Abstract
The angiopoietin (Ang)–Tie pathway is essential for the proper maturation and remodeling of the vasculature. Despite its importance in disease, the mechanisms that control signal transduction through this pathway are poorly understood. Here, we demonstrate that heparan sulfate glycosaminoglycans (HS GAGs) regulate Ang–Tie signaling through direct interactions with both Ang ligands and Tie1 receptors. HS GAGs formed ternary complexes with Ang1 or Ang4 and Tie2 receptors, resulting in potentiation of endothelial survival signaling. In addition, HS GAGs served as ligands for the orphan receptor Tie1. The HS–Tie1 interaction promoted Tie1–Tie2 heterodimerization and enhanced Tie1 stability within the mature vasculature. Loss of HS–Tie1 binding using CRISPR–Cas9-mediated mutagenesis in vivo led to decreased Tie protein levels, pathway suppression and aberrant retinal vascularization. Together, these results reveal that sulfated glycans use dual mechanisms to regulate Ang–Tie signaling and are important for the development and maintenance of the vasculature.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Data generated or analyzed during this study are included in the article and related Supplementary Information or are available from the corresponding author on reasonable request. Publicly available data used in this study include the Tie2 crystal structure (PDB 2GY5), the Tie1 protein sequence (UniProt P35590), the Dec. 2011 murine genome assembly (GRCm38/mm10) and the CHOPCHOP gRNA design tool (https://chopchop.cbu.uib.no/). Source data are provided with this paper.
References
Potente, M., Gerhardt, H. & Carmeliet, P. Basic and therapeutic aspects of angiogenesis. Cell 146, 873–887 (2011).
Carmeliet, P. & Jain, R. K. Molecular mechanisms and clinical applications of angiogenesis. Nature 473, 298–307 (2011).
Adams, R. H. & Alitalo, K. Molecular regulation of angiogenesis and lymphangiogenesis. Nat. Rev. Mol. Cell Biol. 8, 464–478 (2007).
Simons, M., Gordon, E. & Claesson-Welsh, L. Mechanisms and regulation of endothelial VEGF receptor signalling. Nat. Rev. Mol. Cell Biol. 17, 611–625 (2016).
Beenken, A. & Mohammadi, M. The FGF family: biology, pathophysiology and therapy. Nat. Rev. Drug Discov. 8, 235–253 (2009).
Andrae, J., Gallini, R. & Betsholtz, C. Role of platelet-derived growth factors in physiology and medicine. Genes Dev. 22, 1276–1312 (2008).
Capila, I. & Linhardt, R. J. Heparin–protein interactions. Angew. Chem. Int. Ed. Engl. 41, 391–412 (2002).
Xu, D. & Esko, J. D. Demystifying heparan sulfate–protein interactions. Annu. Rev. Biochem. 83, 129–157 (2014).
Poulain, F. E. & Yost, H. J. Heparan sulfate proteoglycans: a sugar code for vertebrate development? Development 142, 3456–3467 (2015).
Bishop, J. R., Schuksz, M. & Esko, J. D. Heparan sulphate proteoglycans fine-tune mammalian physiology. Nature 446, 1030–1037 (2007).
Olczyk, P., Mencner, Ł. & Komosinska-Vassev, K. Diverse roles of heparan sulfate and heparin in wound repair. Biomed. Res. Int. 2015, 549417 (2015).
Forsberg, E. & Kjellen, L. Heparan sulfate: lessons from knockout mice. J. Clin. Invest. 108, 175–180 (2001).
Fuster, M. M. & Wang, L. Endothelial heparan sulfate in angiogenesis. Prog. Mol. Biol. Transl. Sci. 93, 179–212 (2010).
Jakobsson, L. et al. Heparan sulfate in trans potentiates VEGFR-mediated angiogenesis. Dev. Cell 10, 625–634 (2006).
Xu, D., Fuster, M. M., Lawrence, R. & Esko, J. D. Heparan sulfate regulates VEGF165- and VEGF121-mediated vascular hyperpermeability. J. Biol. Chem. 286, 737–745 (2011).
Schlessinger, J. et al. Crystal structure of a ternary FGF–FGFR–heparin complex reveals a dual role for heparin in FGFR binding and dimerization. Mol. Cell 6, 743–750 (2000).
Pellegrini, L., Burke, D. F., von Delft, F., Mulloy, B. & Blundell, T. L. Crystal structure of fibroblast growth factor receptor ectodomain bound to ligand and heparin. Nature 407, 1029–1034 (2000).
Abramsson, A. et al. Defective N-sulfation of heparan sulfate proteoglycans limits PDGF-BB binding and pericyte recruitment in vascular development. Genes Dev. 21, 316–331 (2007).
van Wijk, X. M. R. & van Kuppevelt, T. H. Heparan sulfate in angiogenesis: a target for therapy. Angiogenesis 17, 443–462 (2014).
Augustin, H. G., Koh, G. Y., Thurston, G. & Alitalo, K. Control of vascular morphogenesis and homeostasis through the angiopoietin–Tie system. Nat. Rev. Mol. Cell Biol. 10, 165–177 (2009).
Saharinen, P., Eklund, L. & Alitalo, K. Therapeutic targeting of the angiopoietin–Tie pathway. Nat. Rev. Drug Discov. 16, 635–661 (2017).
Sato, T. N. et al. Distinct roles of the receptor tyrosine kinases Tie-1 and Tie-2 in blood vessel formation. Nature 376, 70–74 (1995).
Puri, M. C., Rossant, J., Alitalo, K., Bernstein, A. & Partanen, J. The receptor tyrosine kinase Tie is required for integrity and survival of vascular endothelial cells. EMBO J. 14, 5884–5891 (1995).
Kim, M. et al. Opposing actions of angiopoietin-2 on Tie2 signaling and FOXO1 activation. J. Clin. Invest. 126, 3511–3525 (2016).
Partanen, J. et al. A novel endothelial cell surface receptor tyrosine kinase with extracellular epidermal growth factor homology domains. Mol. Cell. Biol. 12, 1698–1707 (1992).
Saharinen, P. et al. Multiple angiopoietin recombinant proteins activate the Tie1 receptor tyrosine kinase and promote its interaction with Tie2. J. Cell Biol. 169, 239–243 (2005).
Seegar, T. C. M. et al. Tie1–Tie2 interactions mediate functional differences between angiopoietin ligands. Mol. Cell 37, 643–655 (2010).
Leppänen, V.-M., Saharinen, P. & Alitalo, K. Structural basis of Tie2 activation and Tie2–Tie1 heterodimerization. Proc. Natl Acad. Sci. USA 114, 4376–4381 (2017).
Savant, S. et al. The orphan receptor Tie1 controls angiogenesis and vascular remodeling by differentially regulating Tie2 in tip and stalk cells. Cell Rep. 12, 1761–1773 (2015).
Korhonen, E. A. et al. Tie1 controls angiopoietin function in vascular remodeling and inflammation. J. Clin. Invest. 126, 3495–3510 (2016).
D’Amico, G. et al. Tie1 deletion inhibits tumor growth and improves angiopoietin antagonist therapy. J. Clin. Invest. 124, 824–834 (2014).
La Porta, S. et al. Endothelial Tie1-mediated angiogenesis and vascular abnormalization promote tumor progression and metastasis. J. Clin. Invest. 128, 834–845 (2018).
Brown, J. M. et al. A sulfated carbohydrate epitope inhibits axon regeneration after injury. Proc. Natl Acad. Sci. USA. 109, 4768–4773 (2012).
Rogers, C. J. et al. Elucidating glycosaminoglycan–protein–protein interactions using carbohydrate microarray and computational approaches. Proc. Natl Acad. Sci. USA. 108, 9747–9752 (2011).
Pulsipher, A., Griffin, M. E., Stone, S. E. & Hsieh-Wilson, L. C. Long-lived engineering of glycans to direct stem cell fate. Angew. Chem. Int. Ed. Engl. 54, 1466–1470 (2015).
Powell, A. K., Fernig, D. G. & Turnbull, J. E. Fibroblast growth factor receptors 1 and 2 interact differently with heparin/heparan sulfate: implications for dynamic assembly of a ternary signaling complex. J. Biol. Chem. 277, 28554–28563 (2002).
Teran, M. & Nugent, M. A. Synergistic binding of vascular endothelial growth factor-A and its receptors to heparin selectively modulates complex affinity. J. Biol. Chem. 290, 16451–16462 (2015).
Li, G. et al. Glycosaminoglycanomics of cultured cells using a rapid and sensitive LC-MS/MS approach. ACS Chem. Biol. 10, 1303–1310 (2015).
Griffith, A. R. et al. Predicting glycosaminoglycan surface protein interactions and implications for studying axonal growth. Proc. Natl Acad. Sci. USA. 114, 13697–13702 (2017).
Barton, W. A. et al. Crystal structures of the Tie2 receptor ectodomain and the angiopoietin-2–Tie2 complex. Nat. Struct. Mol. Biol. 13, 524–532 (2006).
Fredriksson, S. et al. Protein detection using proximity-dependent DNA ligation assays. Nat. Biotechnol. 20, 473–477 (2002).
Taichman, D. B. et al. A unique pattern of Tie1 expression in the developing murine lung. Exp. Lung Res. 29, 113–122 (2003).
Coles, C. H. et al. Proteoglycan-specific molecular switch for RPTPσ clustering and neuronal extension. Science 332, 484–488 (2011).
Meyer, R. D., Mohammadi, M. & Rahimi, N. A single amino acid substitution in the activation loop defines the decoy characteristic of VEGFR-1/FLT-1. J. Biol. Chem. 281, 867–875 (2006).
Vander Kooi, C. W. et al. Structural basis for ligand and heparin binding to neuropilin B domains. Proc. Natl Acad. Sci. USA 104, 6152–6157 (2007).
Ibrahimi, O. A., Zhang, F., Hrstka, S. C. L., Mohammadi, M. & Linhardt, R. J. Kinetic model for FGF, FGFR and proteoglycan signal transduction complex assembly. Biochemistry 43, 4724–4730 (2004).
Griffin, M. E. & Hsieh-Wilson, L. C. Glycan engineering for cell and developmental biology. Cell Chem. Biol. 23, 108–121 (2016).
Tully, S. E., Rawat, M. & Hsieh-Wilson, L. C. Discovery of a TNF-α antagonist using chondroitin sulfate microarrays. J. Am. Chem. Soc. 128, 7740–7741 (2006).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Mayo, S. L., Olafson, B. D. & Goddard, W. A. III DREIDING: a generic force field for molecular simulations. J. Phys. Chem. 94, 8897–8909 (1990).
Kutner, R. H., Zhang, X.-Y. & Reiser, J. Production, concentration and titration of pseudotyped HIV-1-based lentiviral vectors. Nat. Protoc. 4, 495–505 (2009).
Montague, T. G., Cruz, J. M., Gagnon, J. A., Church, G. M. & Valen, E. CHOPCHOP: a CRISPR–Cas9 and TALEN web tool for genome editing. Nucleic Acids Res. 42, W401–W407 (2014).
Mashiko, D. et al. Generation of mutant mice by pronuclear injection of circular plasmid expressing Cas9 and single guided RNA. Sci. Rep. 3, 3355 (2013).
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR–Cas-mediated genome engineering. Cell 153, 910–918 (2013).
Acknowledgements
We thank S. Pease and staff of the Caltech Genetically Engineered Mouse Services Core for help with generating the Tie1-2A mouse line and J. Costanza and A. Gomez of the Caltech Office of Laboratory Animal Resources for mouse line care and maintenance. We also thank J. Vielmetter and the Caltech Protein Expression Center of the Beckman Institute for help with conducting the SPR experiments. This work was supported by the NIH (5R01GM093627 and 5R01GM127920 to L.C.H.-W.) and the National Science Foundation (CBET-1805022 to W.A.G., DGE-1144469 to M.E.G. and DGE-1745301 to A.W.S.).
Author information
Authors and Affiliations
Contributions
M.E.G. and L.C.H.-W. conceived the project. Unless otherwise noted, M.E.G. performed the experimental work. A.W.S. conducted some of the microarray, ELISA and cell imaging assays. G.M.M. aided in assay optimization for initial binding experiments and conducted all computational work under the guidance of W.A.G.III. All authors contributed to the design of the experimental and computational work and to data analysis, discussed the results and commented on the manuscript. M.E.G. and L.C.H.-W. wrote the manuscript. L.C.H.-W. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Ang-Tie2 complex formation occurs in a sulfation-dependent manner.
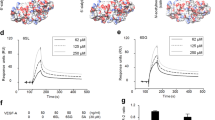
Binding of biotinylated HS sulfation motifs to immobilized Tie2 in the presence or absence of a Ang1 or b Ang4, as detected by ELISA. Removal of the N- and O-sulfate groups (De-N,O), the N-sulfate groups (De-N), the O-sulfate groups (De-O), the 6-O-sulfate groups (De-6O), or the 2-O-sulfate groups (De-2O), or conversion of the N-sulfate to N-acetyl groups (N-Ac) reduced formation of HS-Ang1/4-Tie2 ternary complexes. Data represent mean ± s.e.m.; n = 2 independent replicates.
Extended Data Fig. 2 Orphan receptor Tie1 binds to the CS-E motif.
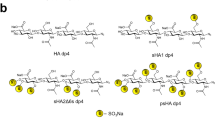
a, Chemical structure of the CS-E sulfation motif in CS polysaccharides. Interaction of CS-E and Tie1 by b ELISA, c glycan microarray, and d surface plasmon resonance. Dissociation constant for Tie1, as determined by ELISA: KD,app = 19.9 nM (10.5 to 53.5 nM); value represents mean (95% CI); graphed data represent n = 2 independent replicates. For glycan microarrays, data represent mean ± s.e.m.; n = 10 individual spots per glycan concentration. Dissociation constant for Tie1, as determined by SPR: KD,app = 14.7 nM (14.6 to 14.8 nM); value represents mean (95% CI). SPR data were fit using a 1:1 Langmuir model shown in red.
Extended Data Fig. 3 HS engages Tie1 in a sulfation-specific manner.
Binding of Tie1-Fc to different HS sulfation motifs, as determined by ELISA. Removal of the N- and O-sulfate groups (De-N,O), the O-sulfate groups (De-O), or the 6-O-sulfate groups (De-6O), or conversion of the N-sulfate to N-acetyl groups (N-Ac) abolished the HS–Tie1 interaction. Removal of the N-sulfate groups (De-N) or the 2-O-sulfate groups (De-2O) greatly reduced HS-Tie1 binding. Data represent mean ± s.e.m.; n = 2 independent replicates.
Extended Data Fig. 4 Soluble Tie1 binds to HUVECs.
Immunofluorescence imaging and orthogonal plane views of HUVECs with cell-associated Tie1-Fc (yellow), scale bar = 5 μm. Red bars indicate cross-sectional views in xz and yz images, which were produced by combining z-stack images using FIJI/ImageJ. The majority of Tie1 appears at the apical or basolateral cell surface. Representative images from four individual cells are shown.
Extended Data Fig. 5 Sulfated HS GAGs engage Tie1 in its N-terminal domains.
a Schematic and b Western blot of overexpressed, purified full-length (Tie1-FL) and C-terminal truncated (Tie-N) Tie1 ectodomains. Colors in (a) correspond to the ribbon diagram of Tie1 in Fig. 4a. Gray sections are derived from the expression plasmid. c, Binding of biotinylated triS-HS to recombinant Tie1-FL and Tie-N by ELISA. Data for (c) were obtained in the same experiment as Fig. 4e. Dissociation constants, as determined by ELISA: KD,app (Tie1-FL) = 6.51 nM (5.58 to 7.63 nM), KD,app (Tie1-N) = 6.26 nM (5.26 to 7.45 nM); values represent mean (95% CI); graphed data represent n = 2 independent replicates.
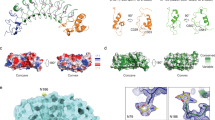
Extended Data Fig. 6 HS GAGs engage Tie1 in an electropositive region of the first N-terminal Ig-like domain.
a Ribbon diagram and b electrostatic potential surface of the Tie2 crystal structure (pdb 2GY5), which lacks the electropositive HS binding site found in Tie1. The red to blue scale in (b) represents relative electrostatic potential (electronegative to electropositive). c, Schematic of the first N-terminal Ig-like domain (Ig1) of Tie1, highlighting the six positively charged amino acids within 5-10 Å of the top 10 highest ranked HS GAG docked poses. All residues are mutated to alanine in the Tie1-6A construct, whereas only the two bolded residues are mutated in Tie1-2A. d, Structural model of the Tie1 N-terminal region (residues 22-447) indicating key amino acid residues involved in HS binding (purple) and those reported to be involved in Tie2 binding (orange)27. Distances are displayed in angstroms between the alpha carbon of each residue. e, Western blot showing expression of secreted Tie1 ectodomain constructs from HEK-293T cells after purification.
Extended Data Fig. 7 The HS-Tie1 interaction is driven primarily by ionic interactions.
a, Relative energetic contributions of van der Waals, hydrogen bonding, and Coulombic forces to the total calculated nonbonding energy are shown for the top 10 HS-Tie1 binding poses. In all cases, Coulombic forces dominated the calculated energy. b, Salt elution profile of Tie1 bound to heparin-sepharose shows loss of the HS-Tie1 interaction with increasing NaCl concentrations, indicative of strong ionic interactions between HS and Tie1. FT = flow-through.
Extended Data Fig. 8 Generation, genotyping, and characterization of Tie1-2A mice.
a, Schematic of the homology-directed repair method used to generate the Tie1-2A mouse line incorporating R38A and R82 A mutations into the endogenous murine Tie1 gene. b, Representative genotyping results from a Tie12A/wt x Tie12A/wt breeding pair after the Tie1 locus was amplified by PCR and digested with StuI. Animal 7 is Tie1wt/wt, animals 1, 2, 3, and 5 are Tie12A/wt, and animals 4 and 6 are Tie12A/2A. c, Quantification of weight differences between Tie12A/2A (2A/2A) and wild-type (wt/wt) 4-month-old, male littermates, n = 5 per genotype. d,e, Quantification of relative vessel area and branchpoints from retinal samples described in Fig. 6; n = 5 per genotype. Data represent mean ± s.e.m.; unpaired, two-tailed Student’s t test.
Extended Data Fig. 9 Tie1 mutation does not affect Erk1/2 phosphorylation in vivo.
a, Representative Western blots and b, quantification of pErk1/2 from lung tissue harvested from 7-day-old pups; n = 6 per genotype. pErk1/2 values were normalized to the corresponding total Erk1/2 levels and are reported relative to the wild-type animals. All gels were transferred to the same blot and imaged to allow for direct comparison between samples. Data represent mean ± s.e.m.; unpaired, two-tailed Student’s t test.
Extended Data Fig. 10 HS GAGs regulate the Ang/Tie signaling pathway.
HS GAGs utilize two distinct binding modes to promote pathway activation. a, Endothelial cell-associated HS GAGs recruit agonistic ligands Ang1 and Ang4 to form HS-Ang-Tie2 complexes and elicit downstream signaling. b, HS GAGs also bind to the orphan receptor Tie1 to promote the Tie1-Tie2 interaction, which maintains Tie2 protein levels and prevents protein turnover at the cell surface to sustain downstream signaling.
Supplementary information
Supplementary Information
Supplementary Figs. 1–4
Supplementary Data
Source Data for Supplementary Fig. 3
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data and unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data and unprocessed western blots.
Source Data Fig. 6
Statistical source data and unprocessed western blots.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data and unprocessed western blot.
Source Data Extended Data Fig. 6
Unprocessed western blot.
Source Data Extended Data Fig. 7
Statistical source data and unprocessed western blot.
Source Data Extended Data Fig. 8
Statistical source data and unprocessed DNA gel.
Source Data Extended Data Fig. 9
Statistical source data and unprocessed western blots.
Rights and permissions
About this article
Cite this article
Griffin, M.E., Sorum, A.W., Miller, G.M. et al. Sulfated glycans engage the Ang–Tie pathway to regulate vascular development. Nat Chem Biol 17, 178–186 (2021). https://doi.org/10.1038/s41589-020-00657-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-00657-7
This article is cited by
-
The niche matters: origin, function and fate of CNS-associated macrophages during health and disease
Acta Neuropathologica (2024)
-
The Provenance, Providence, and Position of Endothelial Cells in Injured Spinal Cord Vascular Pathology
Cellular and Molecular Neurobiology (2023)
-
Gestational chronic intermittent hypoxia induces hypertension, proteinuria, and fetal growth restriction in mice
Sleep and Breathing (2022)
-
A systems biology model of junctional localization and downstream signaling of the Ang–Tie signaling pathway
npj Systems Biology and Applications (2021)