Abstract
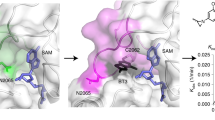
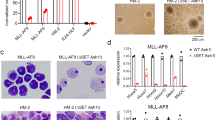
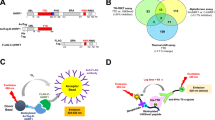
Chemical probes of epigenetic ‘readers’ of histone post-translational modifications (PTMs) have become powerful tools for mechanistic and functional studies of their target proteins in normal physiology and disease pathogenesis. Here we report the development of the first class of chemical probes of YEATS domains, newly identified ‘readers’ of histone lysine acetylation (Kac) and crotonylation (Kcr). Guided by the structural analysis of a YEATS–Kcr complex, we developed a series of peptide-based inhibitors of YEATS domains by targeting a unique π-π-π stacking interaction at the proteins’ Kcr recognition site. Further structure optimization resulted in the selective inhibitors preferentially binding to individual YEATS-containing proteins including AF9 and ENL with submicromolar affinities. We demonstrate that one of the ENL YEATS-selective inhibitors, XL-13m, engages with endogenous ENL, perturbs the recruitment of ENL onto chromatin, and synergizes the BET and DOT1L inhibition-induced downregulation of oncogenes in MLL-rearranged acute leukemia.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Crystal structure data of AF9 YEATS domain bound to inhibitor XL-07i has been deposited in the Protein Data Bank (PDB) under accession code 5YYF. Other data support the findings of this study are included in the article and/or the associated supplementary files, or available from the corresponding authors upon reasonable request.
References
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Kouzarides, T. SnapShot: histone-modifying enzymes. Cell 131, 822 (2007).
Patel, D. J. & Wang, Z. Readout of epigenetic modifications. Annu. Rev. Biochem. 82, 81–118 (2013).
Musselman, C. A., Lalonde, M. E., Côté, J. & Kutateladze, T. G. Perceiving the epigenetic landscape through histone readers. Nat. Struct. Mol. Biol. 19, 1218–1227 (2012).
Taverna, S. D., Li, H., Ruthenburg, A. J., Allis, C. D. & Patel, D. J. How chromatin-binding modules interpret histone modifications: lessons from professional pocket pickers. Nat. Struct. Mol. Biol. 14, 1025–1040 (2007).
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001).
Suganuma, T. & Workman, J. L. Signals and combinatorial functions of histone modifications. Annu. Rev. Biochem. 80, 473–499 (2011).
Chi, P., Allis, C. D. & Wang, G. G. Covalent histone modifications—miswritten, misinterpreted and mis-erased in human cancers. Nat. Rev. Cancer 10, 457–469 (2010).
Bhaumik, S. R., Smith, E. & Shilatifard, A. Covalent modifications of histones during development and disease pathogenesis. Nat. Struct. Mol. Biol. 14, 1008–1016 (2007).
Arrowsmith, C. H., Bountra, C., Fish, P. V., Lee, K. & Schapira, M. Epigenetic protein families: a new frontier for drug discovery. Nat. Rev. Drug. Discov. 11, 384–400 (2012).
Helin, K. & Dhanak, D. Chromatin proteins and modifications as drug targets. Nature 502, 480–488 (2013).
Cole, P. A. Chemical probes for histone-modifying enzymes. Nat. Chem. Biol. 4, 590–597 (2008).
Wagner, J. M., Hackanson, B., Lübbert, M. & Jung, M. Histone deacetylase (HDAC) inhibitors in recent clinical trials for cancer therapy. Clin. Epigenetics 1, 117–136 (2010).
Marmorstein, R. & Zhou, M. M. Writers and readers of histone acetylation: structure, mechanism, and inhibition. Cold Spring Harb. Perspect. Biol. 6, a018762 (2014).
Filippakopoulos, P. & Knapp, S. Targeting bromodomains: epigenetic readers of lysine acetylation. Nat. Rev. Drug. Discov. 13, 337–356 (2014).
Shortt, J., Ott, C. J., Johnstone, R. W. & Bradner, J. E. A chemical probe toolbox for dissecting the cancer epigenome. Nat. Rev. Cancer 17, 160–183 (2017).
Li, Y. et al. AF9 YEATS domain links histone acetylation to DOT1L-mediated H3K79 methylation. Cell 159, 558–571 (2014).
Zhao, D., Li, Y., Xiong, X., Chen, Z. & Li, H. YEATS Domain-A histone acylation reader in health and disease. J. Mol. Biol. 429, 1994–2002 (2017).
Schulze, J. M., Wang, A. Y. & Kobor, M. S. YEATS domain proteins: a diverse family with many links to chromatin modification and transcription. Biochem. Cell. Biol. 87, 65–75 (2009).
Li, Y. et al. Molecular coupling of histone crotonylation and active transcription by AF9 YEATS domain. Mol. Cell 62, 181–193 (2016).
Andrews, F. H. et al. The Taf14 YEATS domain is a reader of histone crotonylation. Nat. Chem. Biol. 12, 396–398 (2016).
Zhao, D. et al. YEATS2 is a selective histone crotonylation reader. Cell Res. 26, 629–632 (2016).
Zhang, Q. et al. Structural insights into histone crotonyl-lysine recognition by the AF9 YEATS domain. Structure 24, 1606–1612 (2016).
Wan, L. et al. ENL links histone acetylation to oncogenic gene expression in acute myeloid leukaemia. Nature 543, 265–269 (2017).
Erb, M. A. et al. Transcription control by the ENL YEATS domain in acute leukaemia. Nature 543, 270–274 (2017).
Mi, W. et al. YEATS2 links histone acetylation to tumorigenesis of non-small cell lung cancer. Nat. Commun. 8, 1088 (2017).
Li, Y., Zhao, D., Chen, Z. & Li, H. YEATS domain: linking histone crotonylation to gene regulation. Transcription 8, 9–14 (2017).
Niphakis, M. J. & Cravatt, B. F. Enzyme inhibitor discovery by activity-based protein profiling. Annu. Rev. Biochem. 83, 341–377 (2014).
Li, X. & Kapoor, T. M. Approach to profile proteins that recognize post-translationally modified histone “tails”. J. Am. Chem. Soc. 132, 2504–2505 (2010).
Yang, T., Liu, Z. & Li, X. D. Developing diazirine-based chemical probes to identify histone modification ‘readers’ and ‘erasers’. Chem. Sci. 6, 1011–1017 (2015).
Jafari, R. et al. The cellular thermal shift assay for evaluating drug target interactions in cells. Nat. Protoc. 9, 2100–2122 (2014).
Martinez Molina, D. & Nordlund, P. The cellular thermal shift assay: a novel biophysical assay for in situ drug target engagement and mechanistic biomarker studies. Annu. Rev. Pharmacol. Toxicol. 56, 141–161 (2016).
Jang, M. K. et al. The bromodomain protein Brd4 is a positive regulatory component of P-TEFb and stimulates RNA polymerase II-dependent transcription. Mol. Cell 19, 523–534 (2005).
Basheer, F. & Huntly, B. J. P. BET bromodomain inhibitors in leukemia. Exp. Hematol. 43, 718–731 (2015).
Fierz, B. & Muir, T. W. Chromatin as an expansive canvas for chemical biology. Nat. Chem. Biol. 8, 417–427 (2012).
Huston, A., Arrowsmith, C. H., Knapp, S. & Schapira, M. Probing the epigenome. Nat. Chem. Biol. 11, 542–545 (2015).
McGaughey, G. B., Gagné, M. & Rappé, A. K. pi-Stacking interactions. Alive and well in proteins. J. Biol. Chem. 273, 15458–15463 (1998).
Cho, K. I., Kim, D. & Lee, D. A feature-based approach to modeling protein-protein interaction hot spots. Nucleic Acids Res. 37, 2672–2687 (2009).
Perlman, E. J. et al. MLLT1 YEATS domain mutations in clinically distinctive favourable histology Wilms tumours. Nat. Commun. 6, 10013 (2015).
Suganuma, T. & Workman, J. L. Crosstalk among histone modifications. Cell 135, 604–607 (2008).
Lee, J. S., Smith, E. & Shilatifard, A. The language of histone crosstalk. Cell 142, 682–685 (2010).
Leach, B. I. et al. Leukemia fusion target AF9 is an intrinsically disordered transcriptional regulator that recruits multiple partners via coupled folding and binding. Structure 21, 176–183 (2013).
Kerry, J. et al. MLL-AF4 Spreading identifies binding sites that are distinct from super-enhancers and that govern sensitivity to DOT1L inhibition in leukemia. Cell Rep. 18, 482–495 (2017).
Kuntimaddi, A. et al. Degree of recruitment of DOT1L to MLL-AF9 defines level of H3K79 di- and tri-methylation on target genes and transformation potential. Cell Rep. 11, 808–820 (2015).
Gilan, O. et al. Functional interdependence of BRD4 and DOT1L in MLL leukemia. Nat. Struct. Mol. Biol. 23, 673–681 (2016).
Bruce, V. J. & McNaughton, B. R. Inside job: methods for delivering proteins to the interior of mammalian cells. Cell Chem. Biol. 24, 924–934 (2017).
Luo, Z., Lin, C. & Shilatifard, A. The super elongation complex (SEC) family in transcriptional control. Nat. Rev. Mol. Cell Biol. 13, 543–547 (2012).
He, N. et al. Human polymerase-associated factor complex (PAFc) connects the super elongation complex (SEC) to RNA polymerase II on chromatin. Proc. Natl. Acad. Sci. USA 108, E636–E645 (2011).
Gates, L. A. et al. Acetylation on histone H3 lysine 9 mediates a switch from transcription initiation to elongation. J. Biol. Chem. 292, 14456–14472 (2017).
Li, X. et al. Quantitative chemical proteomics approach to identify post-translational modification-mediated protein-protein interactions. J. Am. Chem. Soc. 134, 1982–1985 (2012).
Bao, X. et al. Identification of ‘erasers’ for lysine crotonylated histone marks using a chemical proteomics approach. eLife 3, e02999 (2014).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D. Biol. Crystallogr. 66, 22–25 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
Acknowledgements
We acknowledge support from the Hong Kong Research Grants Council Collaborative Research Fund (CRF C7029-15G to X.D.L.), the Areas of Excellence Scheme (AoE/P-705/16 to X.D.L.), the General Research Fund (GRF 17126618, 17125917 and 17303114 to X.D.L.), and the Early Career Scheme (ECS; HKU 709813P to X.D.L.). We acknowledge the National Natural Science Foundation of China (21572191 and 91753130 to X.D.L and 31725014 to H.L.), National Key R&D Program of China (2016YFA0500700 to H.L.), National Institutes of Health (1R01CA204639-01 to C.D.A.), the Leukemia and Lymphoma Society (LLS-SCOR 7006-13 to C.D.A), and funds from The Rockefeller University (to C.D.A.). Y.L. is a Tsinghua Advanced Fellow. L.W. is a fellow of the Jane Coffin Childs Memorial Fund. We acknowledge support from Beijing Metropolis for the Beijing Novo Program (Z181100006218068 to Y.L.) and China Association for Science and Technology for the Young Elite Scientists Sponsorship Program (to Y.L.). We thank the staff members at beamline BL17U1 the Shanghai Synchrotron Radiation Facility and S. Fan at Tsinghua Center for Structural Biology for their assistance in data collection and the China National Center for Protein Sciences Beijing for providing facility support. We thank H. Sun at Department of Chemistry, City University of Hong Kong for providing plasmid of the second BrD of BRD4. We thank A.Y.-H. Leung at Department of Medicine, the University of Hong Kong for providing the MV4;11 cell line.
Author information
Authors and Affiliations
Contributions
X.D.L. conceived the research project. X.L., X.-M.L., H.L., Y.L., and X.D.L. designed the experiments and analyzed the data. X.L., Y.J., Z.L., K.Y.F., and S.H.E.v.d.B. carried out the small-molecule and peptide synthesis. X.L., Y.C., Z.L., G.T., and Y.L. expressed and purified the proteins. X.L. performed the in vitro competition assay and ITC experiments. Y.L. and H.L. resolved the crystal structure and performed in silico modeling studies. X.-M.L. carried out the CETSA, ChIP-qPCR, and RT-qPCR experiments. L.W., C.D.A, and X.S. provided discussions and unpublished preliminary data. H.L. and X.D.L. supervised the work in their respective fields. X.L., Y.L. and X.D.L. wrote the manuscript with inputs from X.-M.L. and H.L.
Corresponding authors
Ethics declarations
Competing interests
X.L. and X.D.L. have filed a patent application (US Provisional Application No. 62/590,690) related to the peptide-based inhibitors reported in this manuscript.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–2, Supplementary Figures 1–12
Supplementary Note
Synthetic Procedures
Rights and permissions
About this article
Cite this article
Li, X., Li, XM., Jiang, Y. et al. Structure-guided development of YEATS domain inhibitors by targeting π-π-π stacking. Nat Chem Biol 14, 1140–1149 (2018). https://doi.org/10.1038/s41589-018-0144-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0144-y
This article is cited by
-
Histone crotonylation of peripheral blood mononuclear cells is a potential biomarker for diagnosis of colorectal cancer
Epigenetics & Chromatin (2023)
-
The ENL YEATS epigenetic reader domain critically links MLL-ENL to leukemic stem cell frequency in t(11;19) Leukemia
Leukemia (2023)
-
Targeting the histone H3 lysine 79 methyltransferase DOT1L in MLL-rearranged leukemias
Journal of Hematology & Oncology (2022)
-
Oncometabolites drive tumorigenesis by enhancing protein acylation: from chromosomal remodelling to nonhistone modification
Journal of Experimental & Clinical Cancer Research (2022)
-
Targeting transcription cycles in cancer
Nature Reviews Cancer (2022)