Abstract
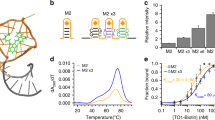
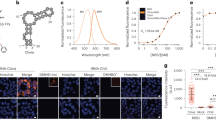
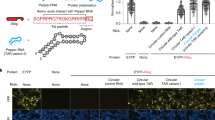
RNAs directly regulate a vast array of cellular processes, emphasizing the need for robust approaches to fluorescently label and track RNAs in living cells. Here, we develop an RNA imaging platform using the cobalamin riboswitch as an RNA tag and a series of probes containing cobalamin as a fluorescence quencher. This highly modular ‘Riboglow’ platform leverages different colored fluorescent dyes, linkers and riboswitch RNA tags to elicit fluorescence turn-on upon binding RNA. We demonstrate the ability of two different Riboglow probes to track mRNA and small noncoding RNA in live mammalian cells. A side-by-side comparison revealed that Riboglow outperformed the dye-binding aptamer Broccoli and performed on par with the gold standard RNA imaging system, the MS2-fluorescent protein system, while featuring a much smaller RNA tag. Together, the versatility of the Riboglow platform and ability to track diverse RNAs suggest broad applicability for a variety of imaging approaches.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Change history
05 August 2019
In the version of this article originally published, numbered compounds were not linked correctly to their respective compound pages. The error has been corrected in the HTML version of this paper.
References
Gerstberger, S., Hafner, M. & Tuschl, T. A census of human RNA-binding proteins. Nat. Rev. Genet. 15, 829–845 (2014).
Buchan, J. R. & Parker, R. Eukaryotic stress granules: the ins and outs of translation. Mol. Cell 36, 932–941 (2009).
Decker, C. J. & Parker, R. P-bodies and stress granules: possible roles in the control of translation and mRNA degradation. Cold Spring Harb. Perspect. Biol. 4, a012286 (2012).
Matera, A. G. & Wang, Z. A day in the life of the spliceosome. Nat. Rev. Mol. Cell Biol. 15, 108–121 (2014).
Tsalikis, J. et al. Intracellular bacterial pathogens trigger the formation of U small nuclear RNA bodies (U bodies) through metabolic stress induction. J. Biol. Chem. 290, 20904–20918 (2015).
Fusco, D. et al. Single mRNA molecules demonstrate probabilistic movement in living mammalian cells. Curr. Biol. 13, 161–167 (2003).
Wu, B., Chen, J. & Singer, R. H. Background free imaging of single mRNAs in live cells using split fluorescent proteins. Sci. Rep. 4, 3615 (2014).
Tutucci, E. et al. An improved MS2 system for accurate reporting of the mRNA life cycle. Nat. Methods 15, 81–89 (2018).
Garcia, J. F. & Parker, R. MS2 coat proteins bound to yeast mRNAs block 5′ to 3′ degradation and trap mRNA decay products: implications for the localization of mRNAs by MS2-MCP system. RNA 21, 1393–1395 (2015).
Wu, B., Eliscovich, C., Yoon, Y. J. & Singer, R. H. Translation dynamics of single mRNAs in live cells and neurons. Science 352, 1430–1435 (2016).
Katz, Z. B. et al. Mapping translation ‘hot-spots’ in live cells by tracking single molecules of mRNA and ribosomes. eLife 5, 1–17 (2016).
Nguyen, D. H., DeFina, S. C., Fink, W. H. & Dieckmann, T. Binding to an RNA aptamer changes the charge distribution and conformation of malachite green. J. Am. Chem. Soc. 124, 15081–15084 (2002).
Babendure, J. R., Adams, S. R. & Tsien, R. Y. Aptamers switch on fluorescence of triphenylmethane dyes. J. Am. Chem. Soc. 125, 14716–14717 (2003).
Arora, A., Sunbul, M. & Jäschke, A. Dual-colour imaging of RNAs using quencher- and fluorophore-binding aptamers. Nucleic Acids Res. 43, e144 (2015).
Sunbul, M. & Jäschke, A. Contact-mediated quenching for RNA imaging in bacteria with a fluorophore-binding aptamer. Angew. Chem. Int. Edn. Engl. 52, 13401–13404 (2013).
Tan, X. et al. Fluoromodules consisting of a promiscuous RNA aptamer and red or blue fluorogenic cyanine dyes: selection, characterization, and bioimaging. J. Am. Chem. Soc. 139, 9001–9009 (2017).
Paige, J. S., Wu, K. Y. & Jaffrey, S. R. RNA mimics of green fluorescent protein. Science 333, 642–646 (2011).
Filonov, G. S., Moon, J. D., Svensen, N. & Jaffrey, S. R. Broccoli: rapid selection of an RNA mimic of green fluorescent protein by fluorescence-based selection and directed evolution. J. Am. Chem. Soc. 136, 16299–16308 (2014).
Dolgosheina, E. V. et al. RNA mango aptamer-fluorophore: a bright, high-affinity complex for RNA labeling and tracking. ACS Chem. Biol. 9, 2412–2420 (2014).
Autour, A. et al. Fluorogenic RNA Mango aptamers for imaging small non-coding RNAs in mammalian cells. Nat. Commun. 9, 656 (2018).
Filonov, G. S. & Jaffrey, S. R. RNA imaging with dimeric Broccoli in live bacterial and mammalian cells. Curr. Protoc. Chem. Biol. 8, 1–28 (2016).
Song, W. et al. Imaging RNA polymerase III transcription using a photostable RNA-fluorophore complex. Nat. Chem. Biol. 13, 1187–1194 (2017).
Ceres, P., Trausch, J. J. & Batey, R. T. Engineering modular ‘ON’ RNA switches using biological components. Nucleic Acids Res. 41, 10449–10461 (2013).
Ceres, P., Garst, A. D., Marcano-Velázquez, J. G. & Batey, R. T. Modularity of select riboswitch expression platforms enables facile engineering of novel genetic regulatory devices. ACS Synth. Biol. 2, 463–472 (2013).
Johnson, J. E. Jr., Reyes, F. E., Polaski, J. T. & Batey, R. T. B12 cofactors directly stabilize an mRNA regulatory switch. Nature 492, 133–137 (2012).
Lee, M. & Grissom, C. B. Design, synthesis, and characterization of fluorescent cobalamin analogues with high quantum efficiencies. Org. Lett. 11, 2499–2502 (2009).
Smeltzer, C. C. et al. Synthesis and characterization of fluorescent cobalamin (CobalaFluor) derivatives for imaging. Org. Lett. 3, 799–801 (2001).
Fedosov, S. N. et al. Application of a fluorescent cobalamin analogue for analysis of the binding kinetics. A study employing recombinant human transcobalamin and intrinsic factor. FEBS J. 273, 4742–4753 (2006).
Chromiński, M. & Gryko, D. “Clickable” vitamin B12 derivative. Chemistry 19, 5141–5148 (2013).
Loska, R., Janiga, A. & Gryko, D. Design and synthesis of protoporphyrin IX/vitamin B-12 molecular hybrids via CuAAC reaction. J. Porphyr. phtalocyanines 17, 104–117 (2013).
Trachman, R. J. III et al. Structural basis for high-affinity fluorophore binding and activation by RNA Mango. Nat. Chem. Biol. 13, 807–813 (2017).
Warner, K. D. et al. Structural basis for activity of highly efficient RNA mimics of green fluorescent protein. Nat. Struct. Mol. Biol. 21, 658–663 (2014).
Jeng, S. C. Y., Chan, H. H. Y., Booy, E. P., McKenna, S. A. & Unrau, P. J. Fluorophore ligand binding and complex stabilization of the RNA Mango and RNA Spinach aptamers. RNA 22, 1884–1892 (2016).
Filonov, G. S., Kam, C. W., Song, W. & Jaffrey, S. R. In-gel imaging of RNA processing using broccoli reveals optimal aptamer expression strategies. Chem. Biol. 22, 649–660 (2015).
Ponchon, L. & Dardel, F. Recombinant RNA technology: the tRNA scaffold. Nat. Methods 4, 571–576 (2007).
Morisaki, T. et al. Real-time quantification of single RNA translation dynamics in living cells. Science 352, 1425–1429 (2016).
Hayashi-Takanaka, Y. et al. Tracking epigenetic histone modifications in single cells using Fab-based live endogenous modification labeling. Nucleic Acids Res. 39, 6475–6488 (2011).
McNeil, P. L. & Warder, E. Glass beads load macromolecules into living cells. J. Cell Sci. 88, 669–678 (1987).
Nelles, D. A. et al. Programmable RNA tracking in live cells with CRISPR/Cas9. Cell 165, 488–496 (2016).
Zurla, C., Lifland, A. W. & Santangelo, P. J. Characterizing mRNA interactions with RNA granules during translation initiation inhibition. PLoS One 6, e19727 (2011).
Grimm, J. B. et al. A general method to fine-tune fluorophores for live-cell and in vivo imaging. Nat. Methods 14, 987–994 (2017).
Kedersha, N., Tisdale, S., Hickman, T. & Anderson, P. Real-time and quantitative imaging of mammalian stress granules and processing bodies. Methods Enzymol. 448, 521–552 (2008).
Wu, B., Chao, J. A. & Singer, R. H. Fluorescence fluctuation spectroscopy enables quantitative imaging of single mRNAs in living cells. Biophys. J. 102, 2936–2944 (2012).
Ni, C. Z. et al. Crystal structure of the MS2 coat protein dimer: implications for RNA binding and virus assembly. Structure 3, 255–263 (1995).
Han, K. Y., Leslie, B. J., Fei, J., Zhang, J. & Ha, T. Understanding the photophysics of the spinach-DFHBI RNA aptamer-fluorogen complex to improve live-cell RNA imaging. J. Am. Chem. Soc. 135, 19033–19038 (2013).
McCloskey, A., Taniguchi, I., Shinmyozu, K. & Ohno, M. hnRNP C tetramer measures RNA length to classify RNA polymerase II transcripts for export. Science 335, 1643–1646 (2012).
Ishikawa, H. et al. Identification of truncated forms of U1 snRNA reveals a novel RNA degradation pathway during snRNP biogenesis. Nucleic Acids Res. 42, 2708–2724 (2014).
Hutten, S., Chachami, G., Winter, U., Melchior, F. & Lamond, A. I. A role for the Cajal-body-associated SUMO isopeptidase USPL1 in snRNA transcription mediated by RNA polymerase II. J. Cell Sci. 127, 1065–1078 (2014).
Nahvi, A., Barrick, J. E. & Breaker, R. R. Coenzyme B12 riboswitches are widespread genetic control elements in prokaryotes. Nucleic Acids Res. 32, 143–150 (2004).
Quadros, E. V. & Sequeira, J. M. Cellular uptake of cobalamin: transcobalamin and the TCblR/CD320 receptor. Biochimie 95, 1008–1018 (2013).
Edwards, A. L., Garst, A. D. & Batey, R. T. Determining structures of RNA aptamers and riboswitches by X-ray crystallography. Methods Mol. Biol. 535, 135–163 (2009).
Polaski, J. T., Holmstrom, E. D., Nesbitt, D. J. & Batey, R. T. Mechanistic insights into cofactor-dependent coupling of RNA folding and mRNA transcription/translation by a cobalamin riboswitch. Cell Rep. 15, 1100–1110 (2016).
Gilbert, S. D. & Batey, R. T. Monitoring RNA-ligand interactions using isothermal titration calorimetry. Methods Mol. Biol. 540, 97–114 (2009).
Lakowicz, J. R. Quenching of Fluorescence. in Principles of Fluorescence Spectroscopy 277–330 (Springer, 2006).
Beckley, S. A. et al. Reduction of target gene expression by a modified U1 snRNA. Mol. Cell. Biol. 21, 2815–2825 (2001).
Wheeler, J. R., Matheny, T., Jain, S., Abrisch, R. & Parker, R. Distinct stages in stress granule assembly and disassembly. eLife 5, 1–25 (2016).
Xi, L., Schmidt, J. C., Zaug, A. J., Ascarrunz, D. R. & Cech, T. R. A novel two-step genome editing strategy with CRISPR-Cas9 provides new insights into telomerase action and TERT gene expression. Genome Biol. 16, 231 (2015).
Shpargel, K. B. & Matera, A. G. Gemin proteins are required for efficient assembly of Sm-class ribonucleoproteins. Proc. Natl Acad. Sci. USA 102, 17372–17377 (2005).
Shpargel, K. B., Ospina, J. K., Tucker, K. E., Matera, A. G. & Hebert, M. D. Control of Cajal body number is mediated by the coilin C-terminus. J. Cell Sci. 116, 303–312 (2003).
Acknowledgements
The authors would like to acknowledge financial support from the Human Frontiers Science Project and NIH Director’s Pioneer Award GM114863 (to A.E.P.). We acknowledge support from the National Science Centre, SYMFONIA DEC-2014/12/W/ST5/00589 to D. Gryko and A.J.W., the National Institutes of Health (5R01 GM073850) to R.T.B., and NSF Physics Frontier Center at JILA (PHY1734006) to R.J.. We thank L. Lavis (Janelia Research Campus) for providing fluorophores JF585, SiR594 and JF646 for Halo staining, S. Jaffrey (Cornell University) and G. Matera (UNC Chapel Hill) for contributing plasmids and S. Shukla and J. Garcia for helpful discussions; T. Stasevich, D. Muhlrad, J. Lee and M. Lo for technical expertise; and to J. Eberhard and J. Gassensmith for helpful discussions. The imaging work was performed at the BioFrontiers Institute Advanced Light Microscopy Core, whose Nikon A1R microscope was acquired by the generous support of the NIST-CU Cooperative Agreement award number 70NANB15H226. R.J is a staff member in the Quantum Physics Division of the National Institute of Standards and Technology (NIST). Certain commercial equipment, instruments, or materials are identified in this paper in order to specify the experimental procedure adequately. Such identification is not intended to imply recommendation or endorsement by the NIST, nor is it intended to imply that the materials or equipment identified are necessarily the best available for the purpose.
Author information
Authors and Affiliations
Contributions
E.B., J.T.P., R.T.B. and A.E.P. conceptualized and designed the study. J.T.P. and R.T.B. rationally designed riboswitch variants. E.B., J.T.P., R.T.B., A.J.W., D.G. and A.E.P. designed organic probes. A.J.W. and M.C. synthesized organic probes. J.T.P. and Z.E.H. purified riboswitch variants for in vitro work. E.B. performed in vitro work, designed and performed cellular work, and analyzed data with input from all authors. D.B. constructed plasmids and assisted with cellular work. S.-T.H. performed in vitro fluorescence lifetime and bleaching experiments. J.R.W. made the Halo-G3BP1 U2-OS cell line. R.P. and R.J. provided critical advice. E.B. and A.E.P. wrote the manuscript with edits from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–13, Supplementary Figures 1–33
Supplementary Note 1
Synthetic Procedures
Rights and permissions
About this article
Cite this article
Braselmann, E., Wierzba, A.J., Polaski, J.T. et al. A multicolor riboswitch-based platform for imaging of RNA in live mammalian cells. Nat Chem Biol 14, 964–971 (2018). https://doi.org/10.1038/s41589-018-0103-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0103-7
This article is cited by
-
Large Stokes shift fluorescent RNAs for dual-emission fluorescence and bioluminescence imaging in live cells
Nature Methods (2023)
-
Visualizing orthogonal RNAs simultaneously in live mammalian cells by fluorescence lifetime imaging microscopy (FLIM)
Nature Communications (2023)
-
Avidity-based bright and photostable light-up aptamers for single-molecule mRNA imaging
Nature Chemical Biology (2023)
-
Intracellular RNA and DNA tracking by uridine-rich internal loop tagging with fluorogenic bPNA
Nature Communications (2023)
-
Fast-exchanging spirocyclic rhodamine probes for aptamer-based super-resolution RNA imaging
Nature Communications (2023)