Abstract
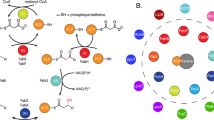
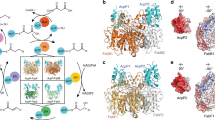
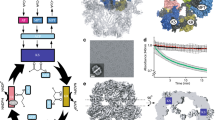
Fatty acid synthases are dynamic ensembles of enzymes that can biosynthesize long hydrocarbon chains efficiently. Here we visualize the interaction between the Escherichia coli acyl carrier protein (AcpP) and β-ketoacyl-ACP-synthase I (FabB) using X-ray crystallography, NMR, and molecular dynamics simulations. We leveraged this structural information to alter lipid profiles in vivo and provide a molecular basis for how protein–protein interactions can regulate the fatty acid profile in E. coli.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
References
Finzel, K., Lee, D. J. & Burkart, M. D. Chembiochem 16, 528–547 (2015)..
Feng, Y. & Cronan, J. E. J. Biol. Chem. 284, 29526–29535 (2009).
Wright, H. T. & Reynolds, K. A. Curr. Opin. Microbiol. 10, 447–453 (2007).
Yu, X., Liu, T., Zhu, F. & Khosla, C. Proc. Natl Acad. Sci. USA 108, 18643–18648 (2011).
Gajewski, J. et al. Nat. Chem. Biol. 13, 363–365 (2017).
Rock, C. & Cronan, J. E. Methods Enzymol. 71, 341–351 (1981).
Kim, Y. & Prestegard, J. H. Biochemistry 28, 8792–8797 (1989).
Roujeinikova, A. et al. J. Mol. Biol. 365, 135–145 (2007).
Majerus, P. W., Alberts, A. W. & Vagelos, P. R. Proc. Natl Acad. Sci. USA 53, 410–417 (1965).
Chan, D. I., Stockner, T., Tieleman, D. P. & Vogel, H. J. J. Biol. Chem. 283, 33620–33629 (2008).
Crosby, J. & Crump, M. P. Nat. Prod. Rep. 29, 1111 (2012).
Thiele, G. A. R. et al. Biochemistry 56, 2533–2536 (2017).
Nguyen, C. et al. Nature 505, 427–431 (2014).
Cronan, J. E. Biochem. J. 460, 157–163 (2014).
Beld, J., Cang, H. & Burkart, M. D. Angew. Chem. Int. Ed. Engl. 53, 14456–14461 (2014).
Worthington, A. S., Rivera, H., Torpey, J. W., Alexander, M. D. & Burkart, M. D. ACS Chem. Biol. 1, 687–691 (2006).
Worthington, A. S. & Burkart, M. D. Org. Biomol. Chem. 4, 44–46 (2006).
Byers, D. M. & Gong, H. Biochem. Cell Biol. 85, 649–662 (2007).
Zhang, L. et al. Cell Res. 26, 1330–1344 (2016).
De Lay, N. R. & Cronan, J. E. J. Bacteriol. 188, 287–296 (2006).
Garwin, J. L. & Cronan, J. E. Jr. J. Bacteriol. 141, 1457–1459 (1980).
Magnuson, K., Jackowski, S., Rock, C. O. & Cronan, J. E. Jr. Microbiol. Rev. 57, 522–542 (1993).
Garwin, J. L., Klages, A. L. & Cronan, J. E. Jr. J. Biol. Chem. 255, 3263–3265 (1980).
Rossini, E., Gajewski, J., Klaus, M., Hummer, G. & Grininger, M. Chem. Commun. 54, 11606–11609 (2018).
De Lay, N. R. & Cronan, J. E. J. Biol. Chem. 282, 20319–20328 (2007).
Kosa, N. M., Haushalter, R. W., Smith, A. R. & Burkart, M. D. Nat. Methods 9, 981–984 (2012).
Battye, T. G. G., Kontogiannis, L., Johnson, O., Powell, H. R. & Leslie, A. G. W. Acta Crystallogr. D 67, 271–281 (2011).
Adams, P. D. et al. Acta Crystallogr. D 66, 213–221 (2010).
Delaglio, F. et al. J. Biomol. NMR 6, 277–293 (1995).
Lee, W., Tonelli, M. & Markley, J. L. Bioinformatics 31, 1325–1327 (2015).
Williamson, M. P. Prog. Nucl. Magn. Reson. Spectrosc. 73, 1–16 (2013).
Pettersen, E. F. et al. J. Comput. Chem. 25, 1605–1612 (2004).
Vanquelef, E. et al. R.E.D. Nucleic Acids Res. 39, W511–W517 (2011).
Acknowledgements
S.C.T. and M.D.B. are supported by NIH GM100305 and GM095970. M.D.B. is also supported by National Science Foundation (NSF) Division of Integrative Organismal Systems (IOS) grant 1516156, and R.L. is also supported by NIH GM093040 and GM079383. This research used resources of the Advanced Light Source, which is a Department of Energy (DOE) Office of Science User Facility under contract no. DE-AC02-05CH11231. The authors thank the staff of beamline 8.2.1 at the Advanced Light Source for support during X-ray diffraction data collection, X. Huang and S. Opella for their guidance and assistance with NMR collection at the UCSD Biomolecular NMR facility, and B. Fuglestad for many helpful discussions on NMR. The authors thank J. E. Cronan for the CY1877 strain. Additional funding from the institutional Chemical and Structural Biology Training Grant (National Institute of General Medical Sciences grant T32GM108561) and the National Science Foundation Graduate Research Fellowship is also acknowledged.
Author information
Authors and Affiliations
Contributions
J.C.M. performed the crystallography and structural analysis, and prepared the manuscript. D.J.L. performed protein NMR, cloning and in vivo complementation, fatty acid analysis, and also prepared the manuscript. D.R.J. performed crystallography and structural analysis. A.J.S. performed MD simulations and analysis. J.B. performed synthesis of the crosslinker and fatty acid analysis. J.F.B. performed structural refinement and validation. J.J.H. performed GCMS and analysis. R.L. provided computational support and supervised MD and dry laboratory work. M.D.B. supervised synthesis, protein NMR, fatty acid complementation, and GCMS analysis. S.C.T. supervised crystallography and wet laboratory work. All authors contributed to editing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–3, Supplementary Figures 1–20
Supplementary Video 1
Principal component analysis of AcpP-FabB molecular dynamics simulations.
Supplementary Video 2
Principal component analysis of AcpP-FabB molecular dynamics simulations.
Rights and permissions
About this article
Cite this article
Milligan, J.C., Lee, D.J., Jackson, D.R. et al. Molecular basis for interactions between an acyl carrier protein and a ketosynthase. Nat Chem Biol 15, 669–671 (2019). https://doi.org/10.1038/s41589-019-0301-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-019-0301-y
This article is cited by
-
Enzymology of assembly line synthesis by modular polyketide synthases
Nature Chemical Biology (2023)
-
Helicobacter pylori FabX contains a [4Fe-4S] cluster essential for unsaturated fatty acid synthesis
Nature Communications (2021)
-
Biochemical and structural characterization of the BioZ enzyme engaged in bacterial biotin synthesis pathway
Nature Communications (2021)
-
Elucidation of transient protein-protein interactions within carrier protein-dependent biosynthesis
Communications Biology (2021)
-
Escherichia coli Nissle 1917 secondary metabolism: aryl polyene biosynthesis and phosphopantetheinyl transferase crosstalk
Applied Microbiology and Biotechnology (2021)