Abstract
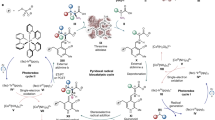
Sugar O-methylation shields algal polysaccharides against microbial hydrolytic enzymes. Here, we describe cytochrome P450 monooxygenases from marine bacteria that, together with appropriate redox-partner proteins, catalyze the oxidative demethylation of 6-O-methyl-d-galactose, which is an abundant monosaccharide of the algal polysaccharides agarose and porphyran. This previously unknown biological function extends the group of carbohydrate-active enzymes to include the class of cytochrome P450 monooxygenases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
08 March 2018
In the version of this article originally published, the line of conditions shown for NADH in Figure 2b was shifted out of place. The error has been corrected in the HTML and PDF versions of the article.
References
Field, C. B., Behrenfeld, M. J., Randerson, J. T. & Falkowski, P. Science 281, 237–240 (1998).
Kraan, S. in Carbohydrates–Comprehensive Studies on Glycobiology and Glycotechnology (ed. Chang, C.-F.) Ch. 22 https://doi.org/10.5772/51572 (INTECH Open Access Publisher, 2012).
Wargacki, A. J. et al. Science 335, 308–313 (2012).
Motone, K., Takagi, T., Sasaki, Y., Kuroda, K. & Ueda, M. J. Biotechnol. 231, 129–135 (2016).
Panagiotopoulos, C., Repeta, D. J., Mathieu, L., Rontani, J. F. & Sempéré, R. Mar. Chem. 154, 34–45 (2013).
Gabrielii, I., Gatenholm, P., Glasser, W. G., Jain, R. K. & Kenne, L. Carbohydr. Polym. 43, 367–374 (2000).
Hehemann, J. H., Kelly, A. G., Pudlo, N. A., Martens, E. C. & Boraston, A. B. Proc. Natl. Acad. Sci. USA 109, 19786–19791 (2012).
Rees, D. A. & Conway, E. Biochem. J. 84, 411–416 (1962).
Chiovitti, A., Bacic, A., Craik, D. J., Kraft, G. T. & Liao, M. L. Carbohydr. Res. 339, 1459–1466 (2004).
Guengerich, F. P. Chem. Res. Toxicol. 14, 611–650 (2001).
Lewis, J. C. et al. Proc. Natl. Acad. Sci. USA 106, 16550–16555 (2009).
Katagiri, M., Ganguli, B. N. & Gunsalus, I. C. J. Biol. Chem. 243, 3543–3546 (1968).
Janusz, G., Kucharzyk, K. H., Pawlik, A., Staszczak, M. & Paszczynski, A. J. Enzyme Microb. Technol. 52, 1–12 (2013).
Hemsworth, G. R. et al. J. Am. Chem. Soc. 135, 6069–6077 (2013).
Levasseur, A., Drula, E., Lombard, V., Coutinho, P. M. & Henrissat, B. Biotechnol. Biofuels 6, 41 (2013).
Yin, D. T. et al. Nat. Commun. 6, 10197 (2015).
Vuong, T. V., Liu, B., Sandgren, M. & Master, E. R. Biomacromolecules 18, 610–616 (2017).
Mann, A. J. et al. Appl. Environ. Microbiol. 79, 6813–6822 (2013).
Girhard, M., Tieves, F., Weber, E., Smit, M. S. & Urlacher, V. B. Appl. Microbiol. Biotechnol. 97, 1625–1635 (2013).
Kim, H. T., Lee, S., Kim, K. H. & Choi, I. G. Bioresour. Technol. 107, 301–306 (2012).
Girhard, M. et al. Microb. Cell Fact. 8, 36 (2009).
Nickerson, D. P. & Wong, L. L. Protein Eng. 10, 1357–1361 (1997).
Li, C. et al. BMC Biotechnol. 11, 92 (2011).
Purdy, M. M., Koo, L. S., Ortiz de Montellano, P. R. & Klinman, J. P. Biochemistry 43, 271–281 (2004).
Omura, T. & Sato, R. J. Biol. Chem. 239, 2370–2378 (1964).
Jefcoate, C. R. Methods Enzymol. 52, 258–279 (1978).
Ruiz-Matute, A. I., Hernández-Hernández, O., Rodríguez-Sánchez, S., Sanz, M. L. & Martínez-Castro, I. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 879, 1226–1240 (2011).
Kumar, S., Stecher, G. & Tamura, K. Mol. Biol. Evol. 33, 1870–1874 (2016).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. J. Mol. Biol. 215, 403–410 (1990).
Edgar, R. C. Nucleic Acids Res. 32, 1792–1797 (2004).
Hehemann, J. H., Smyth, L., Yadav, A., Vocadlo, D. J. & Boraston, A. B. J. Biol. Chem. 287, 13985–13995 (2012).
Finn, R. D. et al. Nucleic Acids Res. 44, D279–D285 (2016). D1.
Yin, Y. et al. Nucleic Acids Res. 40, W445–W451 (2012).
Krzywinski, M. et al. Genome Res. 19, 1639–1645 (2009).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. J. Mol. Biol. 305, 567–580 (2001).
Acknowledgements
We thank the German Research Foundation (DFG) for funding through the Research Unit FOR2406. J.-H.H. acknowledges funding by the Emmy Noether Program of the DFG, grant number HE 7217/1-1. We are also grateful to V. Urlacher (Düsseldorf, Germany) for providing the genes encoding P450cam, PdX, PdR and CPR. We thank D. Nelson (Memphis, USA) for assigning the P450s to a subfamily in the P450 database.
Author information
Authors and Affiliations
Contributions
J.-H.H., T.S. and U.T.B. initiated the study and directed the project. J.E. together with L.R. cloned, expressed and purified the P450 enzymes and performed preliminary studies; H.C.B. and L.R. cloned, expressed and purified all other enzymes and performed all further experiments. S.T. and J.-H.H. performed the computational analysis. L.R., H.C.B., J.-H.H. and U.T.B. prepared the manuscript, which was revised and approved by all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
A correction to this article is available online at https://doi.org/10.1038/s41589-018-0020-9.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1 and 2 and Supplementary Figures 1–7
Rights and permissions
About this article
Cite this article
Reisky, L., Büchsenschütz, H.C., Engel, J. et al. Oxidative demethylation of algal carbohydrates by cytochrome P450 monooxygenases. Nat Chem Biol 14, 342–344 (2018). https://doi.org/10.1038/s41589-018-0005-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0005-8
This article is cited by
-
Unique alcohol dehydrogenases involved in algal sugar utilization by marine bacteria
Applied Microbiology and Biotechnology (2023)
-
Metabolism of a hybrid algal galactan by members of the human gut microbiome
Nature Chemical Biology (2022)
-
North Sea spring bloom-associated Gammaproteobacteria fill diverse heterotrophic niches
Environmental Microbiome (2021)
-
Blastopirellula retiformator sp. nov. isolated from the shallow-sea hydrothermal vent system close to Panarea Island
Antonie van Leeuwenhoek (2020)
-
Bakterielle Mechanismen der marinen Polysaccharidverwertung
BIOspektrum (2020)