Abstract
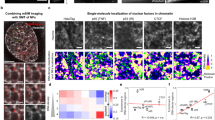
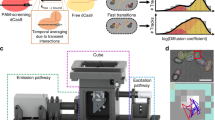
The enormous size of mammalian genomes means that for a DNA-binding protein the number of nonspecific, off-target sites vastly exceeds the number of specific, cognate sites. How mammalian DNA-binding proteins overcome this challenge to efficiently locate their target sites is not known. Here, through live-cell single-molecule tracking, we show that CCCTC-binding factor, CTCF, is repeatedly trapped in small zones that likely correspond to CTCF clusters, in a manner that is largely dependent on an internal RNA-binding region (RBRi). We develop a new theoretical model called anisotropic diffusion through transient trapping in zones to explain CTCF dynamics. Functionally, transient RBRi-mediated trapping increases the efficiency of CTCF target search by ~2.5-fold. Overall, our results suggest a ‘guided’ mechanism where CTCF clusters concentrate diffusing CTCF proteins near cognate binding sites, thus increasing the local ON-rate. We suggest that local guiding may allow DNA-binding proteins to more efficiently locate their target sites.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Raw and processed SPT data is freely available at Zenodo: https://zenodo.org/record/2208323. All cell lines will be provided upon request.
Code availability
Raw code as well as a detailed description of how the data was analyzed is available on GitLab: https://gitlab.com/anders.sejr.hansen/anisotropy. The code for localization and tracking is also available on GitLab: https://gitlab.com/tjian-darzacq-lab/SPT_LocAndTrack. The code for performing Brownian motion simulations (Supplementary Figs. 2a–c and 3) is likewise available on GitLab: https://gitlab.com/tjian-darzacq-lab/simSPT. Finally, the PALM-analysis code is also available on GitLab: https://gitlab.com/anders.sejr.hansen/palm_pipeline/.
References
Mao, Y. S., Zhang, B. & Spector, D. L. Biogenesis and function of nuclear bodies. Trends Genet. 27, 295–306 (2011).
Woringer, M. & Darzacq, X. Protein motion in the nucleus: from anomalous diffusion to weak interactions. Biochem. Soc. Trans. 46, 945–956 (2018).
Metzler, R., Jeon, J.-H., Cherstvy, A. G. & Barkai, E. Anomalous diffusion models and their properties: non-stationarity, non-ergodicity, and ageing at the centenary of single particle tracking. Phys. Chem. Chem. Phys. 16, 24128–24164 (2014).
Höfling, F. & Franosch, T. Anomalous transport in the crowded world of biological cells. Reports Prog. Phys. 76, 046602 (2013).
Rhodes, J., Mazza, D., Nasmyth, K. & Uphoff, S. Scc2/Nipbl hops between chromosomal cohesin rings after loading. eLife 6, e30000 (2017).
McSwiggen, D. T. et al. Evidence for DNA-mediated nuclear compartmentalization distinct from phase separation. eLife 8, e47098 (2019).
Bancaud, A., Lavelle, C., Huet, S. & Ellenberg, J. A fractal model for nuclear organization: current evidence and biological implications. Nucleic Acids Res. 40, 8783–8792 (2012).
Rice, S. A. Diffusion-limited reactions. Compr. Chem. Kinet. 25, 3–46 (1985).
Kapanidis, A. N., Uphoff, S. & Stracy, M. Understanding protein mobility in bacteria by tracking single molecules. J. Mol. Biol. 430, 4443–4455 (2018).
Pulkkinen, O. & Metzler, R. Distance matters: the impact of gene proximity in bacterial gene regulation. Phys. Rev. Lett. 110, 198101 (2017).
Kolesov, G., Wunderlich, Z., Laikova, O. N., Gelfand, M. S. & Mirny, L. A. How gene order is influenced by the biophysics of transcription regulation. Proc. Natl Acad. Sci. USA 104, 13948–13953 (2007).
van den Broek, B. et al. Coiling enhances target localization by proteins. Proc. Natl Acad. Sci. USA 105, 15738–15742 (2008).
Di Stefano, M., Rosa, A., Belcastro, V., di Bernardo, D. & Micheletti, C. Colocalization of coregulated genes: a steered molecular dynamics study of human chromosome 19. PLoS Comput. Biol. https://doi.org/10.1371/journal.pcbi.1003019 (2013).
Bauer, M. & Metzler, R. Generalized facilitated diffusion model for DNA-binding proteins with search and recognition states. Biophys. J. 102, 2321–2330 (2012).
Slutsky, M. & Mirny, L. A. Kinetics of protein-DNA interaction: facilitated target location in sequence-dependent potential. Biophys. J. 87, 4021–4035 (2004).
Lomholt, M., Ambjörnsson, T. & Metzler, R. Optimal target search on a fast-folding polymer chain with volume exchange. Phys. Rev. Lett. 95, 260603 (2005).
Rada-Iglesias, A., Grosveld, F. G. & Papantonis, A. Forces driving the three-dimensional folding of eukaryotic genomes. Mol. Syst. Biol. 14, e8214 (2018).
Hassler, M., Shaltiel, I. A. & Haering, C. H. Towards a unified model of SMC complex function. Curr. Biol. 28, R1266–R1281 (2018).
Hansen, A. S., Pustova, I., Cattoglio, C., Tjian, R. & Darzacq, X. CTCF and cohesin regulate chromatin loop stability with distinct dynamics. eLife 6, e25776 (2017).
Hansen, A. S. et al. Robust model-based analysis of single-particle tracking experiments with spot-on. eLife 7, e33125 (2018).
Elf, J., Li, G.-W. & Xie, X. S. Probing transcription factor dynamics at the single-molecule level in a living cell. Science 316, 1191–1194 (2007).
Di Rienzo, C., Piazza, V., Gratton, E., Beltram, F. & Cardarelli, F. Probing short-range protein Brownian motion in the cytoplasm of living cells. Nat. Commun. 5, 5891 (2014).
Manley, S. et al. High-density mapping of single-molecule trajectories with photoactivated localization microscopy. Nat. Methods 5, 155–157 (2008).
GrimmJ. B. et al. Bright photoactivatable fluorophores for single-molecule imaging. Nat. Methods 13, 985–988 (2016).
Persson, F., Lindén, M., Unoson, C. & Elf, J. Extracting intracellular diffusive states and transition rates from single-molecule tracking data. Nat. Methods 10, 265–269 (2013).
Izeddin, I. et al. Single-molecule tracking in live cells reveals distinct target-search strategies of transcription factors in the nucleus. eLife 2014, 1–27 (2014).
Liao, Y., Yang, S. K., Koh, K., Matzger, A. J. & Biteen, J. S. Heterogeneous single-molecule diffusion in one-, two-, and three-dimensional microporous coordination polymers: directional, trapped, and immobile guests. Nano Lett. 12, 3080–3085 (2012).
Burov, S. et al. Distribution of directional change as a signature of complex dynamics. Proc. Natl Acad. Sci. USA 110, 19689–19694 (2013).
Teves, S. S. et al. A dynamic mode of mitotic bookmarking by transcription factors.eLife 6, 22280 (2016).
Weber, S. C., Spakowitz, A. J. & Theriot, J. A. Bacterial chromosomal loci move subdiffusively through a viscoelastic cytoplasm.Phys. Rev. Lett. 104, 238102 (2010).
Weber, S. C., Thompson M. A., Moerner, W. E., Spakowitz, A. J. & Theriot, J. A. Analytical tools to distinguish the effects of localization error, confinement, and medium elasticity on the velocity autocorrelation function. Biophys. J. 102, 2443–2450 (2012).
Weber, S. C., Theriot, J. A. & Spakowitz, A. J. Subdiffusive motion of a polymer composed of subdiffusive monomers. Phys. Rev. E 82, 11913 (2010).
Amitai, A., Seeber, A., Gasser, S. M. & Holcman, D. Visualization of chromatin decompaction and break site extrusion as predicted by statistical polymer modeling of single-locus trajectories.Cell Rep. 18, 1200–1214 (2017).
Amitai, A. Chromatin configuration affects the dynamics and distribution of a transiently interacting protein. Biophys. J. 114, 766–771 (2018).
Saxton, M. J. A biological interpretation of transient anomalous subdiffusion. I. Qualitative model. Biophysj 92, 1178–1191 (2007).
Metzler, R. & Klafter, J. The random walk’s guide to anomalous diffusion: a fractional dynamics approach. Phys. Rep. 339, 1–77 (2000).
Hansen, A. S. et al. Distinct classes of chromatin loops revealed by deletion of an RNA-binding region in CTCF. Mol. Cell 76, 396–411 (2019).
Saldaña-Meyer, R. et al. CTCF regulates the human p53 gene through direct interaction with its natural antisense transcript, Wrap53. Genes Dev. 28, 723–734 (2014).
Rasko, J. E. J. et al. Cell growth inhibition by the multifunctional multivalent zinc-finger factor CTCF. Cancer Res. 61, 6002–6007 (2001).
Lampo, T. J., Stylianidou, S., Backlund, M. P., Wiggins, P. A. & Spakowitz, A. J. Cytoplasmic RNA-protein particles exhibit non-Gaussian subdiffusive behavior. Biophys. J. 112, 532–542 (2017).
Elmokadem, A. & Yu, J. Optimal drift correction for superresolution localization microscopy with Bayesian inference. Biophys. J. 109, 1772–1780 (2015).
Ester, M., Kriegel, H.-P., Sander, J. & Xu, X. In Proc. 2nd International Conference on Knowledge Discovery and Data Mining (ed Fayyad, Usama M.) 226–231 (1996).
Stone, M. B. & Veatch, S. L. Steady-state cross-correlations for live two-colour super-resolution localization data sets. Nat. Commun. 6, 7347 (2015).
Bronstein, I. et al. Transient anomalous diffusion of telomeres in the nucleus of mammalian cells. Phys. Rev. Lett. 103, 18102 (2009).
Golding, I. & Cox, E. C. Physical nature of bacterial cytoplasm. Phys. Rev. Lett. 96, 98102 (2006).
Saldaña-Meyer, R. et al. RNA interactions are essential for CTCF-mediated genome organization. Mol. Cell 76, 412–422 (2019).
Havlin, S. & Ben-Avraham, D. Diffusion in disordered media. Adv. Phys. 51, 187–292 (2002).
Cencini, M. & Pigolotti, S. Energetic funnel facilitates facilitated diffusion. Nucleic Acids Res. 46, 558–567 (2017).
Mir, M. et al. Dynamic multifactor hubs interact transiently with sites of active transcription in Drosophila embryos. eLife 7, e40497 (2018).
Tsai, A. et al. Nuclear microenvironments modulate transcription from low-affinity enhancers. elife 6, e28975 (2017).
Pettitt, S. J. et al. Agouti C57BL/6N embryonic stem cells for mouse genetic resources. Nat. Methods 6, 493–495 (2009).
Tokunaga, M., Imamoto, N. & Sakata-Sogawa, K. Highly inclined thin illumination enables clear single-molecule imaging in cells. Nat. Methods 5, 159–161 (2008).
Sergé, A., Bertaux, N., Rigneault, H. & Marguet, D. Dynamic multiple-target tracing to probe spatiotemporal cartography of cell membranes. Nat. Methods 5, 687–694 (2008).
Sprague, B. L., Pego, R. L., Stavreva, D. A. & McNally, J. G. Analysis of binding reactions by fluorescence recovery after photobleaching. Biophys. J. 86, 3473–3495 (2004).
Cattoglio, C. et al. Determining cellular CTCF and cohesin abundances to constrain 3D genome models. eLife 8, e40164 (2019).
Rossi, A. M. & Taylor, C. W. Analysis of protein-ligand interactions by fluorescence polarization. Nat. Protoc. 6, 365 (2011).
Mueller, F., Mazza, D., Stasevich, T. J. & McNally, J. G. FRAP and kinetic modeling in the analysis of nuclear protein dynamics: what do we really know? Curr. Opin. Cell Biol. 22, 403–411 (2010).
Acknowledgements
We thank L. Lavis for generously providing JF dyes, M. Woringer for insightful discussions and help with simSPT, A. Tangara and A. Robles for microscope assembly and maintenance, G. Dailey for help and assistance with cloning, K. Heydari at the Li Ka Shing Facility for flow cytometry assistance and A. deHart, L. Witowsky, A. Manford, L. Dahal and A. Basil Heckert for discussions and help with fluorescence polarization experiments. We thank A. Seeber, K. Dao Duc, D. McSwiggen and other members of the Tjian and Darzacq laboratories for comments on the manuscript. This work was performed in part at the CRL Molecular Imaging Center, supported by the Gordon and Betty Moore Foundation. A.S.H. was a postdoctoral fellow of the Siebel Stem Cell Institute and is supported by a National Institutes of Health (NIH) NIGMS K99 Pathway to Independence Award (no. K99GM130896). This work was supported by NIH grant nos. UO1-EB021236 and U54-DK107980 (X.D.), the California Institute of Regenerative Medicine grant no. LA1–08013 (X.D.) and by the Howard Hughes Medical Institute (003061, R.T.).
Author information
Authors and Affiliations
Contributions
A.S.H., A.A., R.T. and X.D. conceived of the project. A.S.H. and A.A. conceived of the ADTZ model. A.S.H. performed the experiments, developed anisotropy analysis pipeline, analyzed the experimental data and performed Brownian motion simulations. A.A. developed the theoretical framework, performed and analyzed model simulations. A.S.H. and C.C. generated the C59D2 ΔRBRi-Halo-CTCF mESC line. C.C. performed in vitro CTCF binding assays. A.S.H. and A.A. drafted the manuscript and all authors edited the manuscript. R.T. and X.D. supervised the project. A.S.H. and A.A. contributed equally to this project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 2 and 3, Figs. 1–15 and Notes 1–3.
Supplementary Table 1
Comparison of vbSPT and Spot-On. Effect of HMM (vbSPT) on filtering out bound population.
Supplementary Video 1
Comparison of vbSPT and Spot-On. Effect of HMM (vbSPT) on filtering out bound population.
Supplementary Video 2
Single Halo-CTCF protein exhibiting anomalous diffusion inside the mESC nucleus.
Supplementary Video 3
Single Halo-CTCF protein exhibiting anomalous diffusion inside the mESC nucleus.
Supplementary Video 4
Single Halo-CTCF protein exhibiting anomalous diffusion inside the mESC nucleus.
Rights and permissions
About this article
Cite this article
Hansen, A.S., Amitai, A., Cattoglio, C. et al. Guided nuclear exploration increases CTCF target search efficiency. Nat Chem Biol 16, 257–266 (2020). https://doi.org/10.1038/s41589-019-0422-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-019-0422-3
This article is cited by
-
Disordered C-terminal domain drives spatiotemporal confinement of RNAPII to enhance search for chromatin targets
Nature Cell Biology (2024)
-
Real-time single-molecule imaging of transcriptional regulatory networks in living cells
Nature Reviews Genetics (2024)
-
Enhancer selectivity in space and time: from enhancer–promoter interactions to promoter activation
Nature Reviews Molecular Cell Biology (2024)
-
Cargo-specific effects of hypoxia on clathrin-mediated trafficking
Pflügers Archiv - European Journal of Physiology (2024)
-
CTCF is a DNA-tension-dependent barrier to cohesin-mediated loop extrusion
Nature (2023)