Abstract
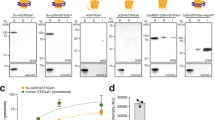
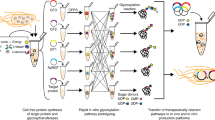
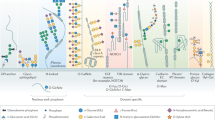
Glycosylation is an abundant post-translational modification that is important in disease and biotechnology. Current methods to understand and engineer glycosylation cannot sufficiently explore the vast experimental landscapes required to accurately predict and design glycosylation sites modified by glycosyltransferases. Here we describe a systematic platform for glycosylation sequence characterization and optimization by rapid expression and screening (GlycoSCORES), which combines cell-free protein synthesis and mass spectrometry of self-assembled monolayers. We produced six N- and O-linked polypeptide-modifying glycosyltransferases from bacteria and humans in vitro and rigorously determined their substrate specificities using 3,480 unique peptides and 13,903 unique reaction conditions. We then used GlycoSCORES to optimize and design small glycosylation sequence motifs that directed efficient N-linked glycosylation in vitro and in the Escherichia coli cytoplasm for three heterologous proteins, including the human immunoglobulin Fc domain. We find that GlycoSCORES is a broadly applicable method to facilitate fundamental understanding of glycosyltransferases and engineer synthetic glycoproteins.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Khoury, G. A., Baliban, R. C. & Floudas, C. A. Proteome-wide post-translational modification statistics: frequency analysis and curation of the swiss-prot database. Sci. Rep. 1, 90 (2011).
Helenius, A. & Aebi, M. Intracellular functions of N-linked glycans. Science 291, 2364–2369 (2001).
Sethuraman, N. & Stadheim, T. A. Challenges in therapeutic glycoprotein production. Curr. Opin. Biotechnol. 17, 341–346 (2006).
Elliott, S. et al. Enhancement of therapeutic protein in vivo activities through glycoengineering. Nat. Biotechnol. 21, 414–421 (2003).
Chung, C. H. et al. Cetuximab-induced anaphylaxis and IgE specific for galactose-α-1,3-galactose. N. Engl. J. Med. 358, 1109–1117 (2008).
Lin, C.-W. et al. A common glycan structure on immunoglobulin G for enhancement of effector functions. Proc. Natl Acad. Sci. USA 112, 10611–10616 (2015).
Clausen, H., Wandall, H.H., Steentoft, C., Stanley, P. & Schnaar, R.L. in Essentials of Glycobiology. (eds. A. Varki et al.) 713–728 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, 2015).
Valderrama-Rincon, J. D. et al. An engineered eukaryotic protein glycosylation pathway in Escherichia coli. Nat. Chem. Biol. 8, 434–436 (2012).
Keys, T. G. & Aebi, M. Engineering protein glycosylation in prokaryotes. Curr. Opin. Syst. Biol. 5, 23–31 (2017).
Wang, L.-X. & Davis, B. G. Realizing the promise of chemical glycobiology. Chem. Sci. 4, 3381–3394 (2013).
Yang, Z. et al. Engineered CHO cells for production of diverse, homogeneous glycoproteins. Nat. Biotechnol. 33, 842–844 (2015).
Li, H. et al. Optimization of humanized IgGs in glycoengineered Pichia pastoris. Nat. Biotechnol. 24, 210–215 (2006).
Xu, Y. et al. A novel enzymatic method for synthesis of glycopeptides carrying natural eukaryotic N-glycans. Chem. Commun. (Camb.) 53, 9075–9077 (2017).
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P. M. & Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42, D490–D495 (2014).
Ban, L. et al. Discovery of glycosyltransferases using carbohydrate arrays and mass spectrometry. Nat. Chem. Biol. 8, 769–773 (2012).
Pathak, S. et al. The active site of O-GlcNAc transferase imposes constraints on substrate sequence. Nat. Struct. Mol. Biol. 22, 744–750 (2015).
Ortiz-Meoz, R. F., Merbl, Y., Kirschner, M. W. & Walker, S. Microarray discovery of new OGT substrates: the medulloblastoma oncogene OTX2 is O-GlcNAcylated. J. Am. Chem. Soc. 136, 4845–4848 (2014).
Robinson, P. V., Tsai, C. T., de Groot, A. E., McKechnie, J. L. & Bertozzi, C. R. Glyco-seek: ultrasensitive detection of protein-specific glycosylation by proximity ligation polymerase chain reaction. J. Am. Chem. Soc. 138, 10722–10725 (2016).
Naegeli, A. et al. Substrate specificity of cytoplasmic N-glycosyltransferase. J. Biol. Chem. 289, 24521–24532 (2014).
Naegeli, A. et al. Molecular analysis of an alternative N-glycosylation machinery by functional transfer from Actinobacillus pleuropneumoniae to Escherichia coli. J. Biol. Chem. 289, 2170–2179 (2014).
Keys, T. G. et al. A biosynthetic route for polysialylating proteins in Escherichia coli. Metab. Eng. 44, 293–301 (2017).
Cuccui, J. et al. The N-linking glycosylation system from Actinobacillus pleuropneumoniae is required for adhesion and has potential use in glycoengineering. Open Biol. 7, 160212 (2017).
Schwarz, F., Fan, Y. Y., Schubert, M. & Aebi, M. Cytoplasmic N-glycosyltransferase of Actinobacillus pleuropneumoniae is an inverting enzyme and recognizes the NX(S/T) consensus sequence. J. Biol. Chem. 286, 35267–35274 (2011).
Song, Q. et al. Production of homogeneous glycoprotein with multi-site modifications by an engineered N-glycosyltransferase mutant. J. Biol. Chem. 292, 8856–8863 (2017).
Gross, J. et al. The Haemophilus influenzae HMW1 adhesin is a glycoprotein with an unusual N-linked carbohydrate modification. J. Biol. Chem. 283, 26010–26015 (2008).
Kawai, F. et al. Structural insights into the glycosyltransferase activity of the Actinobacillus pleuropneumoniae HMW1C-like protein. J. Biol. Chem. 286, 38546–38557 (2011).
Lomino, J. V. et al. A two-step enzymatic glycosylation of polypeptides with complex N-glycans. Bioorg. Med. Chem. 21, 2262–2270 (2013).
Chen, M. M., Glover, K. J. & Imperiali, B. From peptide to protein: comparative analysis of the substrate specificity of N-linked glycosylation in C. jejuni. Biochemistry 46, 5579–5585 (2007).
Fisher, A. C. et al. Production of secretory and extracellular N-linked glycoproteins in Escherichia coli. Appl. Environ. Microbiol. 77, 871–881 (2011).
Carlson, E. D., Gan, R., Hodgman, C. E. & Jewett, M. C. Cell-free protein synthesis: applications come of age. Biotechnol. Adv. 30, 1185–1194 (2012).
Kuo, H. Y., DeLuca, T. A., Miller, W. M. & Mrksich, M. Profiling deacetylase activities in cell lysates with peptide arrays and SAMDI mass spectrometry. Anal. Chem. 85, 10635–10642 (2013).
Kornacki, J. R., Stuparu, A. D. & Mrksich, M. Acetyltransferase p300/CBP associated Factor (PCAF) regulates crosstalk-dependent acetylation of histone H3 by distal site recognition. ACS Chem. Biol. 10, 157–164 (2015).
Kim, J. & Mrksich, M. Profiling the selectivity of DNA ligases in an array format with mass spectrometry. Nucleic Acids Res. 38, e2 (2010).
Laurent, N. et al. Enzymatic glycosylation of peptide arrays on gold surfaces. ChemBioChem 9, 883–887 (2008).
Laurent, N. et al. SPOT synthesis of peptide arrays on self-assembled monolayers and their evaluation as enzyme substrates. ChemBioChem 9, 2592–2596 (2008).
Hussain, M. R., Hoessli, D. C. & Fang, M. N-acetylgalactosaminyltransferases in cancer. Oncotarget 7, 54067–54081 (2016).
Schjoldager, K. T. et al. Probing isoform-specific functions of polypeptide GalNAc-transferases using zinc finger nuclease glycoengineered SimpleCells. Proc. Natl Acad. Sci. USA 109, 9893–9898 (2012).
Yoshida, A., Suzuki, M., Ikenaga, H. & Takeuchi, M. Discovery of the shortest sequence motif for high level mucin-type O-glycosylation. J. Biol. Chem. 272, 16884–16888 (1997).
Gerken, T. A., Raman, J., Fritz, T. A. & Jamison, O. Identification of common and unique peptide substrate preferences for the UDP-GalNAc:polypeptide α-N-acetylgalactosaminyltransferases T1 and T2 derived from oriented random peptide substrates. J. Biol. Chem. 281, 32403–32416 (2006).
Kong, Y. et al. Probing polypeptide GalNAc-transferase isoform substrate specificities by in vitro analysis. Glycobiology 25, 55–65 (2015).
Steentoft, C. et al. Precision mapping of the human O-GalNAc glycoproteome through SimpleCell technology. EMBO J. 32, 1478–1488 (2013).
Wang, A. C., Jensen, E. H., Rexach, J. E., Vinters, H. V. & Hsieh-Wilson, L. C. Loss of O-GlcNAc glycosylation in forebrain excitatory neurons induces neurodegeneration. Proc. Natl Acad. Sci. USA 113, 15120–15125 (2016).
Yang, X. et al. Phosphoinositide signalling links O-GlcNAc transferase to insulin resistance. Nature 451, 964–969 (2008).
Liu, X. et al. A peptide panel investigation reveals the acceptor specificity of O-GlcNAc transferase. FASEB J. 28, 3362–3372 (2014).
Chalkley, R. J., Thalhammer, A., Schoepfer, R. & Burlingame, A. L. Identification of protein O-GlcNAcylation sites using electron transfer dissociation mass spectrometry on native peptides. Proc. Natl Acad. Sci. USA 106, 8894–8899 (2009).
Lazarus, M. B., Nam, Y., Jiang, J., Sliz, P. & Walker, S. Structure of human O-GlcNAc transferase and its complex with a peptide substrate. Nature 469, 564–567 (2011).
Choi, K. J., Grass, S., Paek, S., St Geme, J. W. III & Yeo, H. J. The Actinobacillus pleuropneumoniae HMW1C-like glycosyltransferase mediates N-linked glycosylation of the Haemophilus influenzae HMW1 adhesin. PLoS One 5, e15888 (2010).
Haselberg, R., de Jong, G. J. & Somsen, G. W. Low-flow sheathless capillary electrophoresis-mass spectrometry for sensitive glycoform profiling of intact pharmaceutical proteins. Anal. Chem. 85, 2289–2296 (2013).
Schoborg, J. A. et al. A cell-free platform for rapid synthesis and testing of active oligosaccharyltransferases. Biotechnol. Bioeng. 115, 739–750 (2018).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Gurard-Levin, Z. A., Scholle, M. D., Eisenberg, A. H. & Mrksich, M. High-throughput screening of small molecule libraries using SAMDI mass spectrometry. ACS Comb. Sci. 13, 347–350 (2011).
Goerke, A. R. & Swartz, J. R. Development of cell-free protein synthesis platforms for disulfide bonded proteins. Biotechnol. Bioeng. 99, 351–367 (2008).
Espah Borujeni, A., Channarasappa, A. S. & Salis, H. M. Translation rate is controlled by coupled trade-offs between site accessibility, selective RNA unfolding and sliding at upstream standby sites. Nucleic Acids Res. 42, 2646–2659 (2014).
Martin, R. W. et al. Cell-free protein synthesis from genomically recoded bacteria enables multisite incorporation of noncanonical amino acids. Nat. Commun. 9, 1203 (2018).
Lajoie, M. J. et al. Genomically recoded organisms expand biological functions. Science 342, 357–360 (2013).
Kwon, Y.-C. & Jewett, M. C. High-throughput preparation methods of crude extract for robust cell-free protein synthesis. Sci. Rep. 5, 8663 (2015).
Jewett, M. C. & Swartz, J. R. Mimicking the Escherichia coli cytoplasmic environment activates long-lived and efficient cell-free protein synthesis. Biotechnol. Bioeng. 86, 19–26 (2004).
Jewett, M. C., Calhoun, K. A., Voloshin, A., Wuu, J. J. & Swartz, J. R. An integrated cell-free metabolic platform for protein production and synthetic biology. Mol. Syst. Biol. 4, 220 (2008).
Jewett, M. C. & Swartz, J. R. Rapid expression and purification of 100 nmol quantities of active protein using cell-free protein synthesis. Biotechnol. Prog. 20, 102–109 (2004).
Hong, S. H. et al. Cell-free protein synthesis from a release factor 1 deficient Escherichia coli activates efficient and multiple site-specific nonstandard amino acid incorporation. ACS Synth. Biol. 3, 398–409 (2014).
Jian, W., Edom, R. W., Wang, D., Weng, N. & Zhang, S. W. Relative quantitation of glycoisoforms of intact apolipoprotein C3 in human plasma by liquid chromatography-high-resolution mass spectrometry. Anal. Chem. 85, 2867–2874 (2013).
Acknowledgements
The authors acknowledge J.C. Stark and J. Hershewe for assistance with western blotting, helpful discussions, and sharing of reagents and ideas; S. Habibi for assistance with LC-TOF instrumentation; and A. Karim for helpful conversations. The authors also thank J. Kath for supply of plasmids, advice on protein expression, and critical reading of the manuscript. We also thank A. Natarajan of the Department of Microbiology at Cornell University, T. Jaroentomeechai of the Robert Frederick Smith School of Chemical and Biomolecular Engineering at Cornell University, and J. Janetzko of the Department of Chemistry and Chemical Biology at Harvard University for sharing the ppGalNAcT, Im7, and hOGT source plasmids, respectively. This work made use of the Integrated Molecular Structure Education and Research Center at Northwestern University, which has received support from the state of Illinois, the Northwestern University Office of Research and the Chemistry Department for LC-TOF instrumentation. This material is based upon work supported by the Defense Threat Reduction Agency (HDTRA1-15-10052/P00001), the David and Lucile Packard Foundation, the Dreyfus Teacher-Scholar program, and the National Science Foundation (Graduate Research Fellowship under Grant No. DGE-1324585 and MCB-1413563).
Author information
Authors and Affiliations
Contributions
W.K. and L.L. designed, performed, and analyzed experiments. M.R. designed and optimized experimental protocols. W.L. helped to synthesize peptide libraries. M.M. and M.C.J. directed the studies and interpreted the data. W.K., L.L., M.P.D., M.M., and M.C.J. conceived of the study and wrote the manuscript with assistance from M.R. and W.L.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–5, Supplementary Figures 1–27, Supplementary Note 1
Rights and permissions
About this article
Cite this article
Kightlinger, W., Lin, L., Rosztoczy, M. et al. Design of glycosylation sites by rapid synthesis and analysis of glycosyltransferases. Nat Chem Biol 14, 627–635 (2018). https://doi.org/10.1038/s41589-018-0051-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0051-2
This article is cited by
-
Exposing the molecular heterogeneity of glycosylated biotherapeutics
Nature Communications (2024)
-
Cell-free systems for a multi-pronged approach to next-generation therapeutics and diagnostics
Biotechnology and Bioprocess Engineering (2024)
-
Recent Advances in Rapid Screening Methods for Glycosyltransferases
Catalysis Letters (2024)
-
Molecular basis for bacterial N-glycosylation by a soluble HMW1C-like N-glycosyltransferase
Nature Communications (2023)
-
Rapid biosynthesis of glycoprotein therapeutics and vaccines from freeze-dried bacterial cell lysates
Nature Protocols (2023)