Abstract
G protein-coupled receptors (GPCRs) relay information across cell membranes through conformational coupling between the ligand-binding domain and cytoplasmic signaling domain. In dimeric class C GPCRs, the mechanism of this process, which involves propagation of local ligand-induced conformational changes over 12 nm through three distinct structural domains, is unknown. Here, we used single-molecule FRET and live-cell imaging and found that metabotropic glutamate receptor 2 (mGluR2) interconverts between four conformational states, two of which were previously unknown, and activation proceeds through the conformational selection mechanism. Furthermore, the conformation of the ligand-binding domains and downstream domains are weakly coupled. We show that the intermediate states act as conformational checkpoints for activation and control allosteric modulation of signaling. Our results demonstrate a mechanism for activation of mGluRs where ligand binding controls the proximity of signaling domains, analogous to some receptor kinases. This design principle may be generalizable to other biological allosteric sensors.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The materials and data reported in this study are available from the corresponding author upon reasonable request. The PDB accession codes for the inactive and active structures of human mGluR5 are 6N52 and 6N51. Source data are provided with this paper.
Code availability
The custom codes for data analysis are available from the corresponding author upon reasonable request.
References
Smock, R. G. & Gierasch, L. M. Sending signals dynamically. Science 324, 198–203 (2009).
Changeux, J. P. & Christopoulos, A. Allosteric modulation as a unifying mechanism for receptor function and regulation. Cell 166, 1084–1102 (2016).
Thal, D. M., Glukhova, A., Sexton, P. M. & Christopoulos, A. Structural insights into G-protein-coupled receptor allostery. Nature 559, 45–53 (2018).
Dorsam, R. T. & Gutkind, J. S. G-protein-coupled receptors and cancer. Nat. Rev. Cancer 7, 79–94 (2007).
Zarzycka, B., Zaidi, S. A., Roth, B. L. & Katritch, V. Harnessing ion-binding sites for GPCR pharmacology. Pharmacol. Rev. 71, 571–595 (2019).
Hilger, D., Masureel, M. & Kobilka, B. K. Structure and dynamics of GPCR signaling complexes. Nat. Struct. Mol. Biol. 25, 4–12 (2018).
Latorraca, N. R., Venkatakrishnan, A. J. & Dror, R. O. GPCR dynamics: structures in motion. Chem. Rev. 117, 139–155 (2017).
Ye, L., Van Eps, N., Zimmer, M., Ernst, O. P. & Prosser, R. S. Activation of the A2A adenosine G-protein-coupled receptor by conformational selection. Nature 533, 265–268 (2016).
Nygaard, R. et al. The dynamic process of β2-adrenergic receptor activation. Cell 152, 532–542 (2013).
Vafabakhsh, R., Levitz, J. & Isacoff, E. Y. Conformational dynamics of a class C G-protein-coupled receptor. Nature 524, 497–501 (2015).
Wingler, L. M. et al. Angiotensin analogs with divergent bias stabilize distinct receptor conformations. Cell 176, 468–478 (2019).
Gusach, A. et al. Beyond structure: emerging approaches to study GPCR dynamics. Curr. Opin. Struct. Biol. 63, 18–25 (2020).
Gregorio, G. G. et al. Single-molecule analysis of ligand efficacy in β2AR–G-protein activation. Nature 547, 68–73 (2017).
Suomivuori, C. M. et al. Molecular mechanism of biased signaling in a prototypical G protein-coupled receptor. Science 367, 881–887 (2020).
Niswender, C. M. & Conn, P. J. Metabotropic glutamate receptors: physiology, pharmacology, and disease. Annu. Rev. Pharmacol. Toxicol. 50, 295–322 (2010).
Pin, J. P. & Bettler, B. Organization and functions of mGlu and GABAB receptor complexes. Nature 540, 60–68 (2016).
Koehl, A. et al. Structural insights into the activation of metabotropic glutamate receptors. Nature 567, 79–84 (2019).
Kunishima, N. et al. Structural basis of glutamate recognition by a dimeric metabotropic glutamate receptor. Nature 407, 971–977 (2000).
Wu, H. et al. Structure of a class C GPCR metabotropic glutamate receptor 1 bound to an allosteric modulator. Science 344, 58–64 (2014).
Doumazane, E. et al. Illuminating the activation mechanisms and allosteric properties of metabotropic glutamate receptors. Proc. Natl Acad. Sci. USA 110, E1416–E1425 (2013).
Hlavackova, V. et al. Sequential inter- and intrasubunit rearrangements during activation of dimeric metabotropic glutamate receptor 1. Sci. Signal. 5, ra59 (2012).
Roy, R., Hohng, S. & Ha, T. A practical guide to single-molecule FRET. Nat. Methods 5, 507–516 (2008).
Noren, C. J., Anthony-Cahill, S. J., Griffith, M. C. & Schultz, P. G. A general-method for site-specific incorporation of unnatural amino-acids into proteins. Science 244, 182–188 (1989).
Serfling, R. & Coin, I. Incorporation of unnatural amino acids into proteins expressed in mammalian cells. Methods Enzymol. 580, 89–107 (2016).
Huber, T., Naganathan, S., Tian, H., Ye, S. X. & Sakmar, T. P. Unnatural amino acid mutagenesis of GPCRs using amber codon suppression and bioorthogonal labeling. Method Enzymol. 520, 281–305 (2013).
Presolski, S. I., Hong, V. P. & Finn, M. G. Copper-catalyzed azide-alkyne click chemistry for bioconjugation. Curr. Protoc. Chem. Biol. 3, 153–162 (2011).
Muto, T., Tsuchiya, D., Morikawa, K. & Jingami, H. Structures of the extracellular regions of the group II/III metabotropic glutamate receptors. Proc. Natl. Acad Sci. USA 104, 3759–3764 (2007).
Huang, S. L. et al. Interdomain movements in metabotropic glutamate receptor activation. Proc. Natl Acad. Sci. USA 108, 15480–15485 (2011).
Jain, A. et al. Probing cellular protein complexes using single-molecule pull-down. Nature 473, 484–488 (2011).
Bronson, J. E., Fei, J., Hofman, J. M., Gonzalez, R. L. Jr. & Wiggins, C. H. Learning rates and states from biophysical time series: a Bayesian approach to model selection and single-molecule FRET data. Biophys. J. 97, 3196–3205 (2009).
Zhang, J. et al. Specific structural elements of the T-box riboswitch drive the two-step binding of the tRNA ligand. Elife 7, e39518 (2018).
Nussinov, R., Ma, B. & Tsai, C. J. Multiple conformational selection and induced fit events take place in allosteric propagation. Biophys. Chem. 186, 22–30 (2014).
Manglik, A. et al. Structural insights into the dynamic process of β2-adrenergic receptor signaling. Cell 161, 1101–1111 (2015).
Dror, R. O. et al. Activation mechanism of the β2-adrenergic receptor. Proc. Natl. Acad. Sci. USA 108, 18684–18689 (2011).
Grushevskyi, E. O. et al. Stepwise activation of a class C GPCR begins with millisecond dimer rearrangement. Proc. Natl Acad. Sci. USA 116, 10150–10155 (2019).
Kniazeff, J. et al. Closed state of both binding domains of homodimeric mGlu receptors is required for full activity. Nat. Struct. Mol. Biol. 11, 706–713 (2004).
Xue, L. et al. Major ligand-induced rearrangement of the heptahelical domain interface in a GPCR dimer. Nat. Chem. Biol. 11, 134–140 (2015).
Yanagawa, M., Yamashita, T. & Shichida, Y. Activation switch in the transmembrane domain of metabotropic glutamate receptor. Mol. Pharmacol. 76, 201–207 (2009).
Shaye, H. et al. Structural basis of the activation of a metabotropic GABA receptor. Nature 584, 298–303 (2020).
Foster, D. J. & Conn, P. J. Allosteric modulation of GPCRs: new insights and potential utility for treatment of schizophrenia and other CNS disorders. Neuron 94, 431–446 (2017).
Mijares, A., Lebesgue, D., Wallukat, G. & Hoebeke, J. From agonist to antagonist: Fab fragments of an agonist-like monoclonal anti-β2-adrenoceptor antibody behave as antagonists. Mol. Pharmacol. 58, 373–379 (2000).
Stoneman, M. R. et al. A general method to quantify ligand-driven oligomerization from fluorescence-based images. Nat. Methods 16, 493–496 (2019).
Hern, J. A. et al. Formation and dissociation of M1 muscarinic receptor dimers seen by total internal reflection fluorescence imaging of single molecules. Proc. Natl Acad. Sci. USA 107, 2693–2698 (2010).
Calebiro, D. et al. Single-molecule analysis of fluorescently labeled G-protein-coupled receptors reveals complexes with distinct dynamics and organization. Proc. Natl Acad. Sci. USA 110, 743–748 (2013).
Moller, J. et al. Single-molecule analysis reveals agonist-specific dimer formation of μ-opioid receptors. Nat. Chem. Biol. 16, 946–954 (2020).
Endres, N. F. et al. Conformational coupling across the plasma membrane in activation of the EGF receptor. Cell 152, 543–556 (2013).
Hong, V., Steinmetz, N. F., Manchester, M. & Finn, M. G. Labeling live cells by copper-catalyzed alkyne–azide click chemistry. Bioconjug. Chem. 21, 1912–1916 (2010).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Goodsell, D. S., Autin, L. & Olson, A. J. Illustrate: software for biomolecular illustration. Structure 27, 1716–1720 (2019).
Jain, A., Liu, R., Xiang, Y. K. & Ha, T. Single-molecule pull-down for studying protein interactions. Nat. Protoc. 7, 445–452 (2012).
Acknowledgements
We thank M.R. Schamber, D. Badong, D. May and A.Y. Pen for technical assistance, M. Gallio, J. Marko, A. Mondragon and K. Ragunathan for critical reading of the manuscript, and J. Fei (University of Chicago) for providing MATLAB scripts. This work was supported by the National Institutes of Health grant R01GM140272 (to R.V.) and by The Searle Leadership Fund for the Life Sciences at Northwestern University and by the Chicago Biomedical Consortium with support from the Searle Funds at The Chicago Community Trust (to R.V.). B.W.L. was supported by the National Institute of General Medical Sciences (NIGMS) Training Grant T32GM-008061. This work used resources of the Keck Biophysics Facility supported in part by the NCI CCSG P30 CA060553 grant awarded to the Robert H. Lurie Comprehensive Cancer Center of Northwestern University.
Author information
Authors and Affiliations
Contributions
B.W.L. performed plasmid construction and smFRET experiments. H.S.A. optimized and performed unnatural amino acid labeling and verification and live-cell FRET imaging and analysis. B.W.L. performed smFRET data analysis with assistance from H.S.A and R.V. The paper was written by B.W.L., H.S.A. and R.V.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Site-specific labeling of mGluR2 by click chemistry.
a, Schematic showing site-specific fluorescent labeling of mGluR2, with the unnatural amino acid 4-azido-L-phenylalanine at residue 548, by copper-catalyzed azide-alkyne click reaction. b, Donor (green: Cy3) and acceptor (red: Cy5) fluorophores conjugated to the CRD of inactive mGluR5 (PDB 6N52) at the position corresponding to residue 548 in mGluR2 (top). Representative confocal microscope image of HEK293T cells expressing 548UAA with the cell surface population labeled with donor (green: Cy3) and acceptor (red: Cy5) fluorophores through click chemistry (bottom). Scale bars, 10 µm. c, Unprocessed image of non-reducing 4-20% polyacrylamide gel electrophoresis of cell lysate from HEK293T cells expressing 548UAA and labeled by Cy5-alkyne. The gel is imaged with 633 nm excitation wavelength and 670-BP30 emission filter. Lane A: protein ladder; lane B: cell lysate; lane C: Cy5-alkyne dye. Results are representative of an individual experiment. d, Image of HEK293T cells expressing 548UAA and labeled with donor (green: Cy3) and acceptor (red: Cy5) fluorophores through click chemistry during live-cell FRET experiments. Scale bars, 10 µm. Results are representative of all titration and max response experiments for the 548UAA construct (N=21 independent experiments).
Extended Data Fig. 2 Live-cell ensemble FRET response to orthosteric ligands and a negative allosteric modulator.
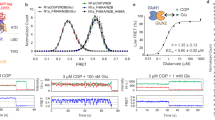
a, Representative donor and acceptor intensities and corresponding FRET signal from live-cell FRET titration experiments for DCG-IV, LY379268 and Glutamate +10 µM Ro64-5229 measured using HEK293T cells expressing 548UAA (top) and corresponding dose-response curves (bottom). Each titration curve is normalized to the 1 mM glutamate response. Data represent mean ± s.e.m. of N=20, 10 and 13 cells for LY379268, DCG-IV and glutamate +10 µM Ro64-5229, respectively, examined over 3 independent experiments. b, Donor and acceptor intensities and corresponding FRET signal in response to saturating concentrations of glutamate (1 mM), DCG-IV (100 µM) and LY379268 (20 µM) in HEK293T cells expressing 548UAA.
Extended Data Fig. 3 Example single-molecule time traces of CRD and VFT domain sensors.
a, Schematic of the single-molecule experiments (top). Representative frame from a single-molecule movie with the donor channel (Cy3) on the left and acceptor channel (Cy5) on the right (bottom). Molecules selected by analysis software for downstream processing are indicated by green circles. Scale bar, 3 µm. b, Example single-molecule time traces of the 548UAA in the absence of glutamate (0 µM) showing donor (green) and acceptor (red) intensities and corresponding FRET (blue). Dashed lines represent 4 distinct FRET states. c, Schematic of the VFT domain conformational sensor (left). Example single-molecule time traces of VFT domain sensor in the absence of glutamate (0 µM) and presence of saturating glutamate (1 mM) showing donor (green) and acceptor (red) intensities and corresponding FRET (blue) (bottom). smFRET population histogram of VFT domain mGluR2 sensor in the inactive (0 µM glutamate; 36 total molecules) and fully active (1 mM glutamate; 24 total molecules) conditions (top right). Data represent mean of N=2 independent experiments.
Extended Data Fig. 4 Example single-molecule time traces of CRD at different glutamate concentrations.
a, Example single-molecule time traces of the 548UAA at intermediate glutamate concentrations. b, Example single-molecule time traces of the 548UAA in saturating glutamate (1 mM). Donor (green) and acceptor (red) intensities and corresponding FRET (blue) are shown. Dashed lines represent 4 distinct FRET states.
Extended Data Fig. 5 mGluR3 undergoes a 4-state activation process.
a, smFRET population histograms of mGluR3 CRD sensor (labeled at residue 557) in the presence of competitive antagonist (LY341495; 221 total molecules) or saturating glutamate (290 total molecules). Data represent mean ± s.e.m. of N=3 independent experiments. Histograms are fitted (red) to 4 Gaussian distributions (blue) centered around 0.25 (state 1), 0.38 (state 2), 0.7 (state 3) and 0.87 (state 4), denoted with dashed lines. b, Example single-molecule time traces of mGluR3 CRD sensor in the presence of antagonist (LY341495) or saturating glutamate (1 mM) showing donor (green) and acceptor (red) intensities and corresponding FRET (blue). Dashed lines represent 4 distinct FRET states. Data was acquired at 100 ms time resolution.
Extended Data Fig. 6 Effect of intersubunit crosslinking on the CRD conformation.
Representative live-cell FRET measurement using HEK293T cells expressing 548UAA with the crosslinking mutation L521C upon application of intermediate (4 µM) and saturating glutamate (1 mM). FRET signal is normalized using initial FRET at time = 0.
Extended Data Fig. 7 smFRET analysis of CRD in the presence of orthosteric agonists.
a, Example single-molecule time traces of 548UAA at intermediate concentrations of DCG-IV (100 nM, top) and LY379268 (2 nM, bottom) showing donor (green) and acceptor (red) intensities and corresponding FRET (blue). Dashed lines represent 4 distinct FRET states. b, smFRET population histograms for 548UAA in the presence of saturating glutamate (1 mM; 152 total molecules), DCG-IV (100 µM; 470 total molecules) and LY379268 (20 µM; 356 total molecules). Data represent mean ± s.e.m. of N=3 independent experiments.
Extended Data Fig. 8 CRDs of mGluR2 are dynamic.
Heatmap illustrating the FRET values sampled by individual 548UAA receptors in the inactive (0 µM glutamate, top) and fully active (1 mM glutamate, bottom) conditions. Each row is the smFRET time trace of a single molecule over 6 seconds. The smFRET traces were smoothed using a 3-point moving average filter. 100 independent molecules are shown for each condition.
Extended Data Fig. 9 Analysis of conformational state 2 and state 4.
Example single-molecule time traces of 548UAA YADA/WT heterodimers at 20 µM glutamate. ATTO488 (light blue), donor (green) and acceptor (red) intensities and the corresponding FRET (blue) are shown. Dashed lines represent 4 distinct FRET states. Data was acquired at 100 ms time resolution.
Extended Data Fig. 10 Characterization of the mGluR2 PAM mutant conformational dynamics.
a, smFRET population histograms of 548UAA PAM mutant (C770A) in the presence of 5 µM (689 total molecules) or 10 µM (370 total molecules) glutamate. Histograms are fitted (red) to 4 Gaussian distributions (blue) centered around 0.31 (state 1), 0.51 (state 2), 0.71 (state 3) and 0.89 (state 4), denoted with dashed lines. Data represent mean ± s.e.m. of N=3 independent experiments. b, Normalized histograms of state 3 (pre-active) and state 4 (active) dwell times (cumulative count) in the presence of 5 or 10 µM glutamate for 548UAA and 548UAA PAM mutant (C770A). Data is fit to a single exponential decay function. Dwell times are from > 80 total molecules per condition from two independent experiments. c, TDPs of 548UAA and 548UAA PAM mutant (C770A) at 5 or 10 µM glutamate. Dashed lines represent 4 distinct FRET states. Transitions are compiled from two independent experiments. d, Dwell times of states 1-4 for 548UAA and 548UAA PAM mutant (C770A) in the presence of 5 or 10 µM glutamate. Average dwell time was calculated by fitting a single exponential decay function to dwell-time histograms for each condition. Dwell times of states 1 and 2 represent the mean of N=2 independent experiments with 86, 83, 91 and 72 total molecules examined for WT (5 µM), WT (10 µM), C770A (5 µM) and C770A (10 µM), respectively. Dwell times of states 3 and 4 represent the mean ± s.e.m. of N=3 independent experiments with 147, 140, 128 and 142 total molecules examined for WT (5 µM), WT (10 µM), C770A (5 µM) and C770A (10 µM), respectively. Transition and dwell-time analysis were performed on 100 ms data.
Supplementary information
Source data
Source Data Extended Data Fig. 1
Unprocessed gel image.
Rights and permissions
About this article
Cite this article
Liauw, B.WH., Afsari, H.S. & Vafabakhsh, R. Conformational rearrangement during activation of a metabotropic glutamate receptor. Nat Chem Biol 17, 291–297 (2021). https://doi.org/10.1038/s41589-020-00702-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-00702-5
This article is cited by
-
Stepwise activation of a metabotropic glutamate receptor
Nature (2024)
-
Fluorescence resonance energy transfer at the single-molecule level
Nature Reviews Methods Primers (2024)
-
Kinetic fingerprinting of metabotropic glutamate receptors
Communications Biology (2023)
-
Homo- and hetero-dimeric subunit interactions set affinity and efficacy in metabotropic glutamate receptors
Nature Communications (2023)
-
Single-molecule visualization of human A2A adenosine receptor activation by a G protein and constitutively activating mutations
Communications Biology (2023)