Abstract
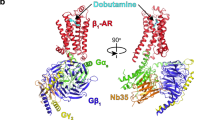
G-protein-coupled receptors (GPCRs) are seven-transmembrane proteins mediating cellular signals in response to extracellular stimuli. Although three-dimensional structures showcase snapshots that can be sampled in the process and nuclear magnetic resonance detects conformational equilibria, the mechanism by which agonist-activated GPCRs interact with various effectors remains elusive. Here, we used paramagnetic nuclear magnetic resonance for leucine amide resonances to visualize the structure of β2-adrenoreceptor in the full agonist-bound state, without thermostabilizing mutations abolishing its activity. The structure exhibited a unique orientation of the intracellular half of the transmembrane helix 6, forming a cluster of G-protein-interacting residues. Furthermore, analyses of efficacy-dependent chemical shifts of the residues near the pivotal PIF microswitch identified an equilibrium among three conformations, including one responsible for the varied signal level in each ligand-bound state. Together, these results provide a structural basis for the dynamic activation of GPCRs and shed light on GPCR-mediated signal transduction.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Atomic coordinates for β2AR-Δ in the fully activated state have been deposited in the PDB under accession code 6KR8. The NMR data and restraints used in the structure calculations have been deposited in the Biological Magnetic Resonance Data Bank under accession number 36284. The other data that support the findings of this study are available from the corresponding author upon reasonable request.
Code availability
All code used in this study is available from the corresponding author upon reasonable request.
References
Rosenbaum, D. M., Rasmussen, S. G. F. & Kobilka, B. K. The structure and function of G-protein-coupled receptors. Nature 459, 356–363 (2009).
Erlandson, S. C., McMahon, C. & Kruse, A. C. Structural basis for G-protein-coupled receptor signaling. Annu. Rev. Biophys. 47, 1–18 (2018).
Flock, T. et al. Selectivity determinants of GPCR: G-protein binding. Nature 545, 317–322 (2017).
Deupi, X. & Kobilka, B. K. Energy landscapes as a tool to integrate GPCR structure, dynamics, and function. Physiol. (Bethesda) 25, 293–303 (2010).
Shimada, I., Ueda, T., Kofuku, Y., Eddy, M. T. & Wüthrich, K. GPCR drug discovery: integrating solution NMR data with crystal and cryo-EM structures. Nat. Rev. Drug Discov. 18, 59–82 (2018).
Xiang, J. et al. Successful strategies to determine high-resolution structures of GPCRs. Trends Pharmacol. Sci. 37, 1055–1069 (2016).
Trzaskowski, B. et al. Action of molecular switches in GPCRs: theoretical and experimental studies. Curr. Med. Chem. 19, 1090–1109 (2012).
Edward Zhou, X., Melcher, K. & Eric Xu, H. Structural biology of G-protein‐coupled receptor signaling complexes. Protein Sci. 28, 487–501 (2019).
Warne, T. et al. The structural basis for agonist and partial agonist action on a β(1)-adrenergic receptor. Nature 469, 241–244 (2011).
Lebon, G. et al. Agonist-bound adenosine A2A receptor structures reveal common features of GPCR activation. Nature 474, 521–525 (2011).
Egloff, P. et al. Structure of signaling-competent neurotensin receptor 1 obtained by directed evolution in Escherichia coli. Proc. Natl Acad. Sci. USA 111, E655–E662 (2014).
Peng, Y. et al. 5-HT2C receptor structures reveal the structural basis of GPCR polypharmacology. Cell 172, 719–730 (2018).
Wacker, D. et al. Structural features for functional selectivity at serotonin receptors. Science 340, 615–619 (2013).
White, J. F. et al. Structure of the agonist-bound neurotensin receptor. Nature 490, 508–513 (2012).
Rasmussen, S. G. F. et al. Structure of a nanobody-stabilized active state of the β(2) adrenoceptor. Nature 469, 175–180 (2011).
Ring, A. M. et al. Adrenaline-activated structure of β2-adrenoceptor stabilized by an engineered nanobody. Nature 502, 575–579 (2013).
Rasmussen, S. G. et al. Crystal structure of the β2 adrenergic receptor-Gs protein complex. Nature 477, 549–555 (2011).
Heydenreich, F. M., Vuckovic, Z., Matkovic, M. & Veprintsev, D. B. Stabilization of G protein-coupled receptors by point mutations. Front. Pharm. 6, 82 (2015).
Kofuku, Y. et al. Efficacy of the β2-adrenergic receptor is determined by conformational equilibrium in the transmembrane region. Nat. Commun. 3, 1045 (2012).
Liu, J. J., Horst, R., Katritch, V., Stevens, R. C. & Wüthrich, K. Biased signaling pathways in β2-adrenergic receptor characterized by 19F-NMR. Science 1106, 1106–1111 (2012).
Solt, A. S. et al. Insight into partial agonism by observing multiple equilibria for ligand-bound and Gs-mimetic nanobody-bound β1-adrenergic receptor. Nat. Commun. 8, 1795 (2017).
Kruse, A. C. et al. Activation and allosteric modulation of a muscarinic acetylcholine receptor. Nature 504, 101–106 (2013).
Manglik, A. et al. Structural Insights into the dynamic process of β2-adrenergic receptor signaling. Cell 161, 1101–1111 (2015).
Gregorio, G. G. et al. Single-molecule analysis of ligand efficacy in β2AR-G-protein activation. Nature 547, 68–73 (2017).
Wingler, L. M. et al. Angiotensin analogs with divergent bias stabilize distinct receptor conformations. Cell 176, 468–478 (2019).
Eddy, M. T. et al. Allosteric coupling of drug binding and intracellular signaling in the A2A adenosine receptor. Cell 172, 68–80 (2018).
Roth, C. B., Hanson, M. A. & Stevens, R. C. Stabilization of the human β2-adrenergic receptor TM4–TM3–TM5 helix interface by mutagenesis of Glu1223.41, a critical residue in GPCR structure. J. Mol. Biol. 376, 1305–1319 (2008).
Gossert, A. D. et al. A simple protocol for amino acid type selective isotope labeling in insect cells with improved yields and high reproducibility. J. Biomol. NMR 51, 449–456 (2011).
Battiste, J. L. & Wagner, G. Utilization of site-directed spin labeling and high-resolution heteronuclear nuclear magnetic resonance for global fold determination of large proteins with limited nuclear Overhauser effect data. Biochemistry 39, 5355–5365 (2000).
Anthis, N. J. & Clore, G. M. Visualizing transient dark states by NMR spectroscopy. Q. Rev. Biophys. 48, 35–116 (2015).
King, G. J. et al. The Arabidopsis B3 domain protein VERNALIZATION1 (VRN1) is involved in processes essential for development, with structural and mutational studies revealing its DNA-binding surface. J. Biol. Chem. 288, 3198–3207 (2013).
Reibarkh, M., Malia, T. J. & Wagner, G. NMR distinction of single- and multiple-mode binding of small-molecule protein ligands. J. Am. Chem. Soc. 128, 2160–2161 (2006).
Hobbs, C. A., Bobay, B. G., Thompson, R. J., Perego, M. & Cavanagh, J. NMR solution structure and DNA-binding model of the DNA-binding domain of competence protein A. J. Mol. Biol. 398, 248–263 (2010).
Takeuchi, K., Imai, M. & Shimada, I. Dynamic equilibrium on DNA defines transcriptional regulation of a multidrug binding transcriptional repressor, LmrR. Sci. Rep. 7, 267 (2017).
Minato, Y., Ueda, T., Machiyama, A., Iwaï, H. & Shimada, I. Dynamic domain arrangement of CheA–CheY complex regulates bacterial thermotaxis, as revealed by NMR. Sci. Rep. 7, 16462 (2017).
Bain, A. D. Chemical exchange in NMR. Prog. Nucl. Magn. Reson. Spectrosc. 43, 63–103 (2003).
Waudby, C. A., Ramos, A., Cabrita, L. D. & Christodoulou, J. Two-dimensional NMR lineshape analysis. Sci. Rep. 6, 24826 (2016).
Hukushima, K. & Nemoto, K. Exchange Monte Carlo method and application to spin glass simulations. J. Phys. Soc. Jpn. 65, 1604–1608 (1996).
Nagata, K., Sugita, S. & Okada, M. Bayesian spectral deconvolution with the exchange Monte Carlo method. Neural Netw. 28, 82–89 (2012).
Fernández, C. & Wider, G. TROSY in NMR studies of the structure and function of large biological macromolecules. Curr. Opin. Struct. Biol. 13, 570–580 (2003).
Lapinaite, A. et al. The structure of the box C/D enzyme reveals regulation of RNA methylation. Nature 502, 519–523 (2013).
Gottstein, D., Reckel, S., Dötsch, V. & Güntert, P. Requirements on paramagnetic relaxation enhancement data for membrane protein structure determination by NMR. Structure 20, 1019–1027 (2012).
Isogai, S. et al. Backbone NMR reveals allosteric signal transduction networks in the β1-adrenergic receptor. Nature 314, 1–17 (2016).
Grzesiek, S., Cordier, F., Jaravine, V. & Barfield, M. Insights into biomolecular hydrogen bonds from hydrogen bond scalar couplings. Prog. Nucl. Magn. Reson. Spectrosc. 45, 275–300 (2004).
Zou, Y., Weis, W. I. & Kobilka, B. K. N-terminal T4 lysozyme fusion facilitates crystallization of a G protein coupled receptor. PLoS One 7, e46039 (2012).
Rosenbaum, D. M. et al. GPCR engineering yields high-resolution structural insights into 2-adrenergic receptor function. Science 318, 1266–1273 (2007).
Fung, J. J. et al. Ligand-regulated oligomerization of β(2)-adrenoceptors in a model lipid bilayer. EMBO J. 28, 3315–3328 (2009).
Nietlispach, D. Suppression of anti-TROSY lines in a sensitivity enhanced gradient selection TROSY scheme. J. Biomol. NMR 31, 161–166 (2005).
Helmus, J. J. & Jaroniec, C. P. NMRglue: an open source Python package for the analysis of multidimensional NMR data. J. Biomol. NMR 55, 355–367 (2013).
Sali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Schwieters, C. D., Bermejo, G. A. & Clore, G. M. Xplor-NIH for molecular structure determination from NMR and other data sources. Protein Sci. 27, 26–40 (2018).
Iwahara, J., Schwieters, C. D. & Clore, G. M. Ensemble approach for NMR structure refinement against 1H paramagnetic relaxation enhancement data arising from a flexible paramagnetic group attached to a macromolecule. J. Am. Chem. Soc. 126, 5879–5896 (2004).
McConnell, H. M. Reaction rates by nuclear magnetic resonance. J. Chem. Phys. 28, 430–431 (1958).
Acknowledgements
This work is supported by The Ministry of Education, Culture, Sports, Science and Technology and the Japan Society for the Promotion of Science KAKENHI grant number JP17H06097 and by the development of innovative drug discovery technologies for middle-sized molecules from the Japan Agency for Medical Research and Development (to I.S.).
Author information
Authors and Affiliations
Contributions
S.I. designed the study, constructed β2AR-Δ and its variants, purified proteins, conducted GTP turnover assays with T.Y., acquired NMR spectra, analyzed the PRE data and calculated the PRE structure and wrote the manuscript. Y.K. established the purification protocol of β2AR at the early stage of the project and constructed the plasmid for the expression of cystathionine-γ-synthase. Y.S. prepared the virus stock for the coexpression of the Gs heterotrimer and cultured the cells by using the virus stock. T.U. performed the exchange Monte Carlo calculation. I.S. designed the study, analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–13 and Supplementary Table 1
Rights and permissions
About this article
Cite this article
Imai, S., Yokomizo, T., Kofuku, Y. et al. Structural equilibrium underlying ligand-dependent activation of β2-adrenoreceptor. Nat Chem Biol 16, 430–439 (2020). https://doi.org/10.1038/s41589-019-0457-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-019-0457-5
This article is cited by
-
Binding kinetics drive G protein subtype selectivity at the β1-adrenergic receptor
Nature Communications (2024)
-
Recognition of methamphetamine and other amines by trace amine receptor TAAR1
Nature (2023)
-
QR code model: a new possibility for GPCR phosphorylation recognition
Cell Communication and Signaling (2022)
-
Structures of β1-adrenergic receptor in complex with Gs and ligands of different efficacies
Nature Communications (2022)
-
Structure-based design of a novel third-generation antipsychotic drug lead with potential antidepressant properties
Nature Neuroscience (2022)