Abstract
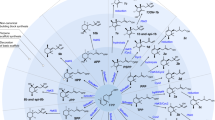
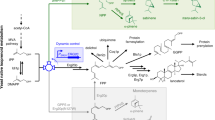
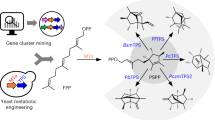
One application of synthetic biology is the redesign of existing biological systems to acquire new functions. In this context, expanding the chemical code underlying key biosynthetic pathways will lead to the synthesis of compounds with new structures and potentially new biological activities. Terpenoids are a large group of specialized metabolites with numerous applications. Yet, being synthesized from five-carbon units, they are restricted to distinct classes that differ by five carbon atoms (C10, C15, C20, etc.). To expand the diversity of terpenoid structures, we engineered yeast cells to synthesize a noncanonical building block with 11 carbons, and produced 40 C11 terpene scaffolds that can form the basis for an entire terpenoid class. By identifying a single-residue switch that converts C10 plant monoterpene synthases to C11-specific enzymes, we engineered dedicated synthases for C11 terpene production. This approach will enable the systematic expansion of the chemical space accessed by terpenoids.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Dewick, P. M. Medicinal natural products: a biosynthetic approach. (Wiley, Chichester, UK, 2009).
Pichersky, E., Noel, J. P. & Dudareva, N. Biosynthesis of plant volatiles: nature’s diversity and ingenuity. Science 311, 808–811 (2006).
Gershenzon, J. & Dudareva, N. The function of terpene natural products in the natural world. Nat. Chem. Biol. 3, 408–414 (2007).
Ignea, C. et al. Reconstructing the chemical diversity of labdane-type diterpene biosynthesis in yeast. Metab. Eng. 28, 91–103 (2015).
Jiang, J., He, X. & Cane, D. E. Biosynthesis of the earthy odorant geosmin by a bifunctional Streptomyces coelicolor enzyme. Nat. Chem. Biol. 3, 711–715 (2007).
Herde, M. et al. Identification and regulation of TPS04/GES, an Arabidopsis geranyllinalool synthase catalyzing the first step in the formation of the insect-induced volatile C16-homoterpene TMTT. Plant Cell 20, 1152–1168 (2008).
Lee, S. et al. Herbivore-induced and floral homoterpene volatiles are biosynthesized by a single P450 enzyme (CYP82G1) in Arabidopsis. Proc. Natl Acad. Sci. USA 107, 21205–21210 (2010).
Magnard, J. L. et al. PLANT VOLATILES. Biosynthesis of monoterpene scent compounds in roses. Science 349, 81–83 (2015).
Richter, A. et al. Characterization of biosynthetic pathways for the production of the volatile homoterpenes DMNT and TMTT in Zea mays. Plant Cell 28, 2651–2665 (2016).
Vedula, L. S. et al. Exploring biosynthetic diversity with trichodiene synthase. Arch. Biochem. Biophys. 466, 260–266 (2007).
Rising, K. A. et al. Formation of a novel macrocyclic alkaloid from the unnatural farnesyl diphosphate analogue anilinogeranyl diphosphate by 5-epi-aristolochene synthase. ACS. Chem. Biol. 10, 1729–1736 (2015).
Karahadian, C., Josephson, D. B. & Lindsay, R. C. Volatile compounds from Penicillium sp. contributing musty-earthy notes to Brie and Camembert cheese flavors. J. Agric. Food. Chem. 33, 339–343 (1985).
Komatsu, M., Tsuda, M., Omura, S., Oikawa, H. & Ikeda, H. Identification and functional analysis of genes controlling biosynthesis of 2-methylisoborneol. Proc. Natl Acad. Sci. USA 105, 7422–7427 (2008).
Wang, C. M. & Cane, D. E. Biochemistry and molecular genetics of the biosynthesis of the earthy odorant methylisoborneol in Streptomyces coelicolor. J. Am. Chem. Soc. 130, 8908–8909 (2008).
Dickschat, J. S. et al. Biosynthesis of the off-flavor 2-methylisoborneol by the myxobacterium Nannocystis exedens. Angew. Chem. Int. Edn Engl. 46, 8287–8290 (2007).
Brock, N. L., Ravella, S. R., Schulz, S. & Dickschat, J. S. A detailed view of 2-methylisoborneol biosynthesis. Angew. Chem. Int. Edn Engl. 52, 2100–2104 (2013).
Dickschat, J. S. Bacterial terpene cyclases. Nat. Prod. Rep. 33, 87–110 (2016).
Chou, W. K., Ikeda, H. & Cane, D. E. Cloning and characterization of Pfl_1841, a 2-methylenebornane synthase in Pseudomonas fluorescens PfO-1. Tetrahedron 67, 6627–6632 (2011).
Giglio, S., Chou, W. K., Ikeda, H., Cane, D. E. & Monis, P. T. Biosynthesis of 2-methylisoborneol in cyanobacteria. Environ. Sci. Technol. 45, 992–998 (2011).
Yamada, Y. et al. Terpene synthases are widely distributed in bacteria. Proc. Natl Acad. Sci. USA 112, 857–862 (2015).
Schumann, R. & Pendleton, P. Dehydration products of 2-methylisoborneol. Water Res. 31, 1243–1246 (1997).
Zhou, Y. J. et al. Modular pathway engineering of diterpenoid synthases and the mevalonic acid pathway for miltiradiene production. J. Am. Chem. Soc. 134, 3234–3241 (2012).
Ignea, C., Pontini, M., Maffei, M. E., Makris, A. M. & Kampranis, S. C. Engineering monoterpene production in yeast using a synthetic dominant negative geranyl diphosphate synthase. ACS Synth. Biol. 3, 298–306 (2014).
Stanley Fernandez, S. M., Kellogg, B. A. & Poulter, C. D. Farnesyl diphosphate synthase. Altering the catalytic site to select for geranyl diphosphate activity. Biochemistry 39, 15316–15321 (2000).
Jiang, Z., Kempinski, C., Bush, C. J., Nybo, S. E. & Chappell, J. Engineering triterpene and methylated triterpene production in plants provides biochemical and physiological insights into terpene metabolism. Plant Physiol. 170, 702–716 (2016).
Springer, M., Weissman, J. S. & Kirschner, M. W. A general lack of compensation for gene dosage in yeast. Mol. Syst. Biol. 6, 368 (2010).
Köksal, M., Chou, W. K., Cane, D. E. & Christianson, D. W. Structure of 2-methylisoborneol synthase from Streptomyces coelicolor and implications for the cyclization of a noncanonical C-methylated monoterpenoid substrate. Biochemistry 51, 3011–3020 (2012).
Chou, W. K., Gould, C. A. & Cane, D. E. Incubation of 2-methylisoborneol synthase with the intermediate analog 2-methylneryl diphosphate. J. Antibiot. (Tokyo) 70, 625–631 (2017).
Citron, C. A., Barra, L., Wink, J. & Dickschat, J. S. Volatiles from nineteen recently genome sequenced actinomycetes. Org. Biomol. Chem. 13, 2673–2683 (2015).
Kampranis, S. C. et al. Rational conversion of substrate and product specificity in a Salvia monoterpene synthase: structural insights into the evolution of terpene synthase function. Plant Cell 19, 1994–2005 (2007).
Phillips, M. A., Wildung, M. R., Williams, D. C., Hyatt, D. C. & Croteau, R. cDNA isolation, functional expression, and characterization of (+)-alpha-pinene synthase and (-)-alpha-pinene synthase from loblolly pine (Pinus taeda): stereocontrol in pinene biosynthesis. Arch. Biochem. Biophys. 411, 267–276 (2003).
Iijima, Y. et al. The biochemical and molecular basis for the divergent patterns in the biosynthesis of terpenes and phenylpropenes in the peltate glands of three cultivars of basil. Plant Physiol. 136, 3724–3736 (2004).
Iijima, Y., Gang, D. R., Fridman, E., Lewinsohn, E. & Pichersky, E. Characterization of geraniol synthase from the peltate glands of sweet basil. Plant Physiol. 134, 370–379 (2004).
Lücker, J. et al. Monoterpene biosynthesis in lemon (Citrus limon). cDNA isolation and functional analysis of four monoterpene synthases. Eur. J. Biochem. 269, 3160–3171 (2002).
Tsaballa, A. et al. Use of the de novo transcriptome analysis of silver-leaf nightshade (Solanum elaeagnifolium) to identify gene expression changes associated with wounding and terpene biosynthesis. BMC Genomics 16, 504 (2015).
Hyatt, D. C. et al. Structure of limonene synthase, a simple model for terpenoid cyclase catalysis. Proc. Natl Acad. Sci. USA 104, 5360–5365 (2007).
Pettersen, E. F. et al. UCSF Chimera--a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Ignea, C. et al. Improving yeast strains using recyclable integration cassettes, for the production of plant terpenoids. Microb. Cell. Fact. 10, 4 (2011).
Genee, H. J. et al. Software-supported USER cloning strategies for site-directed mutagenesis and DNA assembly. ACS Synth. Biol. 4, 342–349 (2015).
Ignea, C. et al. Positive genetic interactors of HMG2 identify a new set of genetic perturbations for improving sesquiterpene production in Saccharomyces cerevisiae. Microb. Cell. Fact. 11, 162 (2012).
Trikka, F. A. et al. Combined metabolome and transcriptome profiling provides new insights into diterpene biosynthesis in S. pomifera glandular trichomes. BMC Genomics 16, 935 (2015).
Finkelstein, J., Antony, E., Hingorani, M. M. & O’Donnell, M. Overproduction and analysis of eukaryotic multiprotein complexes in Escherichia coli using a dual-vector strategy. Anal. Biochem. 319, 78–87 (2003).
Ignea, C. et al. Carnosic acid biosynthesis elucidated by a synthetic biology platform. Proc. Natl Acad. Sci. USA 113, 3681–3686 (2016).
Zaed, A. M. & Sutherland, A. Stereoselective synthesis of the bicyclic guanidine alkaloid (+)-monanchorin. Org. Biomol. Chem. 8, 4394–4399 (2010).
Winterfeldt, E. Applications of diisobutylaluminium hydride (DIBAL) and triisobulyaluminium (TIBA) as reducing agents in organic synthesis. 7, Synthesis 617–630 (1975).
Dixit, V. M., Laskovics, F. M., Noall, W. I. & Poulter, C. D. Tris(tetrabutylammonium) hydrogen pyrophosphate. A new reagent for the preparation of allylic pyrophosphate esters. J. Org. Chem. 46, 1967–1969 (1981).
Miyata, O., Ozawa, Y., Ninomiya, I. & Naito, T. Radical cyclization in heterocycle synthesis. Part 10:1A concise synthesis of (-)-kainic acid via sulfanyl radical addition-cyclization-elimination reaction. Tetrahedron 56, 6199–6207 (2000).
Peitsch, M. C. ProMod and Swiss-Model: Internet-based tools for automated comparative protein modelling. Biochem. Soc. Trans. 24, 274–279 (1996).
Acknowledgements
We would like to thank D. Cane and W. Chou (Brown University, USA) for providing the bacterial construct pET28a(+)/PlGPPMT, C.E. Vickers (University of Queensland, Australia) for providing construct pCEV-G2-Ph/ClLimS, F. Geu-Flores (University of Copenhagen, Denmark) for critical reading of the manuscript, and M. Raadam for technical assistance. We are grateful to D.I. Pattison and E. Lazaridi from the PLEN metabolomics platform for their support. This work was supported by the Greek General Secretariat of Research and Technology (GSRT) grant 11ΣΥΝ_3_770 (to A.M.M. and S.C.K.) and the Novo Nordisk Foundation grants NNF16OC0019554 (to C.I.) and NNF16OC0021760 (to S.C.K.).
Author information
Authors and Affiliations
Contributions
S.C.K. conceived the project and designed experiments; C.I. designed experiments, engineered MIC1 strain, expressed and analyzed C10 and C11 wild-type and mutant synthases in yeast and bacteria, performed mutagenesis of PlGPPMT, PtPinS, SpSabS, ObMyrS, ClLimS, SeCamS, produced the purified proteins, and conducted in vitro enzymatic assays for the determination of the kinetic parameters; M.P. performed expression and analysis of PlGPPMT, SfCinS1 wild-type and mutants, wild-type SpSabS, and conducted yeast optimization for 2meGPP production, M.S.M. performed the chemical synthesis of 2meGPP; M.E.M. and A.M.M. assisted in data analysis; S.C.K. and C.I. analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–9, Supplementary Figures 1–11, Supplementary Notes 1–2
Rights and permissions
About this article
Cite this article
Ignea, C., Pontini, M., Motawia, M.S. et al. Synthesis of 11-carbon terpenoids in yeast using protein and metabolic engineering. Nat Chem Biol 14, 1090–1098 (2018). https://doi.org/10.1038/s41589-018-0166-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0166-5
This article is cited by
-
Engineered enzymes for the synthesis of pharmaceuticals and other high-value products
Nature Synthesis (2024)
-
The microbial biosynthesis of noncanonical terpenoids
Applied Microbiology and Biotechnology (2024)
-
Widespread biosynthesis of 16-carbon terpenoids in bacteria
Nature Chemical Biology (2023)
-
Overexpression of CiMYC2 Transcription Factor from Chrysanthemum indicum var. aromaticum Resulted in Modified Trichome Formation and Terpenoid Biosynthesis in Transgenic Tobacco
Journal of Plant Growth Regulation (2023)
-
Expanding the terpene biosynthetic code with non-canonical 16 carbon atom building blocks
Nature Communications (2022)