Abstract
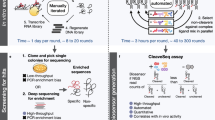
Cellular processes are carried out by many genes, and their study and optimization requires multiple levers by which they can be independently controlled. The most common method is via a genetically encoded sensor that responds to a small molecule. However, these sensors are often suboptimal, exhibiting high background expression and low dynamic range. Further, using multiple sensors in one cell is limited by cross-talk and the taxing of cellular resources. Here, we have developed a directed evolution strategy to simultaneously select for lower background, high dynamic range, increased sensitivity, and low cross-talk. This is applied to generate a set of 12 high-performance sensors that exhibit >100-fold induction with low background and cross-reactivity. These are combined to build a single “sensor array” in the genomes of E. coli MG1655 (wild-type), DH10B (cloning), and BL21 (protein expression). These “Marionette” strains allow for the independent control of gene expression using 12 small-molecule inducers.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Additional data supporting this study are available from the corresponding author upon reasonable request. The sequences of the following plasmids and strains are provided in GenBank: pAJM.711 (PPhlF-YFP) MH101715; pAJM.712 (PCymRC-YFP) MH101716; pAJM.713 (PLuxB-YFP) MH101717; pAJM.714 (PVanCC-YFP) MH101718; pAJM.715 (PTac-YFP) MH101719; pAJM.717 (PTet*-YFP) MH101720; pAJM.716 (PBAD-YFP) MH101721; pAJM.718 (PBetI-YFP) MH101722; pAJM.719 (PTtg-YFP) MH101723; pAJM.1459 (P3B5C-YFP) MH101724; pAJM.721 (PSalTTC-YFP) MH101725; pAJM.944 (PCin-YFP) MH101726; pAJM.847 (PhlFAM + PPhlF-YFP) MH101727; pAJM.657 (CymRAM + PCymRC-YFP) MH101728; pAJM.474 (LuxR + PLuxB-YFP) MH101729; pAJM.773 (VanRAM + PVanCC-YFP) MH101730; pAJM.336 (LacIAM + PTac-YFP) MH101731; pAJM.011 (TetR + PTet*-YFP) MH101732; pAJM.677 (AraCAM + AraE + PBAD-YFP) MH101733; pAJM.683 (BetIAM + PBetI-YFP) MH101734; pAJM.661 (TtgRAM + PTtg-YFP) MH101735; pAJM.690 (PcaUAM + P3B5B-YFP) MH101736; pAJM.771 (NahRAM + PSalTTC-YFP) MH101737; pAJM.1642 (CinRAM + PCin-YFP) MH101738; pAJM.884 (AcuRAM + PAcu-YFP) MH101739; pAJM.969 (MphRAM + EryR + PMph-YFP) MH101740. The following plasmids and strains can be acquired from Addgene: pAJM.711 (PPhlF-YFP) 108512; pAJM.712 (PCymRC-YFP) 108513; pAJM.713 (PLuxB-YFP) 108514; pAJM.714 (PVanCC-YFP) 108515; pAJM.715 (PTac-YFP) 108516; pAJM.717 (PTet*-YFP) 108517; pAJM.716 (PBAD-YFP) 108518; pAJM.718 (PBetI-YFP) 108519; pAJM.719 (PTtg-YFP) 108520; pAJM.1459 (P3B5C-YFP) 108521; pAJM.721 (PSalTTC-YFP) 108522; pAJM.944 (PCin-YFP) 108523; pAJM.847 (PhlFAM + PPhlF-YFP) 108524; pAJM.657 (CymRAM + PCymRC-YFP) 108525; pAJM.474 (LuxR + PLuxB-YFP) 108526; pAJM.773 (VanRAM + PVanCC-YFP) 108527; pAJM.336 (LacIAM + PTac-YFP) 108528; pAJM.011 (TetR + PTet*-YFP) 108529; pAJM.677 (AraCAM + AraE + PBAD-YFP) 108530; pAJM.683 (BetIAM + PBetI-YFP) 108531; pAJM.661 (TtgRAM + PTtg-YFP) 108532; pAJM.690 (PcaUAM + P3B5B-YFP) 108533; pAJM.771 (NahRAM + PSalTTC-YFP) 108534; pAJM.1642 (CinRAM + PCin-YFP) 108535; pAJM.884 (AcuRAM + PAcu-YFP) 108536; pAJM.969 (MphRAM + EryR + PMph-YFP) 108537; sAJM.1504 (Marionette-Clo) 108251; sAJM.1505 (Marionette-Pro) 108253; sAJM.1506 (Marionette-Wild) 108254.
References
Zuo, J. & Chua, N. H. Chemical-inducible systems for regulated expression of plant genes. Curr. Opin. Biotechnol. 11, 146–151 (2000).
Keyes, W. M. & Mills, A. A. Inducible systems see the light. Trends Biotechnol. 21, 53–55 (2003).
Mijakovic, I., Petranovic, D. & Jensen, P. R. Tunable promoters in systems biology. Curr. Opin. Biotechnol. 16, 329–335 (2005).
de Boer, H. A., Comstock, L. J. & Vasser, M. The tac promoter: a functional hybrid derived from the trp and lac promoters. Proc. Natl. Acad. Sci. USA 80, 21–25 (1983).
Guzman, L. M., Belin, D., Carson, M. J. & Beckwith, J. Tight regulation, modulation, and high-level expression by vectors containing the arabinose PBAD promoter. J. Bacteriol. 177, 4121–4130 (1995).
Skerra, A. Use of the tetracycline promoter for the tightly regulated production of a murine antibody fragment in Escherichia coli. Gene 151, 131–135 (1994).
Lutz, R. & Bujard, H. Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1-I2 regulatory elements. Nucleic Acids Res. 25, 1203–1210 (1997).
Urban, J. H. & Vogel, J. Translational control and target recognition by Escherichia coli small RNAs in vivo. Nucleic Acids Res. 35, 1018–1037 (2007).
Cookson, N. A. et al. Queueing up for enzymatic processing: correlated signaling through coupled degradation. Mol. Syst. Biol. 7, 561 (2011).
Elowitz, M. B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000).
Gardner, T. S., Cantor, C. R. & Collins, J. J. Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000).
Golding, I. & Cox, E. C. RNA dynamics in live Escherichia coli cells. Proc. Natl. Acad. Sci. USA 101, 11310–11315 (2004).
Kalir, S. & Alon, U. Using a quantitative blueprint to reprogram the dynamics of the flagella gene network. Cell 117, 713–720 (2004).
Golding, I., Paulsson, J., Zawilski, S. M. & Cox, E. C. Real-time kinetics of gene activity in individual bacteria. Cell 123, 1025–1036 (2005).
Voigt, C. A. Genetic parts to program bacteria. Curr. Opin. Biotechnol. 17, 548–557 (2006).
Caliando, B. J. & Voigt, C. A. Targeted DNA degradation using a CRISPR device stably carried in the host genome. Nat. Commun. 6, 6989 (2015).
Lee, S. K. et al. Directed evolution of AraC for improved compatibility of arabinose- and lactose-inducible promoters. Appl. Environ. Microbiol. 73, 5711–5715 (2007).
Scott, S. R. & Hasty, J. Quorum sensing communication modules for microbial consortia. ACS Synth. Biol. 5, 969–977 (2016).
Tashiro, Y. et al. Directed evolution of the autoinducer selectivity of Vibrio fischeri LuxR. J. Gen. Appl. Microbiol. 62, 240–247 (2016).
Halleran, A. D. & Murray, R. M. Cell-free and in vivo characterization of Lux, Las, and Rpa quorum activation systems in E. coli. ACS Synth. Biol. 7, 752–755 (2018).
Gyorgy, A. et al. Isocost lines describe the cellular economy of genetic circuits. Biophys. J. 109, 639–646 (2015).
Callura, J. M., Cantor, C. R. & Collins, J. J. Genetic switchboard for synthetic biology applications. Proc. Natl. Acad. Sci. USA 109, 5850–5855 (2012).
Bintu, L. et al. Transcriptional regulation by the numbers: models. Curr. Opin. Genet. Dev. 15, 116–124 (2005).
Salis, H., Tamsir, A. & Voigt, C. Engineering bacterial signals and sensors. Contrib. Microbiol. 16, 194–225 (2009).
Daber, R., Sochor, M. A. & Lewis, M. Thermodynamic analysis of mutant lac repressors. J. Mol. Biol. 409, 76–87 (2011).
Gatti-Lafranconi, P., Dijkman, W. P., Devenish, S. R. & Hollfelder, F. A single mutation in the core domain of the lac repressor reduces leakiness. Microb. Cell. Fact. 12, 67 (2013).
Ike, K. et al. Evolutionary design of choline-inducible and -repressible T7-based induction systems. ACS Synth. Biol. 4, 1352–1360 (2015).
Ellefson, J. W., Ledbetter, M. P. & Ellington, A. D. Directed evolution of a synthetic phylogeny of programmable Trp repressors. Nat. Chem. Biol. 14, 361–367 (2018).
Yokobayashi, Y., Weiss, R. & Arnold, F. H. Directed evolution of a genetic circuit. Proc. Natl. Acad. Sci. USA 99, 16587–16591 (2002).
Tang, S. Y., Fazelinia, H. & Cirino, P. C. AraC regulatory protein mutants with altered effector specificity. J. Am. Chem. Soc. 130, 5267–5271 (2008).
Tashiro, Y., Fukutomi, H., Terakubo, K., Saito, K. & Umeno, D. A nucleoside kinase as a dual selector for genetic switches and circuits. Nucleic Acids Res. 39, e12 (2011).
Taylor, N. D. et al. Engineering an allosteric transcription factor to respond to new ligands. Nat. Methods 13, 177–183 (2016).
Maranhao, A. C. & Ellington, A. D. Evolving orthogonal suppressor tRNAs to incorporate modified amino acids. ACS Synth. Biol. 6, 108–119 (2017).
Thyer, R., Filipovska, A. & Rackham, O. Engineered rRNA enhances the efficiency of selenocysteine incorporation during translation. J. Am. Chem. Soc. 135, 2–5 (2013).
Ellefson, J. W. et al. Directed evolution of genetic parts and circuits by compartmentalized partnered replication. Nat. Biotechnol. 32, 97–101 (2014).
Diaz Ricci, J. C. & Hernández, M. E. Plasmid effects on Escherichia coli metabolism. Crit. Rev. Biotechnol. 20, 79–108 (2000).
Kunjapur, A. M. & Prather, K. L. J. Development of a vanillate biosensor for the vanillin biosynthesis pathway in E. coli. https://doi.org/10.1101/375287 (2018).
Khlebnikov, A., Datsenko, K. A., Skaug, T., Wanner, B. L. & Keasling, J. D. Homogeneous expression of the P(BAD) promoter in Escherichia coli by constitutive expression of the low-affinity high-capacity AraE transporter. Microbiology 147, 3241–3247 (2001).
Scott, M., Gunderson, C. W., Mateescu, E. M., Zhang, Z. & Hwa, T. Interdependence of cell growth and gene expression: origins and consequences. Science 330, 1099–1102 (2010).
Lee, J. W. et al. Creating single-copy genetic circuits. Mol. Cell 63, 329–336 (2016).
Barrick, J. E. & Lenski, R. E. Genome dynamics during experimental evolution. Nat. Rev. Genet. 14, 827–839 (2013).
Lee, M. E., Aswani, A., Han, A. S., Tomlin, C. J. & Dueber, J. E. Expression-level optimization of a multi-enzyme pathway in the absence of a high-throughput assay. Nucleic Acids Res. 41, 10668–10678 (2013).
Zelcbuch, L. et al. Spanning high-dimensional expression space using ribosome-binding site combinatorics. Nucleic Acids Res. 41, e98 (2013).
Smanski, M. J. et al. Functional optimization of gene clusters by combinatorial design and assembly. Nat. Biotechnol. 32, 1241–1249 (2014).
Ghodasara, A. & Voigt, C. A. Balancing gene expression without library construction via a reusable sRNA pool. Nucleic Acids Res. 45, 8116–8127 (2017).
Yoon, S. H. et al. Engineering the lycopene synthetic pathway in E. coli by comparison of the carotenoid genes of Pantoea agglomerans and Pantoea ananatis. Appl. Microbiol. Biotechnol. 74, 131–139 (2007).
Alon, U., Surette, M. G., Barkai, N. & Leibler, S. Robustness in bacterial chemotaxis. Nature 397, 168–171 (1999).
Zhang, Y., Lara-Tejero, M., Bewersdorf, J. & Galán, J. E. Visualization and characterization of individual type III protein secretion machines in live bacteria. Proc. Natl. Acad. Sci. USA 114, 6098–6103 (2017).
Larson, M. H. et al. CRISPR interference (CRISPRi) for sequence-specific control of gene expression. Nat. Protoc. 8, 2180–2196 (2013).
Gupta, A., Reizman, I. M., Reisch, C. R. & Prather, K. L. Dynamic regulation of metabolic flux in engineered bacteria using a pathway-independent quorum-sensing circuit. Nat. Biotechnol. 35, 273–279 (2017).
Blattner, F. R. et al. The complete genome sequence of Escherichia coli K-12. Science 277, 1453–1462 (1997).
Kittleson, J. T., Cheung, S. & Anderson, J. C. Rapid optimization of gene dosage in E. coli using DIAL strains. J. Biol. Eng. 5, 10 (2011).
Green, R. & Rogers, E. J. Transformation of chemically competent E. coli. Methods Enzymol. 529, 329–336 (2013).
Meyer, A. J., Ellefson, J. W. & Ellington, A. D. Library generation by gene shuffling. Curr. Protoc. Mol. Biol. 105, 15.12.1–15.12.7 (2014).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 97, 6640–6645 (2000).
Thomason, L. C., Costantino, N. & Court, D. L. E. coli genome manipulation by P1 transduction. Curr. Protoc. Mol. Biol. 79, 1.17.1–1.17.8 (2007).
Li, G. W., Burkhardt, D., Gross, C. & Weissman, J. S. Quantifying absolute protein synthesis rates reveals principles underlying allocation of cellular resources. Cell 157, 624–635 (2014).
Volkmer, B. & Heinemann, M. Condition-dependent cell volume and concentration of Escherichia coli to facilitate data conversion for systems biology modeling. PLoS One 6, e23126 (2011).
Reetz, M. T. & Carballeira, J. D. Iterative saturation mutagenesis (ISM) for rapid directed evolution of functional enzymes. Nat. Protoc. 2, 891–903 (2007).
Salis, H. M., Mirsky, E. A. & Voigt, C. A. Automated design of synthetic ribosome binding sites to control protein expression. Nat. Biotechnol. 27, 946–950 (2009).
Acknowledgements
This work was supported by the US Office of Naval Research Multidisciplinary University Research Initiative grant #N00014-16-1-2388 (A.J.M., T.H.S.-S., E.G., J.Z., and C.A.V.). This work was supported in part by the Koch Institute Support (core) Grant P30-CA14051 from the National Cancer Institute. We would like to thank A.M. Kunjapur and K.L.J. Prather (Department of Chemical Engineering, Massachusetts Institute of Technology) for providing DNA templates for the amplification of PVan, P3B5, vanR, and pcaU and performing the initial characterization of VanR in E. coli. We would also like to thank S. Liu at the MIT-Broad Foundry for assisting in the RNA-seq and ribosome profiling sequencing run.
Author information
Authors and Affiliations
Contributions
A.J.M. and C.A.V. conceived the study and designed the experiments; E.G. performed the mass spectrometry experiments; J.Z. performed the RNA sequencing and ribosome profiling experiments; A.J.M. performed all other experiments. A.J.M., T.H.S.-S., E.G., and J.Z., analyzed the data; A.J.M. and C.A.V. wrote the manuscript with input from all the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–12, Supplementary Figures 1–21, Supplementary Notes 1–4
Rights and permissions
About this article
Cite this article
Meyer, A.J., Segall-Shapiro, T.H., Glassey, E. et al. Escherichia coli “Marionette” strains with 12 highly optimized small-molecule sensors. Nat Chem Biol 15, 196–204 (2019). https://doi.org/10.1038/s41589-018-0168-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0168-3
This article is cited by
-
Genetic toolbox for Photorhabdus and Xenorhabdus: pSEVA based heterologous expression systems and CRISPR/Cpf1 based genome editing for rapid natural product profiling
Microbial Cell Factories (2024)
-
Engineering is evolution: a perspective on design processes to engineer biology
Nature Communications (2024)
-
Snowprint: a predictive tool for genetic biosensor discovery
Communications Biology (2024)
-
Synthetic microbe-to-plant communication channels
Nature Communications (2024)
-
Synthetic microbiology in sustainability applications
Nature Reviews Microbiology (2024)