Abstract
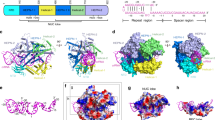
Cas12g, the type V–G CRISPR–Cas effector, is an RNA-guided ribonuclease that targets single-stranded RNA substrate. The CRISPR–Cas12g system offers a potential platform for transcriptome engineering and diagnostic applications. We determined the structures of Cas12g–guide RNA complexes in the absence and presence of target RNA by cryo-EM to a resolution of 3.1 Å and 4.8 Å, respectively. Cas12g adopts a bilobed structure with miniature REC2 and Nuc domains, whereas the guide RNAs fold into a flipped ‘F’ shape, which is primarily recognized by the REC lobe. Target RNA and the CRISPR RNA (crRNA) guide form a duplex that inserts into the central cavity between the REC and NUC lobes, inducing conformational changes in both lobes to activate Cas12g. The structural insights would facilitate the development of Cas12g-based applications.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Cryo-EM reconstructions of Cas12g–sgRNA and Cas12g–sgRNA-target RNA complexes have been deposited in the Electron Microscopy Data Bank under accession numbers EMD-22257 and EMD-22258, respectively. Coordinates for atomic models of Cas12g–sgRNA and Cas12g–sgRNA-target RNA complexes have been deposited in the Protein Data Bank under accession numbers 6XMF and 6XMG, respectively. Source data are provided with this paper.
References
Sorek, R., Lawrence, C. M. & Wiedenheft, B. CRISPR-mediated adaptive immune systems in bacteria and archaea. Annu. Rev. Biochem. 82, 237–266 (2013).
Marraffini, L. A. CRISPR–Cas immunity in prokaryotes. Nature 526, 55–61 (2015).
Jiang, F. & Doudna, J. A. CRISPR–Cas9 structures and mechanisms. Annu. Rev. Biophys. 46, 505–529 (2017).
Hille, F. et al. The biology of CRISPR–Cas: backward and forward. Cell 172, 1239–1259 (2018).
Shmakov, S. et al. Diversity and evolution of class 2 CRISPR–Cas systems. Nat. Rev. Microbiol. 15, 169–182 (2017).
Makarova, K. S. et al. An updated evolutionary classification of CRISPR–Cas systems. Nat. Rev. Microbiol. 13, 722–736 (2015).
Koonin, E. V., Makarova, K. S. & Zhang, F. Diversity, classification and evolution of CRISPR–Cas systems. Curr. Opin. Microbiol. 37, 67–78 (2017).
Barrangou, R. & Doudna, J. A. Applications of CRISPR technologies in research and beyond. Nat. Biotechnol. 34, 933–941 (2016).
Komor, A. C., Badran, A. H. & Liu, D. R. CRISPR-based technologies for the manipulation of eukaryotic genomes. Cell 168, 20–36 (2017).
Cox, D. B. T. et al. RNA editing with CRISPR–Cas13. Science 358, 1019–1027 (2017).
Abudayyeh, O. O. et al. RNA targeting with CRISPR–Cas13. Nature 550, 280–284 (2017).
Terns, M. P. CRISPR-based technologies: impact of RNA-targeting systems. Mol. Cell 72, 404–412 (2018).
Dugar, G. et al. CRISPR RNA-dependent binding and cleavage of endogenous RNAs by the Campylobacter jejuni Cas9. Mol. Cell 69, 893–905 (2018).
Strutt, S. C., Torrez, R. M., Kaya, E., Negrete, O. A. & Doudna, J. A. RNA-dependent RNA targeting by CRISPR–Cas9. eLife 7, e32724 (2018).
Rousseau, B. A., Hou, Z., Gramelspacher, M. J. & Zhang, Y. Programmable RNA cleavage and recognition by a natural CRISPR–Cas9 system from Neisseria meningitidis. Mol. Cell 69, 906–914 (2018).
O’Connell, M. R. et al. Programmable RNA recognition and cleavage by CRISPR/Cas9. Nature 516, 263–266 (2014).
Yan, W. X. et al. Cas13d is a compact RNA-targeting type VI CRISPR effector positively modulated by a WYL-domain-containing accessory protein. Mol. Cell 70, 327–339 (2018).
Smargon, A. A. et al. Cas13b is a type VI-B CRISPR-associated RNA-guided RNase differentially regulated by accessory proteins Csx27 and Csx28. Mol. Cell 65, 618–630 (2017).
Abudayyeh, O. O. et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353, aaf5573 (2016).
Yan, W. X. et al. Functionally diverse type V CRISPR–Cas systems. Science 363, 88–91 (2019).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR–Cas system. Cell 163, 759–771 (2015).
Strecker, J. et al. Engineering of CRISPR–Cas12b for human genome editing. Nat. Commun. 10, 212 (2019).
Teng, F. et al. Repurposing CRISPR–Cas12b for mammalian genome engineering. Cell Discov. 4, 63 (2018).
Liu, J. J. et al. CasX enzymes comprise a distinct family of RNA-guided genome editors. Nature 566, 218–223 (2019).
Chen, J. S. et al. CRISPR–Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science 360, 436–439 (2018).
Li, S.-Y. et al. CRISPR–Cas12a has both cis- and trans-cleavage activities on single-stranded DNA. Cell Res. 28, 491–493 (2018).
Gootenberg, J. S. et al. Multiplexed and portable nucleic acid detection platform with Cas13, Cas12a, and Csm6. Science 360, 439–444 (2018).
Swarts, D. C. & Jinek, M. Mechanistic insights into the cis- and trans-acting DNase activities of Cas12a. Mol. Cell 73, 589–600 (2019).
O’Connell, M. R. Molecular mechanisms of RNA targeting by Cas13-containing type VI CRISPR–Cas systems. J. Mol. Biol. 431, 66–87 (2019).
Konermann, S. et al. Transcriptome engineering with RNA-targeting type VI-D CRISPR effectors. Cell 173, 665–676 (2018).
Dong, D. et al. The crystal structure of Cpf1 in complex with CRISPR RNA. Nature 532, 522–526 (2016).
Gao, P., Yang, H., Rajashankar, K. R., Huang, Z. & Patel, D. J. Type V CRISPR–Cas Cpf1 endonuclease employs a unique mechanism for crRNA-mediated target DNA recognition. Cell Res. 26, 901–913 (2016).
Yamano, T. et al. Crystal structure of Cpf1 in complex with guide RNA and target DNA. Cell 165, 949–962 (2016).
Yamano, T. et al. Structural basis for the canonical and non-canonical PAM recognition by CRISPR–Cpf1. Mol. Cell 67, 633–645 (2017).
Stella, S., Alcón, P. & Montoya, G. Structure of the Cpf1 endonuclease R-loop complex after target DNA cleavage. Nature 546, 559–563 (2017).
Stella, S. et al. Conformational activation promotes CRISPR–Cas12a catalysis and resetting of the endonuclease activity. Cell 175, 1856–1871 (2018).
Swarts, D. C., van der Oost, J. & Jinek, M. Structural basis for guide RNA processing and seed-dependent DNA targeting by CRISPR–Cas12a. Mol. Cell 66, 221–233 (2017).
Nishimasu, H. et al. Structural basis for the altered PAM recognition by engineered CRISPR–Cpf1. Mol. Cell 67, 139–147 (2017).
Zhang, H. et al. Structural basis for the Inhibition of CRISPR–Cas12a by anti-CRISPR proteins. Cell Host Microbe 25, 815–826 (2019).
Yang, H., Gao, P., Rajashankar, K. R. & Patel, D. J. PAM-dependent target DNA recognition and cleavage by C2c1 CRISPR–Cas endonuclease. Cell 167, 1814–1828 (2016).
Liu, L. et al. C2c1-sgRNA complex structure reveals RNA-guided DNA cleavage mechanism. Mol. Cell 65, 310–322 (2017).
Wu, D., Guan, X., Zhu, Y., Ren, K. & Huang, Z. Structural basis of stringent PAM recognition by CRISPR–C2c1 in complex with sgRNA. Cell Res. 27, 705–708 (2017).
Zhang, H., Li, Z., Xiao, R. & Chang, L. Mechanisms for target recognition and cleavage by the Cas12i RNA-guided endonuclease. Nat. Struct. Mol. Biol. 27, 1069–1076 (2020).
Liu, L. et al. The molecular architecture for RNA-guided RNA cleavage by Cas13a. Cell 170, 714–726 (2017).
Swarts, D. C. Stirring up the type V alphabet soup. CRISPR J. 2, 14–16 (2019).
Strecker, J. et al. RNA-guided DNA insertion with CRISPR-associated transposases. Science 365, 48–53 (2019).
Jeon, Y. et al. Direct observation of DNA target searching and cleavage by CRISPR–Cas12a. Nat. Commun. 9, 2777 (2018).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Lander, G. C. et al. Appion: an integrated, database-driven pipeline to facilitate EM image processing. J. Struct. Biol. 166, 95–102 (2009).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Buchan, D. W. A. & Jones, D. T. The PSIPRED protein analysis workbench: 20 years on. Nucleic Acids Res. 47, W402–W407 (2019).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D. Struct. Biol. 74, 531–544 (2018).
Pei, J., Kim, B. H. & Grishin, N. V. PROMALS3D: a tool for multiple protein sequence and structure alignments. Nucleic Acids Res. 36, 2295–2300 (2008).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Acknowledgements
We thank T. Klose and V. Bowman for help with cryo-EM, and S. Wilson for computation. This work made use of the Purdue Cryo-EM Facility. This work was supported by the NIH grant R01GM138675 and a Showalter Trust Research Award to L.C.
Author information
Authors and Affiliations
Contributions
L.C. supervised the study. H.Z., R.X. and Z.L. prepared samples. Z.L., H.Z. and L.C. collected and processed cryo-EM data. Z.L., H.Z., R.X. and R.H. performed biochemical analysis. Z.L., H.Z. and L.C. prepared figures. All authors analyzed the data. Z.L., H.Z. and L.C prepared the manuscript with input from R.X. and R.H.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Sample preparation and cryo-EM of the Cas12g-sgRNA binary complex.
a, Purification of Cas12g. Upper: Size exclusion chromatography (SEC) profile. UV absorbance curves at 280 nm and 260 nm are shown in blue and red, respectively. Lower: SDS-PAGE analysis. b, Circular dichroism (CD) spectra of wild-type and mutants of Cas12g used in this study. c, A representative raw cryo-EM micrograph of the Cas12g-sgRNA complex from a total of 2520 micrographs. d, Representative 2D class averages from a total of 100 images. e, Three major classes from 3D classification. f, 3D refinement for Class I with angular distribution of particles. g, Focused classification around the flexible region of the REC1 domain (REC1220–354). h, Cryo-EM map of class I with each domain of Cas12g and sgRNA colored coded. i, Plot of the global half-map FSC (solid red line), map-to-model FSC (solid orange line), and spread of directional resolution values (±1σ from mean, green dotted lines; the blue bars indicate a histogram of 100 such values evenly sampled over the 3D FSC). FSC plot for the reconstruction suggests an average resolution of 3.1 Å. j, Local resolution map for the reconstruction in f.
Extended Data Fig. 2 Detailed cryo-EM density map of the Cas12g-sgRNA complex with atomic model fitted in.
a, Fitting of the REC1 domain, with density of REC1220–354 from focused classification shown in grey mesh. A representative α-helix from the REC1 domain is shown in detail on the right. b, Secondary structure prediction of REC1220–354 using PSIPRED. c, Fitting of the WED domain. d, Fitting of the RuvC domain. e, Fitting of the REC2 domain. f, Fitting of the Nuc domain. g, Fitting of nucleic acids to the corresponding cryo-EM map, with details of a segment shown as indicated on the right. The atomic models are shown in stick with crRNA and tracrRNA regions colored in orange and yellow, respectively. h, Detailed interactions between scaffolding duplex 1 and Cas12g. Cryo-EM density map is shown in mesh.
Extended Data Fig. 3 Structural comparison between Cas12g and Cas12b.
a, Overall structures of the Cas12g-sgRNA and the Cas12b-sgRNA complexes. b-f, Structural comparison of each domain. The structures are aligned by secondary-structure matching (SSM) in COOT.
Extended Data Fig. 4 Collateral cleavage and stoichiometric titration assays for Cas12g.
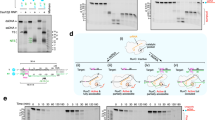
a, Collateral cleavage of unrelated ssDNA by Cas12g. The results shown are representative of three experiments. b, Collateral cleavage of unrelated ssRNA by Cas12g. The results shown are representative of three experiments. c, Stoichiometric titration assay indicates that Cas12g is a multiple turnover enzyme towards RNA substrate. Each point represents the cleavage ratio after 30 minutes at 37 °C. Error bars represent mean ± SD, where n = 3 replicates.
Extended Data Fig. 5 Comparison of the REC1 domain in Cas12g and Cas12b, and crRNA guide mismatch assay for Cas12g.
a, The tracrRNA scaffolding duplex 1 is bound to the concave surface of the REC1 domain in the Cas12g-sgRNA binary complex. b, The crRNA-target DNA duplex is bound to the concave surface of the REC1 domain in the Cas12b-sgRNA-DNA complex. c, The tracrRNA scaffolding duplex 1 is bound to the concave surface of the REC1 domain in the Cas12g-sgRNA-target RNA complex. d, RNA cleavage assay using wild-type and mutant target RNAs. The results shown are representative of three experiments.
Extended Data Fig. 6 Cryo-EM data processing for the Cas12g-sgRNA-target RNA complex.
a, A representative raw cryo-EM micrograph of the Cas12g-sgRNA-target RNA complex from a total of 987 micrographs. b, Representative 2D class averages from a total of 100 images. c, 3D classification. Three major classes were observed. Classes II and III did not result in good reconstructions and likely represent damaged particles. d, 3D refinement for Class I. Angular distribution is shown on the right. e, Plot of the global half-map FSC (solid red line), map-to-model FSC (solid orange line), and spread of directional resolution values (±1σ from mean, green dotted lines; the blue bars indicate a histogram of 100 such values evenly sampled over the 3D FSC). FSC plot for the reconstruction suggests an average resolution of 4.8 Å. f, Local resolution map in two views.
Extended Data Fig. 7 Structural and biochemical analysis of the Cas12g-sgRNA-target RNA complex.
a-b, Structural elements involved in crRNA-target RNA duplex recognition. Three such structural elements are indicated by red lines, including 1) an N-terminal extension enriched in arginines within the WED domain, 2) a loop within the REC2 domain (608-AKKATLQP-615), and 3) REC1220–354. c, Substrate RNA cleavage assay using wild-type Cas12g and Cas12g with mutations in two loops involved in the recognition of the crRNA-target RNA duplex. The results shown are representative of three experiments. d-e, Comparison of the REC1 and REC2 domains for substrate recognition between Cas12g and Cas12b.
Extended Data Fig. 8 Binding assay between the Cas12g-sgRNA complex and target nucleic acids.
Upper: SEC profiles for the Cas12g-sgRNA complex incubated with different targets including target RNA, target RNA with mismatches at positions 16 and 17, target RNA with mismatches at positions 18 and 19, and target ssDNA. UV absorbance curves at 280 nm are shown. Lower: Urea-PAGE (15%) analysis of the elution fractions from SEC as indicated. The results shown are representative of three experiments.
Extended Data Fig. 9 Cryo-EM analysis for the Cas12g-sgRNA-target DNA complex.
a, A representative raw cryo-EM micrograph of the Cas12g-sgRNA-target DNA complex from a total of 1083 micrographs. b, Representative 2D class averages from a total of 100 images. c, 3D classification. Classes 2-6 did not result in good reconstructions and likely represent damaged particles. d, 3D refinement for Class 1 with angular distribution. e, Plot of the global half-map FSC (solid red line), and spread of directional resolution values (±1σ from mean, green dotted lines; the blue bars indicate a histogram of 100 such values evenly sampled over the 3D FSC). FSC plot for the reconstruction suggests an average resolution of 5.8 Å. f, Local resolution map. g, Comparison of the rigid-body fitting between the Cas12g-sgRNA complex and the Cas12g-sgRNA-target RNA complex. Red squares indicate the fitting of the REC1 domain, which shows that the model of the Cas12g-sgRNA complex fits better to the map (left panel). h, Schematic model for Cas12g activation. Target DNA fails to form fully assembled duplex required for Cas12g activation.
Supplementary information
Supplementary Information
Supplementary Fig. 1 and Tables 1 and 2.
Source data
Source Data Fig. 2
Unprocessed gel.
Source Data Fig. 3
Unprocessed gel.
Source Data Fig. 4
Unprocessed gel.
Source Data Fig. 5e,f
Unprocessed gel.
Source Data Extended Data Fig. 1
Unprocessed gel.
Source Data Extended Data Fig. 4a,b
Unprocessed gel.
Source Data Extended Data Fig. 5
Unprocessed gel.
Source Data Extended Data Fig. 7
Unprocessed gel.
Source Data Extended Data Fig. 8
Unprocessed gel.
Rights and permissions
About this article
Cite this article
Li, Z., Zhang, H., Xiao, R. et al. Cryo-EM structure of the RNA-guided ribonuclease Cas12g. Nat Chem Biol 17, 387–393 (2021). https://doi.org/10.1038/s41589-020-00721-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-00721-2
This article is cited by
-
Targeting miRNA by CRISPR/Cas in cancer: advantages and challenges
Military Medical Research (2023)
-
Cas12a2 elicits abortive infection through RNA-triggered destruction of dsDNA
Nature (2023)
-
Programmable RNA detection with CRISPR-Cas12a
Nature Communications (2023)
-
Research progress on nucleic acid detection and genome editing of CRISPR/Cas12 system
Molecular Biology Reports (2023)
-
Structural biology of CRISPR–Cas immunity and genome editing enzymes
Nature Reviews Microbiology (2022)