Abstract
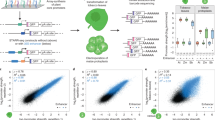
Agricultural biotechnology strategies often require the precise regulation of multiple genes to effectively modify complex plant traits. However, most efforts are hindered by a lack of characterized tools that allow for reliable and targeted expression of transgenes. We have successfully engineered a library of synthetic transcriptional regulators that modulate expression strength in planta. By leveraging orthogonal regulatory systems from Saccharomyces spp., we have developed a strategy for the design of synthetic activators, synthetic repressors, and synthetic promoters and have validated their use in Nicotiana benthamiana and Arabidopsis thaliana. This characterization of contributing genetic elements that dictate gene expression represents a foundation for the rational design of refined synthetic regulators. Our findings demonstrate that these tools provide variation in transcriptional output while enabling the concerted expression of multiple genes in a tissue-specific and environmentally responsive manner, providing a basis for generating complex genetic circuits that process endogenous and environmental stimuli.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The authors declare that the data supporting the findings of this study are available within the article and its Supplementary Information Files are available from the corresponding author upon request. All vectors and resources described are publicly available and can be found through the Inventory of Composable Elements at https://acs-registry.jbei.org/. Once logged in, the plasmids and strains are listed under the JBEI Public Registry tab.
References
Shih, P. M., Liang, Y. & Loqué, D. Biotechnology and synthetic biology approaches for metabolic engineering of bioenergy crops. Plant J. 87, 103–117 (2016).
Hawkins, K. M. & Smolke, C. D. Production of benzylisoquinoline alkaloids in Saccharomyces cerevisiae. Nat. Chem. Biol. 4, 564–573 (2008).
Potvin-Trottier, L., Lord, N. D., Vinnicombe, G. & Paulsson, J. Synchronous long-term oscillations in a synthetic gene circuit. Nature 538, 514–517 (2016).
Brückner, K. et al. A library of synthetic transcription activator-like effector-activated promoters for coordinated orthogonal gene expression in plants. Plant J. 82, 707–716 (2015).
Bashor, C. J. et al. Complex signal processing in synthetic gene circuits using cooperative regulatory assemblies. Science 364, 593–597 (2019).
Farzadfard, F., Perli, S. D. & Lu, T. K. Tunable and multifunctional eukaryotic transcription factors based on CRISPR/Cas. ACS Synth. Biol. 2, 604–613 (2013).
Lebar, T. & Jerala, R. Benchmarking of TALE- and CRISPR/dCas9-based transcriptional regulators in mammalian cells for the construction of synthetic genetic circuits. ACS Synth. Biol. 5, 1050–1058 (2016).
Gaber, R. et al. Designable DNA-binding domains enable construction of logic circuits in mammalian cells. Nat. Chem. Biol. 10, 203–208 (2014).
Khalil, A. S. et al. A synthetic biology framework for programming eukaryotic transcription functions. Cell 150, 647–658 (2012).
Haseloff, J. GFP variants for multispectral imaging of living cells. Methods Cell Biol. 58, 139–151 (1999).
Laplaze, L. et al. GAL4-GFP enhancer trap lines for genetic manipulation of lateral root development in Arabidopsis thaliana. J. Exp. Bot. 56, 2433–2442 (2005).
Gardner, M. J. et al. GAL4 GFP enhancer trap lines for analysis of stomatal guard cell development and gene expression. J. Exp. Bot. 60, 213–226 (2009).
Smale, S. T. & Kadonaga, J. T. The RNA polymerase II core promoter. Annu. Rev. Biochem. 72, 449–479 (2003).
Lubliner, S. et al. Core promoter sequence in yeast is a major determinant of expression level. Genome Res. 25, 1008–1017 (2015).
Pan, S., Czarnecka-Verner, E. & Gurley, W. B. Role of the TATA binding protein–transcription factor IIB interaction in supporting basal and activated transcription in plant cells. Plant Cell 12, 125–135 (2000).
Horikoshi, M. et al. Transcription factor TFIID induces DNA bending upon binding to the TATA element. Proc. Natl Acad. Sci. USA 89, 1060–1064 (1992).
Hampsey, M. Molecular genetics of the RNA polymerase II general transcriptional machinery. Microbiol. Mol. Biol. Rev. 62, 465–503 (1998).
Sadowski, I., Ma, J., Triezenberg, S. & Ptashne, M. GAL4-VP16 is an unusually potent transcriptional activator. Nature 335, 563–564 (1988).
Taylor, I. C., Workman, J. L., Schuetz, T. J. & Kingston, R. E. Facilitated binding of GAL4 and heat shock factor to nucleosomal templates: differential function of DNA-binding domains. Genes Dev. 5, 1285–1298 (1991).
Kiran, K. et al. The TATA-box sequence in the basal promoter contributes to determining light-dependent gene expression in plants. Plant Physiol. 142, 364–376 (2006).
Joshi, C. P. An inspection of the domain between putative TATA box and translation start site in 79 plant genes. Nucleic Acids Res 15, 6643–6653 (1987).
Vaillant, I., Schubert, I., Tourmente, S. & Mathieu, O. MOM1 mediates DNA‐methylation‐independent silencing of repetitive sequences in Arabidopsis. EMBO Rep. 7, 1273–1278 (2006).
Okamoto, H. & Hirochika, H. Silencing of transposable elements in plants. Trends Plant Sci. 6, 527–534 (2001).
Matzke, M. A., Mette, M. F. & Matzke, A. J. Transgene silencing by the host genome defense: implications for the evolution of epigenetic control mechanisms in plants and vertebrates. Plant Mol. Biol. 43, 401–415 (2000).
Kooter, J. M., Matzke, M. A. & Meyer, P. Listening to the silent genes: transgene silencing, gene regulation and pathogen control. Trends Plant Sci. 4, 340–347 (1999).
Mutalik, V. K. et al. Precise and reliable gene expression via standard transcription and translation initiation elements. Nat. Methods 10, 354–360 (2013).
Jensen, P. R. & Hammer, K. The sequence of spacers between the consensus sequences modulates the strength of prokaryotic promoters. Appl. Environ. Microbiol. 64, 82–87 (1998).
Levo, M. & Segal, E. In pursuit of design principles of regulatory sequences. Nat. Rev. Genet. 15, 453–468 (2014).
Harcum, S. W. & Bentley, W. E. Heat-shock and stringent responses have overlapping protease activity in Escherichia coli: implications for heterologous protein yield. Appl. Biochem. Biotechnol. 80, 23–38 (1999).
Denancé, N., Sánchez-Vallet, A., Goffner, D. & Molina, A. Disease resistance or growth: the role of plant hormones in balancing immune responses and fitness costs. Front. Plant Sci. 4, 155 (2013).
Tian, D., Traw, M. B., Chen, J. Q., Kreitman, M. & Bergelson, J. Fitness costs of R-gene-mediated resistance in Arabidopsis thaliana. Nature 423, 74–77 (2003).
Fordyce, P. M. et al. De novo identification and biophysical characterization of transcription-factor binding sites with microfluidic affinity analysis. Nat. Biotechnol. 28, 970–975 (2010).
Baker, C. R., Tuch, B. B. & Johnson, A. D. Extensive DNA-binding specificity divergence of a conserved transcription regulator. Proc. Natl Acad. Sci. USA 108, 7493–7498 (2011).
Baker, C. R., Booth, L. N., Sorrells, T. R. & Johnson, A. D. Protein modularity, cooperative binding, and hybrid regulatory states underlie transcriptional network diversification. Cell 151, 80–95 (2012).
Goff, S. A., Cone, K. C. & Fromm, M. E. Identification of functional domains in the maize transcriptional activator C1: comparison of wild-type and dominant inhibitor proteins. Genes Dev. 5, 298–309 (1991).
Dingwall, C. & Laskey, R. A. Nuclear targeting sequences—a consensus? Trends Biochem. Sci. 16, 478–481 (1991).
Li, T., Stark, M. R., Johnson, A. D. & Wolberger, C. Crystal structure of the MATa1/MATα2 homeodomain heterodimer bound to DNA. Science 270, 262–269 (1995).
Hiratsu, K., Matsui, K., Koyama, T. & Ohme-Takagi, M. Dominant repression of target genes by chimeric repressors that include the EAR motif, a repression domain, in Arabidopsis. Plant J. 34, 733–739 (2003).
Mead, J. et al. Interactions of the Mcm1 MADS box protein with cofactors that regulate mating in yeast. Mol. Cell. Biol. 22, 4607–4621 (2002).
Tan, S. & Richmond, T. J. Crystal structure of the yeast MATα2/MCM1/DNA ternary complex. Nature 391, 660–666 (1998).
Elble, R. & Tye, B. K. Both activation and repression of a-mating-type-specific genes in yeast require transcription factor Mcm1. Proc. Natl Acad. Sci. USA 88, 10966–10970 (1991).
Fagard, M. & Vaucheret, H. (Trans)gene silencing in plants: how many mechanisms? Annu. Rev. Plant Physiol. Plant Mol. Biol. 51, 167–184 (2000).
Morel, J. B., Mourrain, P., Béclin, C. & Vaucheret, H. DNA methylation and chromatin structure affect transcriptional and post-transcriptional transgene silencing in Arabidopsis. Curr. Biol. 10, 1591–1594 (2000).
Matzke, M. A., Primig, M., Trnovsky, J. & Matzke, A. J. Reversible methylation and inactivation of marker genes in sequentially transformed tobacco plants. EMBO J. 8, 643–649 (1989).
Sunilkumar, G., Mohr, L., Lopata-Finch, E., Emani, C. & Rathore, K. S. Developmental and tissue-specific expression of CaMV 35S promoter in cotton as revealed by GFP. Plant Mol. Biol. 50, 463–474 (2002).
Liu, W. et al. Computational discovery of soybean promoter cis-regulatory elements for the construction of soybean cyst nematode-inducible synthetic promoters. Plant Biotechnol. J. 12, 1015–1026 (2014).
Shih, P. M. et al. A robust gene-stacking method utilizing yeast assembly for plant synthetic biology. Nat. Commun. 7, 13215 (2016).
Patron, N. J. et al. Standards for plant synthetic biology: a common syntax for exchange of DNA parts. N. Phytol. 208, 13–19 (2015).
Ham, T. S. et al. Design, implementation and practice of JBEI-ICE: an open source biological part registry platform and tools. Nucleic Acids Res 40, e141 (2012).
Lohr, D., Venkov, P. & Zlatanova, J. Transcriptional regulation in the yeast GAL gene family: a complex genetic network. FASEB J. 9, 777–787 (1995).
Johnston, M. A model fungal gene regulatory mechanism: the GAL genes of Saccharomyces cerevisiae. Microbiological Rev. 51, 458–476 (1987).
Sparkes, I. A., Runions, J., Kearns, A. & Hawes, C. Rapid, transient expression of fluorescent fusion proteins in tobacco plants and generation of stably transformed plants. Nat. Protoc. 1, 2019–2025 (2006).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Jefferson, R. A. Assaying chimeric genes in plants: the GUS gene fusion system. Plant Mol. Biol. Report. 5, 387–405 (1987).
Liu, P.-P., Koizuka, N., Martin, R. C. & Nonogaki, H. The BME3 (Blue Micropylar End 3) GATA zinc finger transcription factor is a positive regulator of Arabidopsis seed germination. Plant J. 44, 960–971 (2005).
Acknowledgements
This work was part of the Department of Energy Early Career Award and the Department of Energy JBEI (http://www.jbei.org) supported by the US Department of Energy, Office of Science, Office of Biological and Environmental Research through contract DE-AC02-05CH11231 (P.M.S., D.L. and H.V.S.) between the Lawrence Berkeley National Laboratory and the US Department of Energy. This work was also funded by BASF, Rec ID 85335789 (P.M.S.). P.M.S. was supported by grant no. 1K99AT009573/R00AT009573 from the National Center for Complementary and Integrative Health at the National Institutes of Health. M.S.B. was supported by a National Science Foundation Graduate Research Fellowship, fellow ID 2018262076.
Author information
Authors and Affiliations
Contributions
P.M.S. and D.L. designed the project. M.S.B., K.M.V. and A.A.R. carried out transient expression experiments in N. benthamiana. K.M.V. and N.M. carried out stable Arabidopsis experiments. M.S.B. and A.Z. performed data analysis. M.S.B. and M.G.T. carried out qPCR experiments. M.S.B., A.Z. and P.M.S. wrote the paper. P.M.S., H.V.S. and D.L. edited the document and provided financial support and supervision.
Corresponding author
Ethics declarations
Competing interests
All authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs 1–21 and Tables 1–3.
Supplementary Dataset 1
Complete parts list with shorthand nomenclature and sequence data.
Supplementary Dataset 2
Expression strengths of all TF/promoter pairs characterized with the trans-element library.
Supplementary Dataset 3
Complete list of all cis-element sequences used in promoter design.
Supplementary Dataset 4
Minimal promoter library with sequences used.
Supplementary Dataset 5
Summary of all parts, constructs and sequences synthesized and assembled for the Gal4 system to investigate CCE and minimal promoters.
Rights and permissions
About this article
Cite this article
Belcher, M.S., Vuu, K.M., Zhou, A. et al. Design of orthogonal regulatory systems for modulating gene expression in plants. Nat Chem Biol 16, 857–865 (2020). https://doi.org/10.1038/s41589-020-0547-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-0547-4
This article is cited by
-
Edible mycelium bioengineered for enhanced nutritional value and sensory appeal using a modular synthetic biology toolkit
Nature Communications (2024)
-
Design principles for synthetic control systems to engineer plants
Plant Cell Reports (2023)
-
Orthogonal control of gene expression in plants using synthetic promoters and CRISPR-based transcription factors
Plant Methods (2022)
-
Metabolite trafficking enables membrane-impermeable-terpene secretion by yeast
Nature Communications (2022)
-
Plant synthetic epigenomic engineering for crop improvement
Science China Life Sciences (2022)