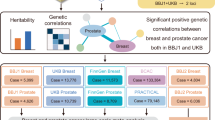
Abstract
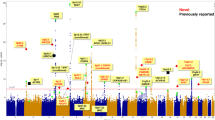
To identify new susceptibility loci to lung cancer among diverse populations, we performed cross-ancestry genome-wide association studies in European, East Asian and African populations and discovered five loci that have not been previously reported. We replicated 26 signals and identified 10 new lead associations from previously reported loci. Rare-variant associations tended to be specific to populations, but even common-variant associations influencing smoking behavior, such as those with CHRNA5 and CYP2A6, showed population specificity. Fine-mapping and expression quantitative trait locus colocalization nominated several candidate variants and susceptibility genes such as IRF4 and FUBP1. DNA damage assays of prioritized genes in lung fibroblasts indicated that a subset of these genes, including the pleiotropic gene IRF4, potentially exert effects by promoting endogenous DNA damage.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The following publicly available datasets were used in this work: dbGaP datasets (Prostate, Lung, Colorectal and Ovarian Cancer Screening Trial, phs000093.v2.p2; FLCCA study, phs000716.v1.p1; EAGLE study, phs000336.v1.p1; NCI study of African-Americans, phs001210.v1.p1; German, SLRI, IARC and MDACC studies, phs000876.v2.p1; Oncoarray study, phs001273.v3.p2; imputed Oncoarray study using HRC reference panel, phs001273.v4.p2; Affymetrix study, phs001681.v1.p1). The ICR study from the 1958 Birth Cohort from the UK does not allow the general upload of findings. Therefore, this dataset is available after request from R. Houlston (Richard.Houlston@icr.ac.uk). The individual-level genotype and phenotype data are available through formal application to the UKBB (https://www.ukbiobank.ac.uk/). The GWAS summary statistics used in validation study were downloaded from FinnGen (https://finngen.gitbook.io/documentation/v/r5/data-download) and the pan-cancer pleiotropy study (https://github.com/Wittelab/pancancer_pleiotropy). The GWAS summary statistics of the candidate 45 variants identified from the discovery phase were obtained following our request from M.Z. and H.S. (China NJMU lung study), T.R. (deCODE and SPAIN lung study) and A.S. and C.L. (INHALE study) and are available in the Supplementary Tables 9 and 10. The eQTL data from GTEx v8 was obtained from https://gtexportal.org/home/datasets. The Icelandic population WGS genetic but not phenotypic data have been deposited at the European Variant Archive under accession code PRJEB15197. Results from GWMA at P ≤ 10−5 are available in the supplementary tables. All sequencing reads were mapped to the GRCh37/hg19 human reference genome. More details of data sources used in this work are provided in the paper and supplementary tables.
Code availability
We performed our analyses using the following publicly available software/packages: SHAPE-IT2 (v2.r790; https://mathgen.stats.ox.ac.uk/genetics_software/shapeit/shapeit.html), McCarthy Group Tools (v4.2.11; https://www.well.ox.ac.uk/~wrayner/tools/), PBWT (https://github.com/richarddurbin/pbwt) and Michigan Imputation Server (https://imputationserver.sph.umich.edu/index.html#!) were used for imputation and phasing; FastPop (https://github.com/biomedicaldatascience/FastPop4) and KING (v2.0, http://people.virginia.edu/~wc9c/KING/) were used for population stratification and relatedness analyses; SAS (v9.4, https://www.sas.com/en_us/home.html), R (v3.6.2, https://cran.r-project.org), PLINK (v1.9 and 2.0, https://www.cog-genomics.org/plink/1.9/ and https://www.cog-genomics.org/plink/2.0/), METASOFT and ForestPMPlot (v2.0.1 and v1.0.3, http://genetics.cs.ucla.edu/meta/) and GCTA (v1.93, https://cnsgenomics.com/software/gcta/) were used for data and statistical analyses; FUMA (v1.3.6, https://fuma.ctglab.nl/), FAVOR (https://favor.genohub.org/), GTEx (v8, https://www.gtexportal.org/home/), coloc (v3.2-1, https://cran.r-project.org/web/packages/coloc/), eCAVIAR (v2, http://zarlab.cs.ucla.edu/tag/caviar/), IPA (https://www.qiagenbioinformatics.com/products/ingenuity-pathway-analysis) and ezQTL (v1.0, https://analysistools.cancer.gov/ezqtl/#/home) were used for post-GWAS analyses; and FlowJo (v10.6, https://www.flowjo.com) was used for single-cell flow cytometry analysis. MANTRA (version 1) is available as a suite of executables on request from the corresponding author (https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3460225/pdf/gepi0035-0809.pdf).
References
Sampson, J. N. et al. Analysis of heritability and shared heritability based on genome-wide association studies for thirteen cancer types. J. Natl Cancer Inst. 107, djv279 (2015).
Bosse, Y. & Amos, C. I. A decade of GWAS results in lung cancer. Cancer Epidemiol. Biomarkers Prev. 27, 363–379 (2018).
Park, S. L., Cheng, I. & Haiman, C. A. Genome-wide association studies of cancer in diverse populations. Cancer Epidemiol. Biomarkers Prev. 27, 405–417 (2018).
Popejoy, A. B. & Fullerton, S. M. Genomics is failing on diversity. Nature 538, 161–164 (2016).
Rosenberg, N. A. et al. Genome-wide association studies in diverse populations. Nat. Rev. Genet. 11, 356–366 (2010).
Schabath, M. B., Cress, D. & Munoz-Antonia, T. Racial and ethnic differences in the epidemiology and genomics of lung cancer. Cancer Control 23, 338–346 (2016).
Asimit, J. L., Hatzikotoulas, K., McCarthy, M., Morris, A. P. & Zeggini, E. Trans-ethnic study design approaches for fine-mapping. Eur. J. Hum. Genet. 24, 1330–1336 (2016).
Conti, D. V. et al. Trans-ancestry genome-wide association meta-analysis of prostate cancer identifies new susceptibility loci and informs genetic risk prediction. Nat. Genet. 53, 65–75 (2021).
Magi, R. et al. Trans-ethnic meta-regression of genome-wide association studies accounting for ancestry increases power for discovery and improves fine-mapping resolution. Hum. Mol. Genet. 26, 3639–3650 (2017).
Li, Y. R. & Keating, B. J. Trans-ethnic genome-wide association studies: advantages and challenges of mapping in diverse populations. Genome Med. 6, 91 (2014).
Morris, A. P. Transethnic meta-analysis of genomewide association studies. Genet. Epidemiol. 35, 809–822 (2011).
Marigorta, U. M. & Navarro, A. High trans-ethnic replicability of GWAS results implies common causal variants. PLoS Genet. 9, e1003566 (2013).
Wang, J. et al. Genetic predisposition to lung cancer: comprehensive literature integration, meta-analysis, and multiple evidence assessment of candidate-gene association studies. Sci. Rep. 7, 8371 (2017).
Bossé, Y. et al. Transcriptome-wide association study reveals candidate causal genes for lung cancer. Int. J. Cancer. 146, 1862–1878 (2020).
Kanwal, M., Ding, X. J. & Cao, Y. Familial risk for lung cancer. Oncol. Lett. 13, 535–542 (2017).
Rashkin, S. R. et al. Pan-cancer study detects genetic risk variants and shared genetic basis in two large cohorts. Nat. Commun. 11, 4423 (2020).
Jiang, X. et al. Shared heritability and functional enrichment across six solid cancers. Nat. Commun. 10, 431 (2019).
McKay, J. D. et al. Large-scale association analysis identifies new lung cancer susceptibility loci and heterogeneity in genetic susceptibility across histological subtypes. Nat. Genet. 49, 1126–1132 (2017).
Li, Y. et al. Genome-wide interaction study of smoking behavior and non-small cell lung cancer risk in Caucasian population. Carcinogenesis 39, 336–346 (2018).
Li, Y. et al. Genetic interaction analysis among oncogenesis-related genes revealed novel genes and networks in lung cancer development. Oncotarget 10, 1760–1774 (2019).
Ji, X. et al. Identification of susceptibility pathways for the role of chromosome 15q25.1 in modifying lung cancer risk. Nat. Commun. 9, 3221 (2018).
Ji, X. et al. Protein-altering germline mutations implicate novel genes related to lung cancer development. Nat. Commun. 11, 2220 (2020).
Amos, C. I. et al. The OncoArray Consortium: a network for understanding the genetic architecture of common cancers. Cancer Epidemiol. Biomarkers Prev. 26, 126–135 (2017).
Byun, J. et al. Genome-wide association study of familial lung cancer. Carcinogenesis 39, 1135–1140 (2018).
Lan, Q. et al. Genome-wide association analysis identifies new lung cancer susceptibility loci in never-smoking women in Asia. Nat. Genet. 44, 1330–1335 (2012).
Kachuri, L. et al. Fine mapping of chromosome 5p15.33 based on a targeted deep sequencing and high density genotyping identifies novel lung cancer susceptibility loci. Carcinogenesis 37, 96–105 (2016).
Zanetti, K. A. et al. Genome-wide association study confirms lung cancer susceptibility loci on chromosomes 5p15 and 15q25 in an African-American population. Lung Cancer 98, 33–42 (2016).
Wang, Y. et al. Rare variants of large effect in BRCA2 and CHEK2 affect risk of lung cancer. Nat. Genet. 46, 736–741 (2014).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Truong, T. et al. Replication of lung cancer susceptibility loci at chromosomes 15q25, 5p15, and 6p21: a pooled analysis from the International Lung Cancer Consortium. J. Natl Cancer Inst. 102, 959–971 (2010).
Zuber, V. et al. Pleiotropic analysis of lung cancer and blood triglycerides. J. Natl Cancer. Inst. 108, djw167 (2016).
Watza, D. et al. COPD-dependent effects of genetic variation in key inflammation pathway genes on lung cancer risk. Int. J. Cancer 147, 747–756 (2020).
Dai, J. et al. Identification of risk loci and a polygenic risk score for lung cancer: a large-scale prospective cohort study in Chinese populations. Lancet Respir. Med. 7, 881–891 (2019).
van Rooij, F. J. A. et al. Genome-wide trans-ethnic meta-analysis identifies seven genetic loci influencing erythrocyte traits and a role for RBPMS in erythropoiesis. Am. J. Hum. Genet. 100, 51–63 (2017).
Li, Y. et al. FastPop: a rapid principal component derived method to infer intercontinental ancestry using genetic data. BMC Bioinformatics 17, 122 (2016).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283 (2016).
Wang, X. Firth logistic regression for rare variant association tests. Front. Genet. 5, 187 (2014).
Ma, C., Blackwell, T., Boehnke, M., Scott, L. J. & Go, T. D. I. Recommended joint and meta-analysis strategies for case-control association testing of single low-count variants. Genet. Epidemiol. 37, 539–550 (2013).
Dey, R. et al. Robust meta-analysis of biobank-based genome-wide association studies with unbalanced binary phenotypes. Genet. Epidemiol. 43, 462–476 (2019).
Han, B. & Eskin, E. Random-effects model aimed at discovering associations in meta-analysis of genome-wide association studies. Am. J. Hum. Genet. 88, 586–598 (2011).
Han, B. & Eskin, E. Interpreting meta-analyses of genome-wide association studies. PLoS Genet. 8, e1002555 (2012).
Bhattacharjee, S. et al. A subset-based approach improves power and interpretation for the combined analysis of genetic association studies of heterogeneous traits. Am. J. Hum. Genet. 90, 821–835 (2012).
Igl, B. W., Konig, I. R. & Ziegler, A. What do we mean by ‘replication’ and ‘validation’ in genome-wide association studies? Hum. Hered. 67, 66–68 (2009).
Spitz, M. R. et al. Role of selected genetic variants in lung cancer risk in African Americans. J. Thorac. Oncol. 8, 391–397 (2013).
Machiela, M. J. & Chanock, S. J. LDlink: a web-based application for exploring population-specific haplotype structure and linking correlated alleles of possible functional variants. Bioinformatics 31, 3555–3557 (2015).
Buniello, A. et al. The NHGRI-EBI GWAS catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 47, D1005–d1012 (2019).
Schumacher, F. R. et al. Genome-wide association study of colorectal cancer identifies six new susceptibility loci. Nat. Commun. 6, 7138 (2015).
Doyle, G. A. et al. In vitro and ex vivo analysis of CHRNA3 and CHRNA5 haplotype expression. PLoS ONE 6, e23373 (2011).
Tanner, J. A. et al. Novel CYP2A6 diplotypes identified through next-generation sequencing are associated with in-vitro and in-vivo nicotine metabolism. Pharmacogenet. Genomics 28, 7–16 (2018).
Kang, E. Y. et al. Meta-analysis identifies gene-by-environment interactions as demonstrated in a study of 4,965 mice. PLoS Genet. 10, e1004022 (2014).
Pena-Chilet, M. et al. Genetic variants in PARP1 (rs3219090) and IRF4 (rs12203592) genes associated with melanoma susceptibility in a Spanish population. BMC Cancer 13, 160 (2013).
Chen, M. H. et al. Trans-ethnic and ancestry-specific blood-cell genetics in 746,667 individuals from 5 global populations. Cell 182, 1198–1213 (2020).
Vuckovic, D. et al. The polygenic and monogenic basis of blood traits and diseases. Cell 182, 1214–1231 (2020).
Astle, W. J. et al. The allelic landscape of human blood cell trait variation and links to common complex disease. Cell 167, 1415–1429 (2016).
Liyanage, U. E. et al. Combined analysis of keratinocyte cancers identifies novel genome-wide loci. Hum. Mol. Genet. 28, 3148–3160 (2019).
Asgari, M. M. et al. Identification of susceptibility loci for cutaneous squamous cell carcinoma. J. Invest. Dermatol. 136, 930–937 (2016).
Chahal, H. S. et al. Genome-wide association study identifies novel susceptibility loci for cutaneous squamous cell carcinoma. Nat. Commun. 7, 12048 (2016).
Liu, M. et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat. Genet. 51, 237–244 (2019).
Kichaev, G. et al. Leveraging polygenic functional enrichment to improve GWAS power. Am. J. Hum. Genet. 104, 65–75 (2019).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Landi, M. T. et al. A genome-wide association study of lung cancer identifies a region of chromosome 5p15 associated with risk for adenocarcinoma. Am. J. Hum. Genet. 85, 679–691 (2009).
McKay, J. D. et al. Lung cancer susceptibility locus at 5p15.33. Nat. Genet. 40, 1404–1406 (2008).
Hung, R. J. et al. Lung cancer risk in never-smokers of European descent is associated with genetic variation in the 5p15.33 TERT-CLPTM1Ll region. J. Thorac. Oncol. 14, 1360–1369 (2019).
Shiraishi, K. et al. A genome-wide association study identifies two new susceptibility loci for lung adenocarcinoma in the Japanese population. Nat. Genet. 44, 900–903 (2012).
Hu, Z. et al. A genome-wide association study identifies two new lung cancer susceptibility loci at 13q12.12 and 22q12.2 in Han Chinese. Nat. Genet. 43, 792–796 (2011).
Hsiung, C. A. et al. The 5p15.33 locus is associated with risk of lung adenocarcinoma in never-smoking females in Asia. PLoS Genet. 6, e1001051 (2010).
Schaid, D. J., Chen, W. & Larson, N. B. From genome-wide associations to candidate causal variants by statistical fine-mapping. Nat. Rev. Genet. 19, 491–504 (2018).
Cannon, M. E. et al. Trans-ancestry Fine mapping and molecular assays identify regulatory variants at the ANGPTL8 HDL-C GWAS. Locus. G3 7, 3217–3227 (2017).
Li, X. et al. Dynamic incorporation of multiple in silico functional annotations empowers rare variant association analysis of large whole-genome sequencing studies at scale. Nat. Genet. 52, 969–983 (2020).
Sun, R. et al. Integration of multiomic annotation data to prioritize and characterize inflammation and immune-related risk variants in squamous cell lung cancer. Genet. Epidemiol. 45, 99–114 (2021).
Li, X. et al. A multi-dimensional integrative scoring framework for predicting functional variants in the human genome. Am. J. Hum. Genet. 109, 446–456 (2022).
Watanabe, K., Taskesen, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8, 1826 (2017).
Elman, J. S. et al. Identification of FUBP1 as a long tail cancer driver and widespread regulator of tumor suppressor and oncogene alternative splicing. Cell Rep. 28, 3435–3449 (2019).
Singer, S. et al. Coordinated expression of stathmin family members by far upstream sequence element-binding protein-1 increases motility in non-small cell lung cancer. Cancer Res. 69, 2234–2243 (2009).
Man, K. et al. The transcription factor IRF4 is essential for TCR affinity-mediated metabolic programming and clonal expansion of T cells. Nat. Immunol. 14, 1155–1165 (2013).
Praetorius, C. et al. A polymorphism in IRF4 affects human pigmentation through a tyrosinase-dependent MITF/TFAP2A pathway. Cell 155, 1022–1033 (2013).
Shaffer, A. L. et al. IRF4 addiction in multiple myeloma. Nature 454, 226–231 (2008).
Do, T. N., Ucisik-Akkaya, E., Davis, C. F., Morrison, B. A. & Dorak, M. T. An intronic polymorphism of IRF4 gene influences gene transcription in vitro and shows a risk association with childhood acute lymphoblastic leukemia in males. Biochim. Biophys. Acta 1802, 292–300 (2010).
Zhang, T. et al. Cell-type-specific eQTL of primary melanocytes facilitates identification of melanoma susceptibility genes. Genome Res. 28, 1621–1635 (2018).
Visser, M., Palstra, R. J. & Kayser, M. Allele-specific transcriptional regulation of IRF4 in melanocytes is mediated by chromatin looping of the intronic rs12203592 enhancer to the IRF4 promoter. Hum. Mol. Genet. 24, 2649–2661 (2015).
Tubbs, A. & Nussenzweig, A. Endogenous DNA damage as a source of genomic instability in cancer. Cell 168, 644–656 (2017).
Xia, J. et al. Bacteria-to-Human protein networks reveal origins of endogenous DNA damage. Cell 176, 127–143 (2019).
Liu, Y. et al. Rare deleterious germline variants and risk of lung cancer. NPJ Precis. Oncol. 5, 12 (2021).
Gomperts, B. N. et al. Evolving concepts in lung carcinogenesis. Semin. Respir. Crit. Care Med. 32, 32–43 (2011).
Miller, Y. E. Pathogenesis of lung cancer: 100 year report. Am. J. Respir. Cell Mol. Biol. 33, 216–223 (2005).
Landi, M. T. et al. Environment And Genetics in Lung cancer Etiology (EAGLE) study: an integrative population-based case-control study of lung cancer. BMC Public Health 8, 203 (2008).
Mitchell, K. A. et al. Relationship between West African ancestry with lung cancer risk and survival in African Americans. Cancer Causes Control 30, 1259–1268 (2019).
Wigginton, J. E., Cutler, D. J. & Abecasis, G. R. A note on exact tests of Hardy–Weinberg equilibrium. Am. J. Hum. Genet. 76, 887–893 (2005).
Graffelman, J. & Moreno, V. The mid p-value in exact tests for Hardy–Weinberg equilibrium. Stat. Appl. Genet. Mol. Biol. 12, 433–448 (2013).
Rafnar, T. et al. Variants associating with uterine leiomyoma highlight genetic background shared by various cancers and hormone-related traits. Nat. Commun. 9, 3636 (2018).
Genomes Project, C. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Hormozdiari, F. et al. Colocalization of GWAS and eQTL signals detects target genes. Am. J. Hum. Genet. 99, 1245–1260 (2016).
Wallace, C. Eliciting priors and relaxing the single causal variant assumption in colocalisation analyses. PLoS Genet. 16, e1008720 (2020).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Acknowledgements
Our study was supported by the National Institutes of Health (NIH) for Integrative Analysis of Lung Cancer Etiology and Risk (U19CA203654) and Sequencing Familial Lung Cancer (R01CA243483). C.I.A. is a Research Scholar of the Cancer Prevention Research Interest of Texas (CPRIT) award (RR170048). Functional studies were partially supported by NIH grants (R01CA250905 (S.M.R), CPRIT RR170048 (C.I.A) and DP1-AG072751 (S.M.R.)). This project was supported by the Cytometry and Cell Sorting Core at Baylor College of Medicine with funding from the CPRIT Core Facility Support Award (CPRIT RP180672) and the NIH (CA125123 and RR024574) as well as the assistance of J.M. Sederstrom. The Resource for the Study of Lung Cancer Epidemiology in North Trent (ReSoLuCENT) study was funded by the Sheffield Hospitals Charity, Sheffield Experimental Cancer Medicine Centre and Weston Park Hospital Cancer Charity. F.T. was supported by a clinical PhD fellowship funded by the Yorkshire Cancer Research/Cancer Research UK Sheffield Cancer Centre. D.M. was supported by Department of Health and Human Services contracts HHSN26820100007C, HHSN268201700012C and 75N92020C00001. J.E.B. was supported by the Intramural Research Program of the National Human Genome Research Institute, NIH. R.W.P. was supported by NIH T32ES027801. J.X. was supported by the National Institute of Environmental Health Sciences of the NIH under Award Number K99ES033259. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH. This work was supported by the Intramural Research Program of the Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH. We acknowledge the participants and investigators of INTEGRAL-ILCCO Consortium, Genetic Epidemiology of Lung Cancer Consortium (GELCC), FinnGen study and Kaiser Permanente Research Bank (KPRB) Genetic Epidemiology Research on Adult Health and Aging (GERA) cohort study.
Author information
Authors and Affiliations
Contributions
J.B., Y. Han, Y. Li and C.I.A. conceived and designed the study. Y. Han and X.X. acquired the data. Y. Han, E.L., J.C. and X.X. performed the analysis. J.X. and Z.S. performed experimental validation. M.Z., W. Zhou, R.S., A.S., C.L., T.R., L.K. and L.C.S. provided substantial support on validation study. J.B., Y. Han, Y. Li, J.X., E.L., J.C. and C.I.A. interpreted the results. J.B., J.X., E.L. and J.C. wrote the first draft of the manuscript. J.B., Y. Han, J.X., E.L., J.C. and X.X. provided supplementary materials. J.B. and C.I.A. provided supervision and contributed to analyses. All authors reviewed and commented on the manuscript and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 LocusZoom regional plots of newly identified cross-ancestry genetic variants.
Newly identified cross-ancestry variant is colored in purple, and colors of other dots indicate linkage disequilibrium measure r2 with the lead variant in purple. a,b, Regional association plots at the CYP8B1 (a) and IRF4 locus (b) in overall lung cancer (Lung). c, Regional association plot at the ACTR2 locus in lung adenocarcinoma (ADE). d, Regional association plot at the LINC01122 locus in lung squamous cell carcinoma (SQC). e, Regional association plot at the IL17RC locus in small cell lung cancer (SCC).
Extended Data Fig. 2 Gating strategies for DNA damage assays.
a–c, Gating strategy, associated with Fig. 3a. (d) histograms of γH2AX in EmGFP-FUBP1 and EmGFP-Tubulin overproducing cells. e–g, Gating strategy, associated with Fig. 3b. h–j, Gating strategy, associated with methods: flow-cytometric DNA damage assays, Q2/Q2 + Q3 calculation in overproduction experiments.
Extended Data Fig. 3 Inference of ancestry membership in three intercontinental populations using FastPop.
The colored points in grey indicate 70,639 individuals from diverse populations. Those in orange, green, and blue denote HapMap 3 samples with European (CEU), East Asian (CHB), African (YRI) ancestry, respectively.
Supplementary information
Supplementary Information
Supplementary Note, Tables 25 and 26 and Figs. 1–4.
Supplementary Tables
Supplementary Tables 1–24.
Rights and permissions
About this article
Cite this article
Byun, J., Han, Y., Li, Y. et al. Cross-ancestry genome-wide meta-analysis of 61,047 cases and 947,237 controls identifies new susceptibility loci contributing to lung cancer. Nat Genet 54, 1167–1177 (2022). https://doi.org/10.1038/s41588-022-01115-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-022-01115-x
This article is cited by
-
Plasma metabolites and risk of seven cancers: a two-sample Mendelian randomization study among European descendants
BMC Medicine (2024)
-
Systematic proteome-wide Mendelian randomization using the human plasma proteome to identify therapeutic targets for lung adenocarcinoma
Journal of Translational Medicine (2024)
-
Impact of individual level uncertainty of lung cancer polygenic risk score (PRS) on risk stratification
Genome Medicine (2024)
-
Age at lung cancer diagnosis in females versus males who never smoke by race and ethnicity
British Journal of Cancer (2024)
-
The role of admixture in the rare variant contribution to inflammatory bowel disease
Genome Medicine (2023)