Abstract
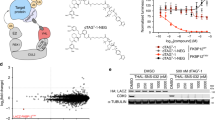
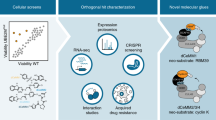
Dissection of complex biological systems requires target-specific control of the function or abundance of proteins. Genetic perturbations are limited by off-target effects, multicomponent complexity, and irreversibility. Most limiting is the requisite delay between modulation to experimental measurement. To enable the immediate and selective control of single protein abundance, we created a chemical biology system that leverages the potency of cell-permeable heterobifunctional degraders. The dTAG system pairs a novel degrader of FKBP12F36V with expression of FKBP12F36V in-frame with a protein of interest. By transgene expression or CRISPR-mediated locus-specific knock-in, we exemplify a generalizable strategy to study the immediate consequence of protein loss. Using dTAG, we observe an unexpected superior antiproliferative effect of pan-BET bromodomain degradation over selective BRD4 degradation, characterize immediate effects of KRASG12V loss on proteomic signaling, and demonstrate rapid degradation in vivo. This technology platform will confer kinetic resolution to biological investigation and provide target validation in the context of drug discovery.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Winter, G. E. et al. Phthalimide conjugation as a strategy for in vivo target protein degradation. Science 348, 1376–1381 (2015).
Lu, J. et al. Hijacking the E3 ubiquitin ligase cereblon to efficiently target BRD4. Chem. Biol. 22, 755–763 (2015).
Zengerle, M., Chan, K. H. & Ciulli, A. Selective small molecule induced degradation of the BET bromodomain protein BRD4. ACS Chem. Biol. 10, 1770–1777 (2015).
Winter, G. E. et al. BET bromodomain proteins function as master transcription elongation factors independent of CDK9 recruitment. Mol. Cell 67, 5–18.e19 (2017).
Bondeson, D. P. et al. Catalytic in vivo protein knockdown by small-molecule PROTACs. Nat. Chem. Biol. 11, 611–617 (2015).
Remillard, D. et al. Degradation of the BAF complex factor BRD9 by heterobifunctional ligands. Angew. Chem. Int. Edn. Engl 56, 5738–5743 (2017).
Bonger, K. M., Chen, L. C., Liu, C. W. & Wandless, T. J. Small-molecule displacement of a cryptic degron causes conditional protein degradation. Nat. Chem. Biol. 7, 531–537 (2011).
Neklesa, T. K. et al. Small-molecule hydrophobic tagging-induced degradation of HaloTag fusion proteins. Nat. Chem. Biol. 7, 538–543 (2011).
Buckley, D. L. et al. HaloPROTACS: Use of small molecule PROTACs to induce degradation of HaloTag fusion proteins. ACS Chem. Biol. 10, 1831–1837 (2015).
Chung, H. K. et al. Tunable and reversible drug control of protein production via a self-excising degron. Nat. Chem. Biol. 11, 713–720 (2015).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat. Methods 6, 917–922 (2009).
Banaszynski, L. A., Chen, L. C., Maynard-Smith, L. A., Ooi, A. G. & Wandless, T. J. A rapid, reversible, and tunable method to regulate protein function in living cells using synthetic small molecules. Cell 126, 995–1004 (2006).
Shoulders, M. D., Ryno, L. M., Cooley, C. B., Kelly, J. W. & Wiseman, R. L. Broadly applicable methodology for the rapid and dosable small molecule-mediated regulation of transcription factors in human cells. J. Am. Chem. Soc. 135, 8129–8132 (2013).
Zhou, Q. et al. A chemical genetics approach for the functional assessment of novel cancer genes. Cancer Res. 75, 1949–1958 (2015).
Clackson, T. et al. Redesigning an FKBP-ligand interface to generate chemical dimerizers with novel specificity. Proc. Natl. Acad. Sci. USA 95, 10437–10442 (1998).
Roberts, J. M. & Bradner, J. E. A bead-based proximity assay for BRD4 ligand discovery. Curr. Protoc. Chem. Biol. 7, 263–278 (2015).
Douglass, E. F. Jr., Miller, C. J., Sparer, G., Shapiro, H. & Spiegel, D. A. A comprehensive mathematical model for three-body binding equilibria. J. Am. Chem. Soc. 135, 6092–6099 (2013).
Lu, G. et al. The myeloma drug lenalidomide promotes the cereblon-dependent destruction of Ikaros proteins. Science 343, 305–309 (2014).
Krönke, J. et al. Lenalidomide causes selective degradation of IKZF1 and IKZF3 in multiple myeloma cells. Science 343, 301–305 (2014).
Schneekloth, J. S. Jr. et al. Chemical genetic control of protein levels: selective in vivo targeted degradation. J. Am. Chem. Soc. 126, 3748–3754 (2004).
Anand, P. et al. BET bromodomains mediate transcriptional pause release in heart failure. Cell 154, 569–582 (2013).
Bradner, J. E., Hnisz, D. & Young, R. A. Transcriptional addiction in cancer. Cell 168, 629–643 (2017).
Delmore, J. E. et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell 146, 904–917 (2011).
Deshpande, A. J., Bradner, J. & Armstrong, S. A. Chromatin modifications as therapeutic targets in MLL-rearranged leukemia. Trends Immunol. 33, 563–570 (2012).
Lovén, J. et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 153, 320–334 (2013).
Shortt, J., Ott, C. J., Johnstone, R. W. & Bradner, J. E. A chemical probe toolbox for dissecting the cancer epigenome. Nat. Rev. Cancer 17, 160–183 (2017).
Filippakopoulos, P. et al. Selective inhibition of BET bromodomains. Nature 468, 1067–1073 (2010).
Tanaka, M. et al. Design and characterization of bivalent BET inhibitors. Nat. Chem. Biol. 12, 1089–1096 (2016).
Sakuma, T., Nakade, S., Sakane, Y., Suzuki, K. T. & Yamamoto, T. MMEJ-assisted gene knock-in using TALENs and CRISPR-Cas9 with the PITCh systems. Nat. Protoc. 11, 118–133 (2016).
Stephen, A. G., Esposito, D., Bagni, R. K. & McCormick, F. Dragging ras back in the ring. Cancer Cell 25, 272–281 (2014).
Cox, A. D., Fesik, S. W., Kimmelman, A. C., Luo, J. & Der, C. J. Drugging the undruggable RAS: mission possible? Nat. Rev. Drug Discov. 13, 828–851 (2014).
Feramisco, J. R., Gross, M., Kamata, T., Rosenberg, M. & Sweet, R. W. Microinjection of the oncogene form of the human H-ras (T-24) protein results in rapid proliferation of quiescent cells. Cell 38, 109–117 (1984).
Shih, C., Padhy, L. C., Murray, M. & Weinberg, R. A. Transforming genes of carcinomas and neuroblastomas introduced into mouse fibroblasts. Nature 290, 261–264 (1981).
Stacey, D. W. & Kung, H. F. Transformation of NIH 3T3 cells by microinjection of Ha-ras p21 protein. Nature 310, 508–511 (1984).
McAlister, G. C. et al. Increasing the multiplexing capacity of TMTs using reporter ion isotopologues with isobaric masses. Anal. Chem. 84, 7469–7478 (2012).
McAlister, G. C. et al. MultiNotch MS3 enables accurate, sensitive, and multiplexed detection of differential expression across cancer cell line proteomes. Anal. Chem. 86, 7150–7158 (2014).
Erickson, B. K. et al. Evaluating multiplexed quantitative phosphopeptide analysis on a hybrid quadrupole mass filter/linear ion trap/orbitrap mass spectrometer. Anal. Chem. 87, 1241–1249 (2015).
Smeal, T., Binetruy, B., Mercola, D. A., Birrer, M. & Karin, M. Oncogenic and transcriptional cooperation with Ha-Ras requires phosphorylation of c-Jun on serines 63 and 73. Nature 354, 494–496 (1991).
Sears, R. et al. Multiple Ras-dependent phosphorylation pathways regulate Myc protein stability. Genes Dev. 14, 2501–2514 (2000).
Chin, Y. R. & Toker, A. The actin-bundling protein palladin is an Akt1-specific substrate that regulates breast cancer cell migration. Mol. Cell 38, 333–344 (2010).
Gilmartin, A. G. et al. GSK1120212 (JTP-74057) is an inhibitor of MEK activity and activation with favorable pharmacokinetic properties for sustained in vivo pathway inhibition. Clin. Cancer Res. 17, 989–1000 (2011).
Robert, C. et al. Improved overall survival in melanoma with combined dabrafenib and trametinib. N. Engl. J. Med. 372, 30–39 (2015).
Nabet, B. et al. Deregulation of the Ras-Erk signaling axis modulates the enhancer landscape. Cell Rep. 12, 1300–1313 (2015).
Natsume, T., Kiyomitsu, T., Saga, Y. & Kanemaki, M. T. Rapid protein depletion in human cells by auxin-inducible degron tagging with short homology donors. Cell Rep. 15, 210–218 (2016).
Weintraub, A. S. et al. YY1 is a structural regulator of enhancer-promoter loops. Cell 171, 1573–1588.e28 (2017).
Bisgrove, D. A., Mahmoudi, T., Henklein, P. & Verdin, E. Conserved P-TEFb-interacting domain of BRD4 inhibits HIV transcription. Proc. Natl. Acad. Sci. USA 104, 13690–13695 (2007).
Schröder, S. et al. Two-pronged binding with bromodomain-containing protein 4 liberates positive transcription elongation factor b from inactive ribonucleoprotein complexes. J. Biol. Chem. 287, 1090–1099 (2012).
Banaszynski, L. A., Sellmyer, M. A., Contag, C. H., Wandless, T. J. & Thorne, S. H. Chemical control of protein stability and function in living mice. Nat. Med. 14, 1123–1127 (2008).
Erb, M. A. et al. Transcription control by the ENL YEATS domain in acute leukaemia. Nature 543, 270–274 (2017).
Huang, H. T. et al. MELK is not necessary for the proliferation of basal-like breast cancer cells. eLife 6, e26693 (2017).
Yang, X. et al. A public genome-scale lentiviral expression library of human ORFs. Nat. Methods 8, 659–661 (2011).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Lovén, J. et al. Revisiting global gene expression analysis. Cell 151, 476–482 (2012).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
We thank N. Kwiatkowski for critical reading of the manuscript, W. Kaelin for sharing referenced cell lines and the dual luciferase plasmids (Dana-Farber Cancer Institute, pLL3.7-EF1a-IRES-Gateway-nluc-2xHA-IRES2-fluc-hCL1-P2A-Puro), R. Kunz and the Thermo Fisher Scientific Center for Multiplexed Proteomics at the Harvard Medical School for the quantitative proteomics and phosphoproteomics assessment, and S. Nabet, A. Aguirre, W. Hahn, and members of the Bradner and Gray laboratories for helpful discussions. This work was supported by an American Cancer Society Postdoctoral Fellowship PF-17-010-01-CDD (B.N.), the Claudia Adams Barr Program in Innovative Basic Cancer Research (D.L.B.), Damon Runyon Cancer Research Foundation DRG-2196-14 (D.L.B.), and generous philanthropic gifts from the Hale Center for Pancreatic Cancer Research and the Katherine L. and Steven C. Pinard Research Fund.
Author information
Authors and Affiliations
Contributions
B.N., J.M.R., and D.L.B. conceived and led the study under the supervision of N.S.G. and J.E.B. D.L.B. and S.D. designed and performed molecule synthesis. J.M.R. constructed the lentiviral and knock-in vector systems. B.N. and J.M.R. designed and performed BRD4 knock-in and target panel studies. B.N. designed and performed KRAS studies. J.M.R. and J.P. designed and performed AlphaScreen assays and IKZF1 dual luciferase assays. S.V. and G.E.W. constructed FKBP12 dual luciferase vectors and B.N., J.M.R., and S.V. performed experiments using these systems. B.N. designed and performed RNA-sequencing experiments and B.N., M.A.E., and M.A.L. performed bioinformatics analyses. B.N., A.Y., A.S., and K.-K.W. designed and performed mouse studies. A.L.L. and T.G.S. assisted in cellular experiments. J.A.P. provided technical advice and data interpretation. J.Q. contributed reagents and technical advice. B.N. and J.E.B. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors claim the following competing financial interests: International Patent Application Nos. PCT/US2016/039048, PCT/US2016/046087, PCT/US2016/046088, PCT/US2016/046089, each filed in the name of Dana-Farber Cancer Institute, Inc. D.L.B., J.P., and A.S. are now employees of Novartis. G.E.W. is a consultant for C4 Therapeutics. N.S.G. is a Scientific Founder and member of the Scientific Advisory Board of Syros Pharmaceuticals, C4 Therapeutics, and Petra Pharmaceuticals and is the inventor on IP licensed to these entities. J.E.B. is a Scientific Founder of Syros Pharmaceuticals, SHAPE Pharmaceuticals, Acetylon Pharmaceuticals, Tensha Therapeutics (now Roche), and C4 Therapeutics and is the inventor on IP licensed to these entities. J.E.B. is now an executive and shareholder in Novartis AG.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Table 1–6, Supplementary Figures 1–15
Supplementary Note
Synthetic procedures and characterization of dFKBP-1, dTAG-7, dTAG-13, dTAG-48, dTAG-51, bio-SLF, bio-Thal
Supplementary Dataset 1
Entire list of normalized and scaled quantitative mass spectrometry-based proteomics data. Triplicate values from biologically independent samples of normalized percent relative abundance of quantified proteins are presented for NIH/3T3 cells expressing FKBP12F36V-KRASG12V treated with DMSO, 1 µM dTAG-13 for one hour, and 1 µM dTAG-13 for four hours.
Supplementary Dataset 2
Entire list of normalized and scaled quantitative mass spectrometry-based phosphoserine/threonine data. Triplicate values from biologically independent samples of normalized percent relative abundance of quantified phosphosites are presented for NIH/3T3 cells expressing FKBP12F36V-KRASG12V treated with DMSO, 1 µM dTAG-13 for one hour, and 1 µM dTAG-13 for four hours.
Supplementary Dataset 3
Entire list of normalized and scaled quantitative mass spectrometry-based phosphotyrosine data. Triplicate values from biologically independent samples of normalized percent relative abundance of quantified phosphosites are presented for NIH/3T3 cells expressing FKBP12F36V-KRASG12V treated with DMSO, 1 µM dTAG-13 for one hour, and 1 µM dTAG-13 for four hours.
Supplementary Dataset 4
ERCC spike-in normalized FPKM values of top 200 upregulated and 200 downregulated expressed transcripts upon comparison of mock transduced (control) NIH/3T3 cells or NIH/3T3 cells expressing FKBP12F36V-KRASG12V treated with DMSO. Triplicate FPKM values from biologically independent samples are presented for control NIH/3T3 cells treated with DMSO and NIH/3T3 cells expressing FKBP12F36V-KRASG12V treated with DMSO, 1 µM dTAG-13, or 10 nM trametinib. Dataset accompanies heatmap in Fig. 5c.
Rights and permissions
About this article
Cite this article
Nabet, B., Roberts, J.M., Buckley, D.L. et al. The dTAG system for immediate and target-specific protein degradation. Nat Chem Biol 14, 431–441 (2018). https://doi.org/10.1038/s41589-018-0021-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0021-8
This article is cited by
-
HiHo-AID2: boosting homozygous knock-in efficiency enables robust generation of human auxin-inducible degron cells
Genome Biology (2024)
-
Context-specific functions of chromatin remodellers in development and disease
Nature Reviews Genetics (2024)
-
Matrin3 mediates differentiation through stabilizing chromatin loop-domain interactions and YY1 mediated enhancer-promoter interactions
Nature Communications (2024)
-
Structural insights into histone exchange by human SRCAP complex
Cell Discovery (2024)
-
Proteolysis-targeting chimeras with reduced off-targets
Nature Chemistry (2024)