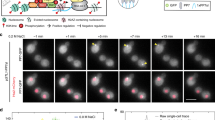
Abstract
Gene expression in mammalian cells results from coordinated protein-driven processes guided by diverse mechanisms of regulation, including protein–protein interactions, protein localization, DNA modifications and chromatin rearrangement. Regulation of gene expression is particularly important in stress-response pathways. To address the need to monitor chromosomal gene expression generating a readily detectable signal output that recapitulates gene expression dynamics, we developed a gene signal amplifier platform that links transcriptional and post-translational regulation of a fluorescent output to the expression of a chromosomal target gene. We generated a multiplex reporter system for monitoring markers of the unfolded protein response, a complex signal transduction pathway that remodels gene expression in response to proteotoxic stress in the endoplasmic reticulum. By recapitulating the transcriptional and translational control mechanisms underlying the expression of a target gene with high sensitivity, this platform provides a technology for monitoring gene expression with superior sensitivity and dynamic resolution.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The authors declare that data supporting the finding of this study are available within the article and its Supplementary Information. Additional data are available from the corresponding author upon reasonable request.
References
Hotamisligil, G. S. & Davis, R. J. Cell signaling and stress responses. Cold Spring Harb. Perspect. Biol. 8, a006072 (2016).
Hetz, C. & Saxena, S. ER stress and the unfolded protein response in neurodegeneration. Nat. Rev. Neurol. 13, 477–491 (2017).
Volpe, C. M. O., Villar-Delfino, P. H., Dos Anjos, P. M. F. & Nogueira-Machado, J. A. Cellular death, reactive oxygen species (ROS) and diabetic complications. Cell Death Dis. 9, 119 (2018).
Buono, R. & Longo, V. D. Starvation, stress resistance, and cancer. Trends Endocrinol. Metab. 29, 271–280 (2018).
Tahmasebi, S., Khoutorsky, A., Mathews, M. B. & Sonenberg, N. Translation deregulation in human disease. Nat. Rev. Mol. Cell Biol. 19, 791–807 (2018).
Butte, A. The use and analysis of microarray data. Nat. Rev. Drug Discov. 1, 951–960 (2002).
Nolan, T., Hands, R. E. & Bustin, S. A. Quantification of mRNA using real-time RT-PCR. Nat. Protoc. 1, 1559–1582 (2006).
Draghici, S., Khatri, P., Eklund, A. C. & Szallasi, Z. Reliability and reproducibility issues in DNA microarray measurements. Trends Genet. 22, 101–109 (2006).
Catarino, R. R. & Stark, A. Assessing sufficiency and necessity of enhancer activities for gene expression and the mechanisms of transcription activation. Genes Dev. 32, 202–223 (2018).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Hetz, C. The unfolded protein response: controlling cell fate decisions under ER stress and beyond. Nat. Rev. Mol. Cell Biol. 13, 89–102 (2012).
Shoulders, M. D. et al. Stress-independent activation of XBP1s and/or ATF6 reveals three functionally diverse ER proteostasis environments. Cell Rep. 3, 1279–1292 (2013).
Walter, F., Schmid, J., Düssmann, H., Concannon, C. G. & Prehn, J. H. M. Imaging of single cell responses to ER stress indicates that the relative dynamics of IRE1/XBP1 and PERK/ATF4 signalling rather than a switch between signalling branches determine cell survival. Cell Death Differ. 22, 1502–1516 (2015).
Walter, P. & Ron, D. The unfolded protein response: from stress pathway to homeostatic regulation. Science 334, 1081–1086 (2011).
Haze, K., Yoshida, H., Yanagi, H., Yura, T. & Mori, K. Mammalian transcription factor ATF6 is synthesized as a transmembrane protein and activated by proteolysis in response to endoplasmic reticulum stress. Mol. Biol. Cell 10, 3787–3799 (1999).
Klemm, S. L., Shipony, Z. & Greenleaf, W. J. Chromatin accessibility and the regulatory epigenome. Nat. Rev. Genet. 20, 207–220 (2019).
Soutourina, J. Transcription regulation by the Mediator complex. Nat. Rev. Mol. Cell Biol. 19, 262–274 (2018).
Schoenfelder, S. & Fraser, P. Long-range enhancer–promoter contacts in gene expression control. Nat. Rev. Genet. 20, 437–455 (2019).
Han, J. et al. ER-stress-induced transcriptional regulation increases protein synthesis leading to cell death. Nat. Cell Biol. 15, 481–490 (2013).
Adachi, Y. et al. ATF6 is a transcription factor specializing in the regulation of quality control proteins in the endoplasmic reticulum. Cell Struct. Funct. 33, 75–89 (2008).
Bochkov, Y. & Palmenberg, A. Translational efficiency of EMCV IRES in bicistronic vectors is dependent upon IRES sequence and gene location. Biotechniques 41, 283–292 (2006).
Mizuguchi, H., Xu, Z., Ishii-Watabe, A., Uchida, E. & Hayakawa, T. IRES-dependent second gene expression is significantly lower than cap-dependent first gene expression in a bicistronic vector. Mol. Ther. 1, 376–382 (2000).
Lee, A.-H., Iwakoshi, N. N. & Glimcher, L. H. XBP-1 regulates a subset of endoplasmic reticulum resident chaperone genes in the unfolded protein response. Mol. Cell. Biol. 23, 7448–7459 (2003).
Yoshida, H. et al. ATF6 activated by proteolysis binds in the presence of NF-Y (CBF) directly to the cis-acting element responsible for the mammalian unfolded protein response. Mol. Cell. Biol. 20, 6755–6767 (2000).
Vattem, K. M. & Wek, R. C. Reinitiation involving upstream ORFs regulates ATF4 mRNA translation in mammalian cells. Proc. Natl Acad. Sci. USA 101, 11269–11274 (2004).
Wang, Y. et al. Activation of ATF6 and an ATF6 DNA binding site by the endoplasmic reticulum stress response. J. Biol. Chem. 275, 27013–27020 (2000).
Dricu, A., Carlberg, M., Wang, M. & Larsson, O. Inhibition of N-linked glycosylation using tunicamycin causes cell death in malignant cells: role of down-regulation of the insulin-like growth factor 1 receptor in induction of apoptosis. Cancer Res. 57, 543–548 (1997).
Lytton, J., Westlin, M. & Hanley, M. R. Thapsigargin inhibits the sarcoplasmic or endoplasmic reticulum Ca-ATPase family of calcium pumps. J. Biol. Chem. 266, 17067–17071 (1991).
Gossen, M. & Bujard, H. Tight control of gene expression in mammalian cells by tetracycline-responsive promoters. Proc. Natl Acad. Sci. USA 89, 5547–5551 (1992).
Weber, W. et al. Macrolide-based transgene control in mammalian cells and mice. Nat. Biotechnol. 20, 901–907 (2002).
Zhao, W., Pferdehirt, L. & Segatori, L. Quantitatively predictable control of cellular protein levels through proteasomal degradation. ACS Synth. Biol. 7, 540–552 (2018).
Oslowski, C. M. & Urano, F. Measuring ER stress and the unfolded protein response using mammalian tissue culture system. Methods Enzymol. 490, 71–92 (2011).
Raser, J. M. & O’Shea, E. K. Noise in gene expression: origins, consequences, and control. Science 309, 2010–2013 (2005).
Adamson, B. et al. A multiplexed single-cell CRISPR screening platform enables systematic dissection of the unfolded protein response. Cell 167, 1867–1882.e21 (2016).
Rong, J. et al. Cell-based high-throughput luciferase reporter gene assays for identifying and profiling chemical modulators of endoplasmic reticulum signaling protein, IRE1. J. Biomol. Screen. 20, 1232–1245 (2015).
Lindholm, D., Wootz, H. & Korhonen, L. ER stress and neurodegenerative diseases. Cell Death Differ. 13, 385–392 (2006).
Venkatesan, A. & Dasgupta, A. Novel fluorescence-based screen to identify small synthetic internal ribosome entry site elements. Mol. Cell. Biol. 21, 2826–2837 (2001).
Fernandez-Rodriguez, J. & Voigt, C. A. Post-translational control of genetic circuits using Potyvirus proteases. Nucleic Acids Res. 44, 6493–6502 (2016).
Wang, B., Barahona, M. & Buck, M. Engineering modular and tunable genetic amplifiers for scaling transcriptional signals in cascaded gene networks. Nucleic Acids Res. 42, 9484–9492 (2014).
Wan, X. et al. Cascaded amplifying circuits enable ultrasensitive cellular sensors for toxic metals. Nat. Chem. Biol. 15, 540–548 (2019).
Hörner, M. & Weber, W. Molecular switches in animal cells. FEBS Lett. 586, 2084–2096 (2012).
Younger, A. K. D. et al. Development of novel metabolite-responsive transcription factors via transposon-mediated protein fusion. Protein Eng. Des. Sel. 31, 55–63 (2018).
Muyldermans, S. Nanobodies: natural single-domain antibodies. Annu. Rev. Biochem. 82, 775–797 (2013).
Virant, D. et al. A peptide tag-specific nanobody enables high-quality labeling for dSTORM imaging. Nat. Commun. 9, 1–14 (2018).
Jayanthi, B. E. K., Zhao, W. & Segatori, L. Input-dependent post-translational control of the reporter output enhances dynamic resolution of mammalian signaling systems. Methods Enzymol. 622, 1–27 (2019).
Qin, J. Y. et al. Systematic comparison of constitutive promoters and the doxycycline-inducible promoter. PLoS ONE 5, 3–6 (2010).
Panganiban, R. A. et al. Genome-wide CRISPR screen identifies suppressors of endoplasmic reticulum stress-induced apoptosis. Proc. Natl Acad. Sci. USA 116, 13384–13393 (2019).
Potting, C. et al. Genome-wide CRISPR screen for PARKIN regulators reveals transcriptional repression as a determinant of mitophagy. Proc. Natl Acad. Sci. USA 115, E180–E189 (2017).
Pusapati, G. V. et al. CRISPR screens uncover genes that regulate target cell sensitivity to the morphogen Sonic Hedgehog. Dev. Cell 44, 113–129.e8 (2018).
Donati, G., Imbriano, C. & Mantovani, R. Dynamic recruitment of transcription factors and epigenetic changes on the ER stress response gene promoters. Nucleic Acids Res. 34, 3116–3127 (2006).
Zhao, W., Bonem, M., McWhite, C., Silberg, J. J. & Segatori, L. Sensitive detection of proteasomal activation using the Deg-On mammalian synthetic gene circuit. Nat. Commun. 5, 3612 (2014).
Kramer, B. P. et al. An engineered epigenetic transgene switch in mammalian cells. Nat. Biotechnol. 22, 867–870 (2004).
Geraerts, M., Willems, S., Baekelandt, V., Debyser, Z. & Gijsbers, R. Comparison of lentiviral vector titration methods. BMC Biotechnol. 6, 34 (2006).
Acknowledgements
This work was funded by the National Science Foundation (grant no. MCB-1615562, grant no. CBET-1805317 and grant no. CBET-1930149) and the Welch Foundation (grant no. C-1824), and was conducted in part using resources of the Shared Equipment Authority at Rice University. This project was supported by the Cytometry and Cell Sorting Core at Baylor College of Medicine, with funding from the CPRIT Core Facility Support Award (grant no. CPRIT-RP180672), and by the NIH (grant no. P30 CA125123 and grant no. S10 RR024574).
Author information
Authors and Affiliations
Contributions
C.A.O. and L.S. conceived the projected. C.A.O., B.B. and S.D.P. performed the experiments and C.A.O. and B.B. analyzed the data. C.A.O. and A.L.Y. generated the mathematical model. C.A.O. and L.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–3, Figs. 1–18 and Note.
Rights and permissions
About this article
Cite this article
Origel Marmolejo, C.A., Bachhav, B., Patibandla, S.D. et al. A gene signal amplifier platform for monitoring the unfolded protein response. Nat Chem Biol 16, 520–528 (2020). https://doi.org/10.1038/s41589-020-0497-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-0497-x
This article is cited by
-
Customizable gene sensing and response without altering endogenous coding sequences
Nature Chemical Biology (2024)
-
Engineering a synthetic gene circuit for high-performance inducible expression in mammalian systems
Nature Communications (2024)