto, K. & Tochikura, T. Induction and purification of endo-β-N-acetylglucosaminidase from Arthrobacter protophormiae grown in ovalbumin. Appl. Environ. Microbiol. 55, 3107–3112 (1989).
Yamamoto, K., Kadowaki, S., Fujisaki, M., Kumagai, H. & Tochikura, T. Novel specificities of Mucor hiemalis endo-β-N-acetylglucosaminidase acting complex asparagine-linked oligosaccharides. Biosci. Biotech. Biochem. 58, 72–77 (1994).
Tarentino, A. L. & Plummer, T. H. Enzymatic deglycosylation of asparagine-linked glycans: purification, properties, and specificity of oligosaccharide-cleaving enzymes from Flavobacterium meningosepticum.Methods Enzymol. 230, 44–57 (1994).
Muramatsu, H. et al. Molecular cloning and expression of endo-β-N-acetylglucosaminidase D, which acts on the core structure of complex type asparagine-linked oligosaccharides. J. Biochem. 129, 923–928 (2001).
Plummer, J. T. H., Phelan, A. W. & Tarentino, A. L. Porcine fibrinogen glycopeptides: substrates for detecting endo-β-N-acetylglucosaminidases F2 and F3. Anal. Biochem. 235, 98–101 (1996).
Lin, W., Du, Y., Zhu, Y. & Chen, X. A cis-membrane FRET-based method for protein-specific imaging of cell-surface glycans. J. Am. Chem. Soc. 136, 679–687 (2014).
Huang, W. et al. Glycosynthases enable a highly efficient chemoenzymatic synthesis of N-glycoproteins carrying intact natural N-glycans. J. Am. Chem. Soc. 131, 2214–2223 (2009).
Eshima, Y., Higuchi, Y., Kinoshita, T., Nakakita, S. & Takegawa, K. Transglycosylation activity of glycosynthase mutants of endo-β-N-acetylglucosaminidase from Coprinopsis cinerea. PLoS ONE 10, e0132859 (2015).
Murakami, S. et al. Identification and characterization of endo-β-N-acetylglucosaminidase from methylotrophic yeast Ogataea minuta. Glycobiology 23, 736–744 (2013).
Whittaker, J. W. Free radical catalysis by galactose oxidase. Chem. Rev. 103, 2347–2364 (2003).
Waddling, C. A., Plummer, T. H., Tarentino, A. L. & Van Roey, P. Structural basis for the substrate specificity of endo-β-N-acetylglucosaminidase F3. Biochemistry 39, 7878–7885 (2000).
Hamouda, H. et al. Rapid analysis of cell surface N-glycosylation from living cells using mass spectrometry. J. Proteome Res. 13, 6144–6151 (2014).
Burford, N. T. et al. Identification of selective agonists and positive allosteric modulators for micro- and delta-opioid receptors from a single high-throughput screen. J. Biomol. Screen. 19, 1255–1265 (2014).
Xie, R., Hong, S., Feng, L., Rong, J. & Chen, X. Cell-selective metabolic glycan labeling based on ligand-targeted liposomes. J. Am. Chem. Soc. 134, 9914–9917 (2012).
Xiao, H., Woods, E. C., Vukojicic, P. & Bertozzi, C. R. Precision glycocalyx editing as a strategy for cancer immunotherapy. Proc. Natl Acad. Sci. USA 113, 10304–10309 (2016).
Tong, X., Li, T., Li, C. & Wang, L. X. Generation and comparative kinetic analysis of new glycosynthase mutants from Streptococcus pyogenes endoglycosidases for antibody glycoengineering. Biochemistry 57, 5239–5246 (2018).
Umekawa, M. et al. Mutants of Mucor hiemalis endo-β-N-acetylglucosaminidase show enhanced transglycosylation and glycosynthase-like activities. J. Biol. Chem. 283, 4469–4479 (2008).
Fujita, K. et al. Synthesis of neoglycoenzymes with homogeneous N-linked oligosaccharides using immobilized endo-β-N-acetylglucosaminidase A. Biochem. Biophys. Res. Commun. 267, 134–138 (2000).
Sun, B. et al. A simplified procedure for gram-scale production of sialylglycopeptide (SGP) from egg yolks and subsequent semi-synthesis of Man3GlcNAc oxazoline. Carbohydr. Res. 396, 62–69 (2014).
Dell, A. et al. Mass spectrometry of carbohydrate-containing biopolymers. Methods Enzymol. 230, 108–132 (1994).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (no. 21877116, to W.H.), the National Science & Technology Major Project ‘Key New Drug Creation and Manufacturing Program’ of China (no. 2018ZX09711002-006, to W.H.) and the China Postdoctoral Science Foundation (no. BX20180339, to F.T., and no. 2018M642120, to F.T.). We thank R. Huang’s colleagues for their kind help with imaging studies. We also thank P. Wu of the Scripps Research Institute for providing us with Lec4 CHO cells.
Author information
Authors and Affiliations
Contributions
F.T. prepared the N-glycan substrates, performed cell-surface N-glycan editing by Endo-F3, conducted the N-glycan profiling by MALDI-TOF and carried out the OPRD1 internalization assay. M.Z. contributed to initial condition optimization and the proof of concept, performed N-glycan editing by Endo-M, constructed the OPRD1-expressing Lec4 CHO cell line and finished the cAMP assay. K.Q. provided the Endo-F3 and its mutant enzymes and performed Lec4 cell N-glycan editing. W.S. prepared the azido-complex-type glycan substrate (Az-CT-Ox). A.Y. and F.T. performed the FRET experiments. Y.Y. provided the Endo-A, Endo-M and its mutant enzymes, and PNGase F. L.Y. helped with western blots and FACS data acquisition and discussion. D.G., L.Z. and Y.T. helped with synthesis of chemical compounds and intermediates. L.Z. and H.Y. helped with OPRD1 expression. Y.C. and H.Z. assisted with MALDI-TOF determination. R.H. helped with imaging experiments. W.H. conceived the original idea and supervised the research. F.T., M.Z. and W.H. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2, Figs. 1–40 and Notes 1 and 2.
Supplementary Data
OPRD1 GenBank file.
Rights and permissions
About this article
Cite this article
Tang, F., Zhou, M., Qin, K. et al. Selective N-glycan editing on living cell surfaces to probe glycoconjugate function. Nat Chem Biol 16, 766–775 (2020). https://doi.org/10.1038/s41589-020-0551-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-0551-8
This article is cited by
-
Cut-insert-stitch editing reaction (CIStER) sequence for surgical chemical glycan editing
Communications Chemistry (2024)
-
Carbohydrate-active enzymes (CAZymes) in the gut microbiome
Nature Reviews Microbiology (2022)
-
Deciphering protein post-translational modifications using chemical biology tools
Nature Reviews Chemistry (2020)
-
Selective cell-surface N-glycan editing
Nature Methods (2020)