Abstract
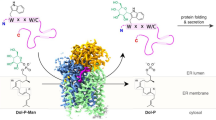
Protein glycosylation regulates many cellular processes. Numerous glycosyltransferases with broad substrate specificities have been structurally characterized. A novel inverting glycosyltransferase, EarP, specifically transfers rhamnose from dTDP-β-l-rhamnose to Arg32 of bacterial translation elongation factor P (EF-P) to activate its function. Here we report a crystallographic study of Neisseria meningitidis EarP. The EarP structure contains two tandem Rossmann-fold domains, which classifies EarP in glycosyltransferase superfamily B. In contrast to other structurally characterized protein glycosyltransferases, EarP binds the entire β-sheet structure of EF-P domain I through numerous interactions that specifically recognize its conserved residues. Thus Arg32 is properly located at the active site, and causes structural change in a conserved dTDP-β-l-rhamnose-binding loop of EarP. Rhamnosylation by EarP should occur via an SN2 reaction, with Asp20 as the general base. The Arg32 binding and accompanying structural change of EarP may induce a change in the rhamnose-ring conformation suitable for the reaction.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Lairson, L. L., Henrissat, B., Davies, G. J. & Withers, S. G. Glycosyltransferases: structures, functions, and mechanisms. Annu. Rev. Biochem. 77, 521–555 (2008).
Coutinho, P. M., Deleury, E., Davies, G. J. & Henrissat, B. An evolving hierarchical family classification for glycosyltransferases. J. Mol. Biol. 328, 307–317 (2003).
Liu, J. & Mushegian, A. Three monophyletic superfamilies account for the majority of the known glycosyltransferases. Protein Sci. 12, 1418–1431 (2003).
Cantarel, B. L. et al. The Carbohydrate-Active EnZymes database (CAZy): an expert resource for glycogenomics. Nucleic Acids Res. 37, D233–D238 (2009).
Hurtado-Guerrero, R. & Davies, G. J. Recent structural and mechanistic insights into post-translational enzymatic glycosylation. Curr. Opin. Chem. Biol. 16, 479–487 (2012).
Fritz, T. A., Raman, J. & Tabak, L. A. Dynamic association between the catalytic and lectin domains of human UDP-GalNAc:polypeptide alpha-N-acetylgalactosaminyltransferase-2. J. Biol. Chem. 281, 8613–8619 (2006).
Lazarus, M. B., Nam, Y., Jiang, J., Sliz, P. & Walker, S. Structure of human O-GlcNAc transferase and its complex with a peptide substrate. Nature 469, 564–567 (2011).
Lizak, C., Gerber, S., Numao, S., Aebi, M. & Locher, K. P. X-ray structure of a bacterial oligosaccharyltransferase. Nature 474, 350–355 (2011).
Yu, H. et al. Notch-modifying xylosyltransferase structures support an SNi-like retaining mechanism. Nat. Chem. Biol. 11, 847–854 (2015).
Valero-González, J. et al. A proactive role of water molecules in acceptor recognition by protein O-fucosyltransferase 2. Nat. Chem. Biol. 12, 240–246 (2016).
Li, Z. et al. Recognition of EGF-like domains by the Notch-modifying O-fucosyltransferase POFUT1. Nat. Chem. Biol. 13, 757–763 (2017).
Yu, H. et al. Structural analysis of Notch-regulating Rumi reveals basis for pathogenic mutations. Nat. Chem. Biol. 12, 735–740 (2016).
Lassak, J. et al. Arginine-rhamnosylation as new strategy to activate translation elongation factor P. Nat. Chem. Biol. 11, 266–270 (2015).
Rajkovic, A. et al. Cyclic rhamnosylated elongation factor P establishes antibiotic resistance in Pseudomonas aeruginosa. MBio 6, e00823 (2015).
Yanagisawa, T. et al. Neisseria meningitidis translation elongation factor P and its active-site arginine residue are essential for cell viability. PLoS One 11, e0147907 (2016).
Li, X. et al. Resolving the α-glycosidic linkage of arginine-rhamnosylated translation elongation factor P triggers generation of the first ArgRha specific antibody. Chem. Sci 7, 6995–7001 (2016).
Doerfel, L. K. et al. EF-P is essential for rapid synthesis of proteins containing consecutive proline residues. Science 339, 85–88 (2013).
Ude, S. et al. Translation elongation factor EF-P alleviates ribosome stalling at polyproline stretches. Science 339, 82–85 (2013).
Yanagisawa, T., Sumida, T., Ishii, R., Takemoto, C. & Yokoyama, S. A paralog of lysyl-tRNA synthetase aminoacylates a conserved lysine residue in translation elongation factor P. Nat. Struct. Mol. Biol. 17, 1136–1143 (2010).
Roy, H. et al. The tRNA synthetase paralog PoxA modifies elongation factor-P with (R)-β-lysine. Nat. Chem. Biol. 7, 667–669 (2011).
Navarre, W. W. et al. PoxA, yjeK, and elongation factor P coordinately modulate virulence and drug resistance in Salmonella enterica. Mol. Cell 39, 209–221 (2010).
De Bruyn, A. & Anteunis, M. 1H-N.m.r. study of L-rhamnose, methyl alpha-L-rhamnopyranoside, and 4-o-beta-D-galactopranosyl-L-rhamnose in deuterium oxide. Carbohydr. Res. 47, 158–163 (1976).
McGeachin, H. M. & Beevers, C. A. The crystal structure of α-rhamnose monohydrate. Acta Crystallogr. 10, 227–232 (1957).
Lira-Navarrete, E. et al. Structural insights into the mechanism of protein O-fucosylation. PLoS One 6, e25365 (2011).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Hanawa-Suetsugu, K. et al. Crystal structure of elongation factor P from Thermus thermophilus HB8. Proc. Natl. Acad. Sci. USA 101, 9595–9600 (2004).
Holm, L. & Rosenström, P. Dali server: conservation mapping in 3D. Nucleic Acids Res. 38, W545–W549 (2010).
Brazier-Hicks, M. et al. Characterization and engineering of the bifunctional N- and O-glucosyltransferase involved in xenobiotic metabolism in plants. Proc. Natl. Acad. Sci. USA 104, 20238–20243 (2007).
Isiorho, E. A., Jeon, B. S., Kim, N. H., Liu, H. W. & Keatinge-Clay, A. T. Structural studies of the spinosyn forosaminyltransferase, SpnP. Biochemistry 53, 4292–4301 (2014).
Li, L. et al. Crystal structure of Medicago truncatula UGT85H2—insights into the structural basis of a multifunctional (iso)flavonoid glycosyltransferase. J. Mol. Biol. 370, 951–963 (2007).
Mulichak, A. M., Lu, W., Losey, H. C., Walsh, C. T. & Garavito, R. M. Crystal structure of vancosaminyltransferase GtfD from the vancomycin biosynthetic pathway: interactions with acceptor and nucleotide ligands. Biochemistry 43, 5170–5180 (2004).
Hayward, S., Kitao, A. & Berendsen, H. J. Model-free methods of analyzing domain motions in proteins from simulation: a comparison of normal mode analysis and molecular dynamics simulation of lysozyme. Proteins 27, 425–437 (1997).
Hol, W. G. The role of the alpha-helix dipole in protein function and structure. Prog. Biophys. Mol. Biol. 45, 149–195 (1985).
Cremer, D. & Pople, J. A. General definition of ring puckering coordinates. J. Am. Chem. Soc. 97, 1354–1358 (1975).
Isiorho, E. A., Liu, H. W. & Keatinge-Clay, A. T. Structural studies of the spinosyn rhamnosyltransferase, SpnG. Biochemistry 51, 1213–1222 (2012).
Chen, C. I. et al. Structure of human POFUT2: insights into thrombospondin type 1 repeat fold and O-fucosylation. EMBO J. 31, 3183–3197 (2012).
Rocha, J. et al. Structure of Arabidopsis thaliana FUT1 reveals a variant of the GT-B class fold and provides insight into xyloglucan fucosylation. Plant Cell 28, 2352–2364 (2016).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. in Methods in Enzymology: Macromolecular Crystallography, Part A, Vol. 276 (eds. Carter, C.W. Jr. & Sweet, R.M.) 307–326 (Academic Press, San Diego, 1997).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Sheldrick, G. M. A short history of SHELX. Acta Crystallogr. A 64, 112–122 (2008).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Kwak, H. G. & Dohmae, N. Proteomic characterization of histone variants in the mouse testis by mass spectrometry-based top-down analysis. Biosci. Trends 10, 357–364 (2016).
Frisch, M. J. et al. Gaussian 09 (Gaussian, Inc., Wallingford, CT, USA, 2009).
Acknowledgements
We thank the staff of beamlines BL32XU and BL26B2 at SPring-8 (Harima, Japan) and beamline BL5A at the Photon Factory (Tsukuba, Japan). We thank T. Terada, T. Imada, and T. Nakayama (RIKEN) for clerical assistance. We thank K. Ohtsuki, M. Usui, and A. Abe (RIKEN) for mass spectrometry analysis, and M. Kuratani and N. Sakai (RIKEN) for technical assistance. This work was supported by the Platform Project for Supporting Drug Discovery and Life Science Research from the Japan Agency for Medical Research and Development (AMED) (to S.Y.), and by JSPS KAKENHI (grant number 16K05859 to T.Y.). This research used the computational resources of the supercomputer HOKUSAI (RIKEN Advanced Center for Computing and Communications).
Author information
Authors and Affiliations
Contributions
S.Y., H.T., and T.Y. conceived the project. T. Sengoku, S.Y., and T.Y. designed the experiments. T. Sengoku and T.Y. prepared protein samples. T. Sengoku crystallized proteins and performed crystallographic experiments and analyses. Y.H. assisted with the crystallographic analyses. N.D., T. Suzuki, and T.Y. performed mass spectrometry experiments. T.Y. performed ITC experiments. T. Sengoku, C.W., and T.H. performed the MD simulation, with advice on the sugar conformation from Y.Y. All authors analyzed the data. T. Sengoku, S.Y., and T.Y. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Table 1 and Supplementary Figures 1–15
Rights and permissions
About this article
Cite this article
Sengoku, T., Suzuki, T., Dohmae, N. et al. Structural basis of protein arginine rhamnosylation by glycosyltransferase EarP. Nat Chem Biol 14, 368–374 (2018). https://doi.org/10.1038/s41589-018-0002-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-018-0002-y