Abstract
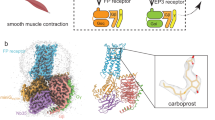
Prostaglandin E receptor EP4, a G-protein-coupled receptor, is involved in disorders such as cancer and autoimmune disease. Here, we report the crystal structure of human EP4 in complex with its antagonist ONO-AE3-208 and an inhibitory antibody at 3.2 Å resolution. The structure reveals that the extracellular surface is occluded by the extracellular loops and that the antagonist lies at the interface with the lipid bilayer, proximal to the highly conserved Arg316 residue in the seventh transmembrane domain. Functional and docking studies demonstrate that the natural agonist PGE2 binds in a similar manner. This structural information also provides insight into the ligand entry pathway from the membrane bilayer to the EP4 binding pocket. Furthermore, the structure reveals that the antibody allosterically affects the ligand binding of EP4. These results should facilitate the design of new therapeutic drugs targeting both orthosteric and allosteric sites in this receptor family.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The atomic coordinates and structure factor files for the EP4–Fab001, EP4–Fab001_Br, and Fab001 have been deposited in the Protein Data Bank with accession codes 5YWY, 5YHL, and 5YFI, respectively. The raw diffraction images have been deposited in Zenodo data repository (https://doi.org/10.5281/zenodo.1173791).
References
Hirata, T. & Narumiya, S. Prostanoid receptors. Chem. Rev. 111, 6209–6230 (2011).
Fujino, H. & Regan, J. W. EP(4) prostanoid receptor coupling to a pertussis toxin-sensitive inhibitory G protein. Mol. Pharmacol. 69, 5–10 (2006).
Buchanan, F. G. et al. Role of beta-arrestin 1 in the metastatic progression of colorectal cancer. Proc. Natl. Acad. Sci. USA 103, 1492–1497 (2006).
Yokoyama, U., Iwatsubo, K., Umemura, M., Fujita, T. & Ishikawa, Y. The prostanoid EP4 receptor and its signaling pathway. Pharmacol. Rev. 65, 1010–1052 (2013).
Yao, C. et al. Prostaglandin E2-EP4 signaling promotes immune inflammation through Th1 cell differentiation and Th17 cell expansion. Nat. Med. 15, 633–640 (2009).
Libioulle, C. et al. Novel Crohn disease locus identified by genome-wide association maps to a gene desert on 5p13.1 and modulates expression of PTGER4. PLoS. Genet. 3, e58 (2007).
Shi, Y. et al. A genome-wide association study identifies new susceptibility loci for non-cardia gastric cancer at 3q13.31 and 5p13.1. Nat. Genet. 43, 1215–1218 (2011).
Hinds, D. A. et al. A genome-wide association meta-analysis of self-reported allergy identifies shared and allergy-specific susceptibility loci. Nat. Genet. 45, 907–911 (2013).
Markovič, T., Jakopin, Ž., Dolenc, M. S. & Mlinarič-Raščan, I. Structural features of subtype-selective EP receptor modulators. Drug Discov. Today 22, 57–71 (2017).
Ward, C. L. et al. First clinical experience with ONO-4232: a randomized, double-blind, placebo-controlled healthy volunteer study of a novel lusitropic agent for acutely decompensated heart failure. Clin. Ther. 38, 1109–1121 (2016).
Watanabe, Y. et al. KAG-308, a newly-identified EP4-selective agonist shows efficacy for treating ulcerative colitis and can bring about lower risk of colorectal carcinogenesis by oral administration. Eur. J. Pharmacol. 754, 179–189 (2015).
Rausch-Derra, L., Huebner, M., Wofford, J. & Rhodes, L. A prospective, randomized, masked, placebo-controlled multisite clinical study of grapiprant, an EP4 prostaglandin receptor antagonist (PRA), in dogs with osteoarthritis. J. Vet. Intern. Med. 30, 756–763 (2016).
Bao, X. et al. Combination of EP4 antagonist and checkpoint inhibitors promotes anti-tumor effector T cells in preclinical tumor models. J. Immunother. Cancer 3, 350 (2015).
Kobayashi, T. et al. Identification of domains conferring ligand binding specificity to the prostanoid receptor. Studies on chimeric prostacyclin/prostaglandin D receptors. J. Biol. Chem. 272, 15154–15160 (1997).
Stillman, B. A., Audoly, L. & Breyer, R. M. A conserved threonine in the second extracellular loop of the human EP2 and EP4 receptors is required for ligand binding. Eur. J. Pharmacol. 357, 73–82 (1998).
Narumiya, S., Sugimoto, Y. & Ushikubi, F. Prostanoid receptors: structures, properties, and functions. Physiol. Rev. 79, 1193–1226 (1999).
Shiroishi, M. et al. Platform for the rapid construction and evaluation of GPCRs for crystallography in Saccharomyces cerevisiae. Microb. Cell. Fact. 11, 78 (2012).
Vaidehi, N., Grisshammer, R. & Tate, C. G. How can mutations thermostabilize G-protein-coupled receptors? Trends Pharmacol. Sci. 37, 37–46 (2016).
Yasuda, S. et al. Hot-spot residues to be mutated common in G protein-coupled receptors of class A: identification of thermostabilizing mutations followed by determination of three-dimensional structures for two example receptors. J. Phys. Chem. B 121, 6341–6350 (2017).
Ballesteros, J. A. & Weinstein, H. Integrated methods for the construction of three dimensional models and computational probing of structure-function relations in G protein-coupled receptors. Methods Neurosci. 25, 366–428 (1995).
Katritch, V. et al. Allosteric sodium in class A GPCR signaling. Trends. Biochem. Sci. 39, 233–244 (2014).
Takayama, K., Shimizu, T., Urushibata, Y. & Sugimoto, Y. Antibody against human prostaglandin E2 receptor EP4. US patent 20130197199 (2013).
Kabashima, K. et al. The prostaglandin receptor EP4 suppresses colitis, mucosal damage and CD4 cell activation in the gut. J. Clin. Invest. 109, 883–893 (2002).
Venkatakrishnan, A. J. et al. Molecular signatures of G-protein-coupled receptors. Nature 494, 185–194 (2013).
Hanson, M. A. et al. Crystal structure of a lipid G protein-coupled receptor. Science 335, 851–855 (2012).
Chrencik, J. E. et al. Crystal structure of antagonist bound human lysophosphatidic acid receptor 1. Cell 161, 1633–1643 (2015).
Shao, Z. et al. High-resolution crystal structure of the human CB1 cannabinoid receptor. Nature 540, 602–606 (2016).
Srivastava, A. et al. High-resolution structure of the human GPR40 receptor bound to allosteric agonist TAK-875. Nature 513, 124–127 (2014).
Palczewski, K. et al. Crystal structure of rhodopsin: a G protein-coupled receptor. Science 289, 739–745 (2000).
Kappel, K., Miao, Y. & McCammon, J. A. Accelerated molecular dynamics simulations of ligand binding to a muscarinic G-protein-coupled receptor. Q. Rev. Biophys. 48, 479–487 (2015).
Stanley, N., Pardo, L. & Fabritiis, G. D. The pathway of ligand entry from the membrane bilayer to a lipid G protein-coupled receptor. Sci. Rep. 6, 22639 (2016).
Mutoh, M. et al. Involvement of prostaglandin E receptor subtype EP4 in colon carcinogenesis. Cancer Res. 62, 28–32 (2002).
Kedzie, K. M., Donello, J. E., Krauss, H. A., Regan, J. W. & Gil, D. W. A single amino-acid substitution in the EP2 prostaglandin receptor confers responsiveness to prostacyclin analogs. Mol. Pharmacol. 54, 584–590 (1998).
Audoly, L. & Breyer, R. M. Substitution of charged amino acid residues in transmembrane regions 6 and 7 affect ligand binding and signal transduction of the prostaglandin EP3 receptor. Mol. Pharmacol. 51, 61–68 (1997).
Stitham, J., Stojanovic, A., Merenick, B. L., O’Hara, K. A. & Hwa, J. The unique ligand-binding pocket for the human prostacyclin receptor. Site-directed mutagenesis and molecular modeling. J. Biol. Chem. 278, 4250–4257 (2003).
Neuschäfer-Rube, F., Engemaier, E., Koch, S., Böer, U. & Püschel, G. P. Identification by site-directed mutagenesis of amino acids contributing to ligand-binding specificity or signal transduction properties of the human FP prostanoid receptor. Biochem. J. 371, 443–449 (2003).
Funk, C. D., Furci, L., Moran, N. & Fitzgerald, G. A. Point mutation in the seventh hydrophobic domain of the human thromboxane A2 receptor allows discrimination between agonist and antagonist binding sites. Mol. Pharmacol. 44, 934–939 (1993).
Natarajan, C., Hata, A. N., Hamm, H. E., Zent, R. & Breyer, R. M. Extracellular loop II modulates GTP sensitivity of the prostaglandin EP3 receptor. Mol. Pharmacol. 83, 206–216 (2013).
Margan, D., Borota, A., Mracec, M. & Mracec, M. 3D homology model of the human prostaglandin E2 receptor EP4 subtype. Rev. Roum. Chim. 57, 39–44 (2012).
Zare, B., Madadkar-Sobhani, A., Dastmalchi, S. & Mahmoudian, M. Prediction of the human EP1 receptor binding site by homology modeling and molecular dynamics simulation. Sci. Pharm. 79, 793–816 (2011).
Hutchings, C. J., Koglin, M., Olson, W. C. & Marshall, F. H. Opportunities for therapeutic antibodies directed at G-protein-coupled receptors. Nat. Rev. Drug. Discov. 16, 787–810 (2017).
Cheng, R. K. Y. et al. Structural insight into allosteric modulation of protease-activated receptor 2. Nature 545, 112–115 (2017).
Zhang, H. et al. Structure of the full-length glucagon class B G-protein-coupled receptor. Nature 546, 259–264 (2017).
Leduc, M. et al. Functional selectivity of natural and synthetic prostaglandin EP4 receptor ligands. J. Pharmacol. Exp. Ther. 331, 297–307 (2009).
Inoue, A. et al. TGFα shedding assay: an accurate and versatile method for detecting GPCR activation. Nat. Methods 9, 1021–1029 (2012).
Nomura, Y. et al. The intervening removable affinity tag (iRAT) production system facilitates Fv antibody fragment-mediated crystallography. Protein Sci. 25, 2268–2276 (2016).
Caffrey, M. & Cherezov, V. Crystallizing membrane proteins using lipidic mesophases. Nat. Protoc. 4, 706–731 (2009).
Ueno, G. et al. Remote access and automation of SPring-8 MX beamlines. AIP Conf. Proc. 1741, 050021 (2016).
Yamashita, K., Hirata, K. & Yamamoto, M. KAMO: towards automated data processing for microcrystals. Acta Crystallogr. D Struct. Biol. 74, 441–449 (2018).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Foadi, J. et al. Clustering procedures for the optimal selection of data sets from multiple crystals in macromolecular crystallography. Acta Crystallogr. D. Biol. Crystallogr. 69, 1617–1632 (2013).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Sheldrick, G. M. A short history of SHELX. Acta Crystallogr. A. 64, 112–122 (2008).
Thorn, A. & Sheldrick, G. M. ANODE: anomalous and heavy-atom density calculation. J. Appl. Crystallogr. 44, 1285–1287 (2011).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Halgren, T. A. Identifying and characterizing binding sites and assessing druggability. J. Chem. Inf. Model. 49, 377–389 (2009).
Sherman, W., Day, T., Jacobson, M. P., Friesner, R. A. & Farid, R. Novel procedure for modeling ligand/receptor induced fit effects. J. Med. Chem. 49, 534–553 (2006).
Harder, E. et al. OPLS3: a force field providing broad coverage of drug-like small molecules and proteins. J. Chem. Theory. Comput. 12, 281–296 (2016).
Case, D. A. et al. AMBER 11. (University of California, San Francisco, 2010).
Acknowledgements
We are grateful to Ono Pharmaceutical Company for supplying EP4 antagonists; to the beamline scientists at BL32XU and BL41XU of SPring-8 (Hyogo, Japan) for their technical assistance during data collection; to A. Inoue at Tohoku University for the TGF-α shedding assay; to B.K. Kobilka (Tsinghua University and Stanford University), W. Shihoya and R. Taniguchi (The University of Tokyo) for their useful comments; and to H. Tsujimoto, M. Sasanuma and members of the Iwata lab at Kyoto University for technical assistance. DNA sequencing analysis was performed at the Medical Research Support Center, Graduate School of Medicine, Kyoto University. This work was supported by the Strategic Basic Research Program, JST (S.I.); the Toray Science Foundation (T.K.); the Takeda Science Foundation (T.K. and R.S.); the Naito Foundation (T.K.); Koyanagi Foundation (T.K.); the Platform for Drug Discovery, Informatics, and Structural Life Science (PDIS) funded by the Ministry of Education, Culture, Sports, Science and Technology of Japan (MEXT) and the Japan Agency for Medical Research and Development (AMED) (T.K., T.M., M. Shiroishi, T.H. and M.Y.); Core Research for Evolutional Science and Technology (CREST) funded by AMED (Y. Su., S.N. and T.K.); the ImPACT Program of the Council for Science, Technology and Innovation (Cabinet Office, Government of Japan; T.M. and M.K.); MEXT as a “Priority Issue on Post-K computer” (Building Innovative Drug Discovery Infrastructure Through Functional Control of Biomolecular Systems) (hp160213) (T. Hi.); and the Japan Society for the Promotion of Science (JSPS) KAKENHI (Grant Nos. 15K08268 to R.S., 15J00102 to K.M., 15J04343 to S.H., 15H06862 to K.Y., and 15H05905 to Y. Su.). K.M. and S.H. are recipients of JSPS postdoctoral fellowships. X-ray crystallographic data were collected at SPring-8 (Proposal Nos. 2013A1379, 2013B1184, 2013B1092, 2014A1301, 2014B1355, 2014B1273, 2015A1080, 2015A1044, 2015B1092, 2015B2044, and 2015B2080).
Author information
Authors and Affiliations
Contributions
T.K., Y.T., S.I., and S.N. designed the project. Initial trials of EP4 were conducted by Y.T., H.A., and T. Nakane. Purification and crystallization of EP4 and EP4–Fab001 were performed by Y.T. and Y. Sekiguchi. The thermostabilizing mutation Gly1063.39Arg was discovered by S. Yasuda, Y.K., T.M., and M.K. using a theoretical strategy developed by M.K., S. Yasuda, and T.M. The alanine-scanning mutations were designed by T.K., Y.T., K.M., and T. Nakane. The construction and binding assays of EP4 mutants were performed by K.M. and Y.T. FSEC-TS was performed by K.M., Y.T. and Y.H. TGF-α shedding assays were performed by K.M. ITC experiments were performed by M. Shiroishi. The generation, expression, purification, and evaluation of the antibody were performed by K. Ta., Y. Su., T.S., Y.U., T.I., and K. Tsu. Fab and Fv fragments were prepared by Y.T., N.N., Y. Sekiguchi, Y.H., and Y. Shiimura. The synthesis of CHEMBL1644016 was performed by S. Yoshida, T. Ku., and T.H. The data collection was performed by Y.T., R.S., K.Y., and K.H., and supervised by S.I. and M.Y. Structure determination and refinement were performed by S.H., R.S., K.Y., K.H., Y.T., T. Nakane, and T. Nakagita. The molecular dynamics simulations and computational modeling were performed by T. Hi and M. Sato. The manuscript was prepared by Y.T., T.K., S.I., K.M., S.H., and R.S., and all authors discussed the results and commented on the manuscript. The research was supervised by T.K., R.S., S.I., and S.N.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11, Supplementary Tables 1–2
Supplementary Video 1
Molecular dynamics simulations of ligand access of EP4
Rights and permissions
About this article
Cite this article
Toyoda, Y., Morimoto, K., Suno, R. et al. Ligand binding to human prostaglandin E receptor EP4 at the lipid-bilayer interface. Nat Chem Biol 15, 18–26 (2019). https://doi.org/10.1038/s41589-018-0131-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0131-3
This article is cited by
-
Docking for EP4R antagonists active against inflammatory pain
Nature Communications (2023)
-
Structural basis of α1A-adrenergic receptor activation and recognition by an extracellular nanobody
Nature Communications (2023)
-
Structures of human prostaglandin F2α receptor reveal the mechanism of ligand and G protein selectivity
Nature Communications (2023)
-
Structural basis of antibody inhibition and chemokine activation of the human CC chemokine receptor 8
Nature Communications (2023)
-
Fly casting with ligand sliding and orientational selection supporting complex formation of a GPCR and a middle sized flexible molecule
Scientific Reports (2022)