Abstract
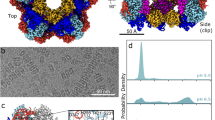
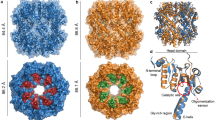
The flagellar hook protein FlgE from spirochaete bacteria self-catalyzes the formation of an unusual inter-subunit lysinoalanine (Lal) crosslink that is critical for cell motility. Unlike other known examples of Lal biosynthesis, conserved cysteine and lysine residues in FlgE spontaneously react to form Lal without the involvement of additional enzymes. Oligomerization of FlgE via its D0 and Dc domains drives assembly of the crosslinking site at the D1–D2 domain interface. Structures of the FlgED2 domain, dehydroalanine (DHA) intermediate and Lal crosslinked FlgE subunits reveal successive snapshots of the reaction. Cys178 flips from a buried configuration to release hydrogen sulfide (H2S/HS−) and produce DHA. Interface residues provide hydrogen bonds to anchor the active site, facilitate β-elimination of Cys178 and polarize the peptide backbone to activate DHA for reaction with Lys165. Cysteine-reactive molecules accelerate DHA formation, whereas nucleophiles can intercept the DHA intermediate, thereby indicating a potential for Lal crosslink inhibitors to combat spirochaetal diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Coordinates and structure files for WT FlgED2, DHA FlgED2 and Lal crosslinked FlgED1D2:D2 have been deposited to the Protein Data Bank with the following accession codes: 6NDW (WT FlgED2), 6NDT (DHA FlgED2) and 6NDX (FlgED1D2:D2 Lal crosslinked dimer). Raw MS data for all mutants and X-ray diffraction images are available from the corresponding author upon reasonable request. Constructs encoding for full-length and truncated Td FlgE variants are available from the corresponding author upon reasonable request.
References
Kang, H. J. & Baker, E. N. Intramolecular isopeptide bonds: protein crosslinks built for stress? Trends Biochem. Sci. 36, 229–237 (2011).
Walden, M., Crow, A., Nelson, M. D. & Banfield, M. J. Intramolecular isopeptide but not internal thioester bonds confer proteolytic and significant thermal stability to the S. pyogenes pilus adhesin Spy0125. Proteins Struct. Funct. Bioinforma. 82, 517–527 (2014).
Kwon, H. et al. Autocatalytically generated Thr-Gln ester bond crosslinks stabilize the repetitive Ig-domain shaft of a bacterial cell surface adhesin. PNAS 111, 1367–1372 (2014).
Baker, E. N., Squire, C. J. & Young, P. G. Self-generated covalent crosslinks in the cell-surface adhesins of Gram-positive bacteria. Biochem. Soc. Trans. 43, 787–794 (2015).
Popa, M. P., McKelvey, T. A., Hempel, J. & Hendrix, R. W. Bacteriophage HK97 structure: wholesale covalent crosslinking between the major head shell subunits. J. Virol. 65, 3227–3237 (1991).
Williams, M. & Baxter, R. The structure and function of thioester-containing proteins in arthropods. Biophys. Rev. 6, 261–272 (2014).
Miller, M. R. et al. Spirochaete flagella hook proteins self-catalyse a lysinoalanine covalent crosslink for motility. Nat. Microbiol. 1, 16134 (2016).
Charon, N. W. et al. The unique paradigm of spirochete motility and chemotaxis. Annu. Rev. Microbiol. 66, 349–370 (2012).
Wolgemuth, C. W. Flagellar motility of the pathogenic spirochetes. Semin. Cell Dev. Biol. 46, 104–112 (2015).
Shibata, S. et al. FliK regulates flagellar hook length as an internal ruler. Mol. Microbiol. 64, 1404–1415 (2007).
Zhao, X. et al. Cryoelectron tomography reveals the sequential assembly of bacterial flagella in Borrelia burgdorferi. Proc. Natl Acad. Sci. USA 110, 14390–14395 (2013).
Kojima, S. & Blair, D. F. The bacterial flagellar motor: structure and function of a complex molecular machine. Int. Rev. Cytol. 233, 93–134 (2004).
Samatey, F. A. et al. Structure of the bacterial flagellar hook and implication for the molecular universal joint mechanism. Nature 431, 1062–1068 (2004).
Zhao, X., Norris, S. J. & Liu, J. Molecular architecture of the bacterial flagellar motor in cells. Biochemistry 53, 4323–4333 (2014).
Alegre-Cebollada, J., Badilla, C. L. & Fernández, J. M. Isopeptide bonds block the mechanical extension of pili in pathogenic Streptococcus pyogenes. J. Biol. Chem. 285, 11235–11242 (2010).
Dierkes, L. E., Peebles, C. L., Firek, B. A., Hendrix, R. W. & Duda, R. L. Mutational analysis of a conserved glutamic acid required for self-catalyzed crosslinking of bacteriophage HK97 capsids. J. Virol. 83, 2088–2098 (2009).
Ross, P. D. et al. Crosslinking renders bacteriophage HK97 capsid maturation irreversible and effects an essential stabilization. EMBO J. 24, 1352–1363 (2005).
Duda, R. L. et al. Structure and energetics of encapsidated DNA in bacteriophage HK97 studied by scanning calorimetry and cryo-electron microscopy. J. Mol. Biol. 391, 471–483 (2009).
Friedman, M. Lysinoalanine in food and in antimicrobial proteins. Adv. Exp. Med. Biol. 459, 145–159 (1999).
Repka, L. M., Chekan, J. R., Nair, S. K. & van der Donk, W. A. Mechanistic understanding of lanthipeptide biosynthetic enzymes. Chem. Rev. 117, 5457–5520 (2017).
Huo, L., Ökesli, A. A., Zhao, M. & Van Der Donk, W. A. Insights into the biosynthesis of duramycin. Appl. Environ. Microbiol. 83, 2698–2714 (2017).
Ökesli, A., Cooper, L. E., Fogle, E. J. & van der Donk, W. A. Nine post-translational modifications during the biosynthesis of cinnamycin. J. Am. Chem. Soc. 133, 13753–13760 (2011).
An, L. et al. Substrate-assisted enzymatic formation of lysinoalanine in duramycin. Nat. Chem. Biol. 14, 928–933 (2018).
Miller, K. A. et al. Initial characterization of the FlgE hook high molecular weight complex of Borrelia burgdorferi. PLoS ONE 9, e98338 (2014).
Lukszo, J., Patterson, D., Albericio, F. & Kates, S. A. 3-(1-Piperidinyl)alanine formation during the preparation of C-terminal cysteine peptides with the Fmoc/t-Bu strategy. Lett. Pept. Sci. 3, 157–166 (1996).
Imada, K. Bacterial flagellar axial structure and its construction. Biophys. Rev. 10, 559–570 (2018).
Homma, M., DeRosier, D. J. & Macnab, R. M. Flagellar hook and hook-associated proteins of Salmonella typhimurium and their relationship to other axial components of the flagellum. J. Mol. Biol. 213, 819–832 (1990).
Moriya, N., Minamino, T., Hughes, K. T., Macnab, R. M. & Namba, K. The type III flagellar export specificity switch is dependent on flik ruler and a molecular clock. J. Mol. Biol. 359, 466–477 (2006).
Moriya, N. et al. Role of the Dc domain of the bacterial hook protein FlgE in hook assembly and function. Biophysics 9, 63–72 (2013).
Fujii, T., Kato, T. & Namba, K. Specific arrangement of alpha-helical coiled coils in the core domain of the bacterial flagellar hook for the universal joint function. Structure 17, 1485–1493 (2009).
Matsunami, H., Barker, C. S., Yoon, Y.-H., Wolf, M. & Samatey, F. A. Complete structure of the bacterial flagellar hook reveals extensive set of stabilizing interactions. Nat. Commun. 7, 13425 (2016).
Yoon, Y.-H. et al. Structural insights into bacterial flagellar hooks similarities and specificities. Sci. Rep. 6, 35552 (2016).
Grabarek, Z. & Gergely, J. Zero-length crosslinking procedure with the use of active esters. Anal. Biochem. 185, 131–135 (1990).
Brennan, D. F. & Barford, D. Eliminylation: a post-translational modification catalyzed by phosphothreonine lyases. Trends Biochem. Sci. 34, 108–114 (2009).
Zhou, G., Wang, H., Ma, Y., Chen, X. & An, N. B. D. fluorophore-based colorimetric and fluorescent chemosensor for hydrogen sulfide and its application for bioimaging. Tetrahedron 69, 867–870 (2013).
Lynch, M. J. & Crane, B. R. Design, validation, and application of an enzyme coupled hydrogen sulfide detection assay. Biochemistry 59, 474–483 (2019).
Kido, Y., Yoon, Y. H. & Samatey, F. A. Crystallization of a 79 kDa fragment of the hook protein FlgE from Campylobacter jejuni. Acta Crystallogr. Sect. F. 67, 1653–1657 (2011).
Chalker, J. M. et al. Methods for converting cysteine to dehydroalanine on peptides and proteins. Chem. Sci. 2, 1666 (2011).
Galan, S. R. G. et al. Post-translational site-selective protein backbone α-deuteration. Nat. Chem. Biol. 14, 955–963 (2018).
Shaikh, T. R. et al. A partial atomic structure for the flagellar hook of Salmonella typhimurium. Proc. Natl Acad. Sci. USA 102, 1023–1028 (2005).
Bharadwaj, K. C. Acryl activation by intramolecular hydrogen bond: Morita Baylis Hillman reaction of acrylamide with broad substrate scope. Chem. Sel. 2, 5384–5389 (2017).
Chevance, F. F. V. & Hughes, K. T. Coordinating assembly of a bacterial macromolecular machine. Nat. Rev. Microbiol. 6, 455–465 (2008).
Glavas, S. & Tanner, M. E. Catalytic acid/base residues of glutamate racemase. Biochemistry 38, 4106–4113 (1999).
Roberts, M. C., Chung, W. O. & Roe, D. E. Characterization of tetracycline and erythromycin resistance determinants in Treponema denticola. Antimicrob. Agents Chemother. 40, 1690–1694 (1996).
Mitjà, O. et al. Re-emergence of yaws after single mass azithromycin treatment followed by targeted treatment: a longitudinal study. Lancet 391, 1599–1607 (2018).
Nikolaidis, I., Favini-Stabile, S. & Dessen, A. Resistance to antibiotics targeted to the bacterial cell wall. Protein Sci. 23, 243–259 (2014).
Bernardes, G. J. L., Chalker, J. M., Errey, J. C. & Davis, B. G. Facile conversion of cysteine and alkyl cysteines to dehydroalanine on protein surfaces: versatile and switchable access to functionalized proteins. J. Am. Chem. Soc. 130, 5052–5053 (2008).
Dadová, J., Galan, S. R. & Davis, B. G. Synthesis of modified proteins via functionalization of dehydroalanine. Curr. Opin. Chem. Biol. 46, 71–81 (2018).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. Sect. D. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Cryst. 66, 486–501 (2010).
Emsley, P. et al. COOT: model-building tools for molecular graphics. Acta Crystallogr. Sect. D. 60, 2126–2132 (2004).
The PyMOL Molecular Graphics System, V.2.0 (Schrödinger, LLC, 2019).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Yang, Y., Thannhauser, T. W., Li, L. & Zhang, S. Development of an integrated approach for evaluation of 2-D gel image analysis: Impact of multiple proteins in single spots on comparative proteomics in conventional 2-D gel/MALDI workflow. Electrophoresis 28, 2080–2094 (2007).
Thomas, C. J., Cleland, T. P., Zhang, S., Gundberg, C. M. & Vashishth, D. Identification and characterization of glycation adducts on osteocalcin. Anal. Biochem. 525, 46–53 (2017).
Yang, Y., Anderson, E. & Zhang, S. Evaluation of six sample preparation procedures for qualitative and quantitative proteomics analysis of milk fat globule membrane. Electrophoresis 39, 2332–2339 (2018).
Acknowledgements
This work was supported by NIH grant no. R35 122535 (B.R.C.), the CBI Training grant nos. T32 GM008500 (M.J.L. and B.R.C.), NIH R01-DE023431 (N.C., M.M. and C.L.), AI078958 (C.L.) and NIH SIG grant no. 1S10 OD017992-01 (S.Z.). CHESS is supported by the NSF and NIH/NIGMS (no. DMR-1332208). MacCHESS is supported by NIH/NIGMS (no. GM-103485). Remote data collection was performed at the NE-CAT beamlines (no. GM124165) using an Eiger detector (OD021527) at the Advanced Photon Source (DE-AC02-06CH11357). The authors would like to thank H. Le for assistance with LC–MS experiments, A. Bilwes-Crane for editing the manuscript, the Cornell Proteomic and MS Facility for providing the mass spectrometry data and E. Anderson and R. Bahwal for technical assistance with MS sample preparation, data acquisition and analysis.
Author information
Authors and Affiliations
Contributions
B.R.C., M.J.L., N.W.C., C.L. and M.M. conceived the study. K.Z. generated an initial construct encoding for Td FlgE. M.J.L. performed the crystallography, X-ray diffraction experiments and associated structural analysis, site-directed mutagenesis, EDC crosslinking and Lal crosslinking SDS–PAGE assays. M.J. carried out preliminary work that identified NEM and DTNB as FlgE crosslink enhancers. M.J.L. prepared samples for MS. S.Z. performed MS data acquisition and associated samples analysis. M.J.L. and B.R.C. prepared figures. M.J.L., B.R.C. and N.W.C. wrote the manuscript. B.R.C., N.W.C., M.J.L., C.L., M.J., M.M. and S.Z. edited the manuscript. B.R.C., N.W.C., M.M. and C.L. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplemental Figs. 1–13 and Supplemental Table 1.
Rights and permissions
About this article
Cite this article
Lynch, M.J., Miller, M., James, M. et al. Structure and chemistry of lysinoalanine crosslinking in the spirochaete flagella hook. Nat Chem Biol 15, 959–965 (2019). https://doi.org/10.1038/s41589-019-0341-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-019-0341-3
This article is cited by
-
An overview of the structure and function of the flagellar hook FlgE protein
World Journal of Microbiology and Biotechnology (2023)