Abstract
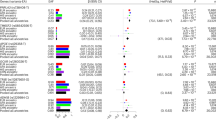
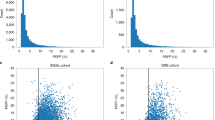
Nonalcoholic fatty liver disease (NAFLD) is a growing cause of chronic liver disease. Using a proxy NAFLD definition of chronic elevation of alanine aminotransferase (cALT) levels without other liver diseases, we performed a multiancestry genome-wide association study (GWAS) in the Million Veteran Program (MVP) including 90,408 cALT cases and 128,187 controls. Seventy-seven loci exceeded genome-wide significance, including 25 without prior NAFLD or alanine aminotransferase associations, with one additional locus identified in European American-only and two in African American-only analyses (P < 5 × 10−8). External replication in histology-defined NAFLD cohorts (7,397 cases and 56,785 controls) or radiologic imaging cohorts (n = 44,289) replicated 17 single-nucleotide polymorphisms (SNPs) (P < 6.5 × 10−4), of which 9 were new (TRIB1, PPARG, MTTP, SERPINA1, FTO, IL1RN, COBLL1, APOH and IFI30). Pleiotropy analysis showed that 61 of 77 multiancestry and all 17 replicated SNPs were jointly associated with metabolic and/or inflammatory traits, revealing a complex model of genetic architecture. Our approach integrating cALT, histology and imaging reveals new insights into genetic liability to NAFLD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The full summary-level association data from the multiancestry, EA, AA, HISP and ASN analyses from this report are available through dbGAP under accession number phs001672.v7.p1 (Veterans Administration MVP Summary Results from Omics Studies). Source data are provided with this paper.
Code availability
Imputation was performed using MiniMac4 and EAGLE v2. Association analysis was performed using PLINK2a. Post-GWAS processing software includes LD Hub v1.9.3, METAL v2011-03-25, DEPICT v140721, LDSC v1.0, GREGOR v4.0, HiCUP v0.8, STRING v11 and Ensembl Variant Effect Predictor with assembly GRCh37.p13 as outlined in Methods. Clear code for analysis is available at the associated website of each software package. Additional analyses were performed in R-4.1, Bioconductor v3.140 and R packages corrcoverage, CHiCAGO and OmnipathR, for which code can be found in their associated vignettes.
Change history
02 March 2023
In the PDF version of this article originally published, the second through fourth text columns of Methods text were interchanged and have now been corrected in the PDF version of the paper online.
References
Asrani, S. K., Devarbhavi, H., Eaton, J. & Kamath, P. S. Burden of liver diseases in the world. J. Hepatol. 70, 151–171 (2019).
Younossi, Z., Anstee, Q. M. & Marietti, M. Global burden of NAFLD and NASH: trends, predictions, risk factors and prevention. Nat. Rev. Gastroenterol. Hepatol. 15, 11–20 (2018).
Estes, C., Razavi, H., Loomba, R., Younossi, Z. & Sanyal, A. J. Modeling the epidemic of nonalcoholic fatty liver disease demonstrates an exponential increase in burden of disease. Hepatology 67, 123–133 (2018).
Carr, R. M., Oranu, A. & Khungar, V. Nonalcoholic fatty liver disease: pathophysiology and management. Gastroenterol. Clin. North Am. 45, 639–652 (2016).
Chalasani, N. et al. The diagnosis and management of nonalcoholic fatty liver disease: practice guidance from the American Association for the Study of Liver Diseases. Hepatology 67, 328–357 (2018).
Sookoian, S. & Pirola, C. J. Genetic predisposition in nonalcoholic fatty liver disease. Clin. Mol. Hepatol. 23, 1–12 (2017).
Romeo, S. et al. Genetic variation in PNPLA3 confers susceptibility to nonalcoholic fatty liver disease. Nat. Genet. 40, 1461–1465 (2008).
Chambers, J. C. et al. Genome-wide association study identifies loci influencing concentrations of liver enzymes in plasma. Nat. Genet. 43, 1131–1138 (2011).
Speliotes, E. K. et al. Genome-wide association analysis identifies variants associated with nonalcoholic fatty liver disease that have distinct effects on metabolic traits. PLoS Genet. 7, e1001324 (2011).
Emdin, C. A. et al. A missense variant in mitochondrial amidoxime reducing component 1 gene and protection against liver disease. PLoS Genet. 16, e1008629 (2020).
Anstee, Q. M. et al. Genome-wide association study of non-alcoholic fatty liver and steatohepatitis in a histologically characterised cohort. J. Hepatol. 73, 505–515 (2020).
Gaziano, J. M. et al. Million Veteran Program: a mega-biobank to study genetic influences on health and disease. J. Clin. Epidemiol. 70, 214–223 (2016).
Husain, N. et al. Nonalcoholic fatty liver disease (NAFLD) in the Veterans Administration population: development and validation of an algorithm for NAFLD using automated data. Aliment Pharm. Ther. 40, 949–954 (2014).
Serper, M. et al. Validating a non-invasive non-alcoholic fatty liver phenotype in the Million Veteran Program. PLoS One 15, e0237430 (2020).
de Vries, P. S. et al. Multiancestry genome-wide association study of lipid levels incorporating gene-alcohol interactions. Am. J. Epidemiol. 188, 1033–1054 (2019).
Kozlitina, J. et al. Exome-wide association study identifies a TM6SF2 variant that confers susceptibility to nonalcoholic fatty liver disease. Nat. Genet. 46, 352–356 (2014).
Abul-Husn, N. S. et al. A protein-truncating HSD17B13 variant and protection from chronic liver disease. N. Engl. J. Med. 378, 1096–1106 (2018).
Young, K. A. et al. Genome-wide association study identifies loci for liver enzyme concentrations in Mexican Americans: the GUARDIAN Consortium. Obes. (Silver Spring) 27, 1331–1337 (2019).
Namjou, B. et al. GWAS and enrichment analyses of non-alcoholic fatty liver disease identify new trait-associated genes and pathways across eMERGE Network. BMC Med. 17, 135 (2019).
Chalasani, N. et al. Genome-wide association study identifies variants associated with histologic features of nonalcoholic Fatty liver disease. Gastroenterology 139, 1567–1576 (2010). 1576 e1-6.
Chen, V. L. et al. Genome-wide association study of serum liver enzymes implicates diverse metabolic and liver pathology. Nat. Commun. 12, 816 (2021).
Pazoki, R. et al. Genetic analysis in European ancestry individuals identifies 517 loci associated with liver enzymes. Nat. Commun. 12, 2579 (2021).
Stephens, C. R. et al. The impact of education and age on metabolic disorders. Front Public Health 8, 180 (2020).
Wakefield, J. Bayes factors for genome-wide association studies: comparison with P-values. Genet. Epidemiol. 33, 79–86 (2009).
Baxter, M. et al. Phenotypic and functional analyses show stem cell-derived hepatocyte-like cells better mimic fetal rather than adult hepatocytes. J. Hepatol. 62, 581–589 (2015).
Brancale, J. & Vilarinho, S. A single cell gene expression atlas of 28 human livers. J. Hepatol. 75, 219–220 (2021).
Goldstein, J. A. et al. LabWAS: novel findings and study design recommendations from a meta-analysis of clinical labs in two independent biobanks. PLoS Genet. 16, e1009077 (2020).
Sliz, E. et al. NAFLD risk alleles in PNPLA3, TM6SF2, GCKR and LYPLAL1 show divergent metabolic effects. Hum. Mol. Genet 27, 2214–2223 (2018).
Stender, S. et al. Relationship between genetic variation at PPP1R3B and levels of liver glycogen and triglyceride. Hepatology 67, 2182–2195 (2018).
Mehta, M. B. et al. Hepatic protein phosphatase 1 regulatory subunit 3B (Ppp1r3b) promotes hepatic glycogen synthesis and thereby regulates fasting energy homeostasis. J. Biol. Chem. 292, 10444–10454 (2017).
Brouwers, M., Jacobs, C., Bast, A., Stehouwer, C. D. A. & Schaper, N. C. Modulation of glucokinase regulatory protein: a double-edged sword? Trends Mol. Med. 21, 583–594 (2015).
Fagerberg, L. et al. Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol. Cell Proteom. 13, 397–406 (2014).
Duff, M. O. et al. Genome-wide identification of zero nucleotide recursive splicing in Drosophila. Nature 521, 376–379 (2015).
Jamialahmadi, O. et al. Exome-wide association study on alanine aminotransferase identifies sequence variants in the GPAM and APOE associated with fatty liver disease. Gastroenterology 160, 1634–1646 e7 (2021).
Hammond, L. E. et al. Mitochondrial glycerol-3-phosphate acyltransferase-deficient mice have reduced weight and liver triacylglycerol content and altered glycerolipid fatty acid composition. Mol. Cell. Biol. 22, 8204–8214 (2002).
Cuchel, M. et al. Inhibition of microsomal triglyceride transfer protein in familial hypercholesterolemia. N. Engl. J. Med. 356, 148–156 (2007).
Soubeyrand, S., Martinuk, A. & McPherson, R. TRIB1 is a positive regulator of hepatocyte nuclear factor 4-alpha. Sci. Rep. 7, 5574 (2017).
Laudadio, I. et al. A feedback loop between the liver-enriched transcription factor network and miR-122 controls hepatocyte differentiation. Gastroenterology 142, 119–129 (2012).
Kim, J. Y., Han, Y. H., Nam, M. W., Kim, H. J. & Lee, M. O. RORalpha suppresses interleukin-6-mediated hepatic acute phase response. Sci. Rep. 9, 11798 (2019).
Laatsch, A. et al. Low density lipoprotein receptor-related protein 1 dependent endosomal trapping and recycling of apolipoprotein E. PLoS One 7, e29385 (2012).
Sanyal, A. J. et al. Pioglitazone, vitamin E, or placebo for nonalcoholic steatohepatitis. N. Engl. J. Med. 362, 1675–1685 (2010).
Musso, G., Cassader, M., Paschetta, E. & Gambino, R. Thiazolidinediones and advanced liver fibrosis in nonalcoholic steatohepatitis: a meta-analysis. JAMA Intern Med. 177, 633–640 (2017).
Ratziu, V. et al. Long-term efficacy of rosiglitazone in nonalcoholic steatohepatitis: results of the fatty liver improvement by rosiglitazone therapy (FLIRT 2) extension trial. Hepatology 51, 445–453 (2010).
Cusi, K. et al. Long-term pioglitazone treatment for patients with nonalcoholic steatohepatitis and prediabetes or type 2 diabetes mellitus: a randomized trial. Ann. Intern Med. 165, 305–315 (2016).
Tilg, H., Adolph, T. E. & Moschen, A. R. Multiple parallel hits hypothesis in nonalcoholic fatty liver disease: revisited after a decade. Hepatology 73, 833–842 (2021).
Hamada, M., Tsunakawa, Y., Jeon, H., Yadav, M. K. & Takahashi, S. Role of MafB in macrophages. Exp. Anim. 69, 1–10 (2020).
Klarin, D. et al. Genetics of blood lipids among ~300,000 multi-ethnic participants of the Million Veteran Program. Nat. Genet. 50, 1514–1523 (2018).
Stoffel, W. et al. Obesity resistance and deregulation of lipogenesis in Delta6-fatty acid desaturase (FADS2) deficiency. EMBO Rep. 15, 110–120 (2014).
Mirea, A. M., Tack, C. J., Chavakis, T., Joosten, L. A. B. & Toonen, E. J. M. IL-1 family cytokine pathways underlying NAFLD: towards new treatment strategies. Trends Mol. Med. 24, 458–471 (2018).
Miao, Z. et al. Identification of 90 NAFLD GWAS loci and establishment of NAFLD PRS and causal role of NAFLD in coronary artery disease. HGG Adv. 3, 100056 (2022).
Kranzler, H. R. et al. Genome-wide association study of alcohol consumption and use disorder in 274,424 individuals from multiple populations. Nat. Commun. 10, 1499 (2019).
Justice, A. C. et al. AUDIT-C and ICD codes as phenotypes for harmful alcohol use: association with ADH1B polymorphisms in two US populations. Addiction 113, 2214–2224 (2018).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Hutchinson, A., Watson, H. & Wallace, C. Improving the coverage of credible sets in Bayesian genetic fine-mapping. PLoS Comput. Biol. 16, e1007829 (2020).
MacLean, M. T. et al. Quantification of abdominal fat from computed tomography using deep learning and its association with electronic health records in an academic biobank. J. Am. Med. Inform. Assoc. 28, 1178–1187 (2021).
Haas, M. E. et al. Machine learning enables new insights into genetic contributions to liver fat accumulation. Cell Genom. 1, 100066 (2021).
Wattacheril, J. et al. Genome-wide associations related to hepatic histology in nonalcoholic fatty liver disease in Hispanic boys. J. Pediatr. 190, 100–107 (2017).
Patton, H. M. et al. Clinical correlates of histopathology in pediatric nonalcoholic steatohepatitis. Gastroenterology 135, 1961–1971 (2008).
Lin, H. J. et al. Home use of a compact, 12lead ECG recording system for newborns. J. Electrocardiol. 53, 89–94 (2019).
Weinshilboum, R. M. & Wang, L. Pharmacogenomics: precision medicine and drug response. Mayo Clin. Proc. 92, 1711–1722 (2017).
Simon, J. A. et al. Phenotypic predictors of response to simvastatin therapy among African-Americans and Caucasians: the Cholesterol and Pharmacogenetics (CAP) study. Am. J. Cardiol. 97, 843–850 (2006).
Hardy, T. et al. The European NAFLD Registry: a real-world longitudinal cohort study of nonalcoholic fatty liver disease. Contemp. Clin. Trials 98, 106175 (2020).
Angulo, P. et al. Liver fibrosis, but no other histologic features, is associated with long-term outcomes of patients with nonalcoholic fatty liver disease. Gastroenterology 149, 389–397 (2015).
Dewey, F. E. et al. Distribution and clinical impact of functional variants in 50,726 whole-exome sequences from the DiscovEHR study. Science 354, aaf6814 (2016).
Harrison, S. A. et al. Selonsertib for patients with bridging fibrosis or compensated cirrhosis due to NASH: Results from randomized phase III STELLAR trials. J. Hepatol. 73, 26–39 (2020).
Roden, D. M. et al. Development of a large-scale de-identified DNA biobank to enable personalized medicine. Clin. Pharmacol. Ther. 84, 362–369 (2008).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Roadmap Epigenomics Consortium et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Finucane, H. K. et al. Heritability enrichment of specifically expressed genes identifies disease-relevant tissues and cell types. Nat. Genet. 50, 621–629 (2018).
CARDIoGRAMplusC4D Consortium et al. Large-scale association analysis identifies new risk loci for coronary artery disease. Nat. Genet. 45, 25–33 (2013).
Fehrmann, R. S. et al. Gene expression analysis identifies global gene dosage sensitivity in cancer. Nat. Genet. 47, 115–125 (2015).
Cahoy, J. D. et al. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J. Neurosci. 28, 264–278 (2008).
Heng, T. S. & Painter, M. W., Immunological Genome Project Consortium. The Immunological Genome Project: networks of gene expression in immune cells. Nat. Immunol. 9, 1091–1094 (2008).
Pers, T. H. et al. Biological interpretation of genome-wide association studies using predicted gene functions. Nat. Commun. 6, 5890 (2015).
Schmidt, E. M. et al. GREGOR: evaluating global enrichment of trait-associated variants in epigenomic features using a systematic, data-driven approach. Bioinformatics 31, 2601–2606 (2015).
McLaren, W. et al. The ensembl variant effect predictor. Genome Biol. 17, 122 (2016).
1000 Genomes Project Consortium et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Chesi, A. et al. Genome-scale Capture-C promoter interactions implicate effector genes at GWAS loci for bone mineral density. Nat. Commun. 10, 1260 (2019).
Pashos, E. E. et al. Large, diverse population cohorts of hiPSCs and derived hepatocyte-like cells reveal functional genetic variation at blood lipid-associated loci. Cell Stem Cell 20, 558–570 (2017).
Caliskan, M. et al. Genetic and epigenetic fine mapping of complex trait associated loci in the human liver. Am. J. Hum. Genet 105, 89–107 (2019).
Wingett, S. et al. HiCUP: pipeline for mapping and processing Hi-C data. F1000Res 4, 1310 (2015).
Cairns, J. et al. CHiCAGO: robust detection of DNA looping interactions in Capture Hi-C data. Genome Biol. 17, 127 (2016).
Szklarczyk, D. et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613 (2019).
Gagliano Taliun, S. A. et al. Exploring and visualizing large-scale genetic associations by using PheWeb. Nat. Genet. 52, 550–552 (2020).
Denny, J. C. et al. PheWAS: demonstrating the feasibility of a phenome-wide scan to discover gene-disease associations. Bioinformatics 26, 1205–1210 (2010).
Loh, P. R. et al. Reference-based phasing using the Haplotype Reference Consortium panel. Nat. Genet. 48, 1443–1448 (2016).
Zhou, W. et al. Efficiently controlling for case-control imbalance and sample relatedness in large-scale genetic association studies. Nat. Genet. 50, 1335–1341 (2018).
Elsworth, B. et al. The MRC IEU OpenGWAS data infrastructure. Preprint at bioRxiv, https://doi.org/10.1101/2020.08.10.244293 (2020).
Shin, S. et al. CREB mediates the insulinotropic and anti-apoptotic effects of GLP-1 signaling in adult mouse beta-cells. Mol. Metab. 3, 803–812 (2014).
Kettunen, J. et al. Genome-wide study for circulating metabolites identifies 62 loci and reveals novel systemic effects of LPA. Nat. Commun. 7, 11122 (2016).
Hemani, G. et al. The MR-Base platform supports systematic causal inference across the human phenome. Elife 7, e34408 (2018).
Sun, B. B. et al. Genomic atlas of the human plasma proteome. Nature 558, 73–79 (2018).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Giri, A. et al. Trans-ethnic association study of blood pressure determinants in over 750,000 individuals. Nat. Genet. 51, 51–62 (2019).
Pulit, S. L. et al. Meta-analysis of genome-wide association studies for body fat distribution in 694 649 individuals of European ancestry. Hum. Mol. Genet 28, 166–174 (2019).
Teumer, A. et al. Genome-wide association meta-analyses and fine-mapping elucidate pathways influencing albuminuria. Nat. Commun. 10, 4130 (2019).
Tin, A. et al. Target genes, variants, tissues and transcriptional pathways influencing human serum urate levels. Nat. Genet. 51, 1459–1474 (2019).
Wuttke, M. et al. A catalog of genetic loci associated with kidney function from analyses of a million individuals. Nat. Genet. 51, 957–972 (2019).
Astle, W. J. et al. The allelic landscape of human blood cell trait variation and links to common complex disease. Cell 167, 1415–1429 (2016).
Guo, H. et al. Integration of disease association and eQTL data using a Bayesian colocalisation approach highlights six candidate causal genes in immune-mediated diseases. Hum. Mol. Genet 24, 3305–3313 (2015).
Lambert, S. A. et al. The human transcription factors. Cell 175, 598–599 (2018).
Garcia-Alonso, L., Holland, C. H., Ibrahim, M. M., Turei, D. & Saez-Rodriguez, J. Benchmark and integration of resources for the estimation of human transcription factor activities. Genome Res. 29, 1363–1375 (2019).
Turei, D., Korcsmaros, T. & Saez-Rodriguez, J. OmniPath: guidelines and gateway for literature-curated signaling pathway resources. Nat. Methods 13, 966–967 (2016).
Ceccarelli, F., Turei, D., Gabor, A. & Saez-Rodriguez, J. Bringing data from curated pathway resources to Cytoscape with OmniPath. Bioinformatics 36, 2632–2633 (2020).
Acknowledgements
This research is based on data from the MVP, Office of Research and Development, Veterans Health Administration and was supported by award MVP000. This publication does not represent the views of the Department of Veterans Affairs, the US Food and Drug Administration or the US Government. This research was also supported by funding from the Department of Veterans Affairs awards I01- BX003362 (P.S.T. and K.-M.C.) and I01 BX003341 (H.R.K. co-principal investigator) and the VA Informatics and Computing Infrastructure VA HSR RES 130457 (S.L.D. and J.A.L.). B.F.V. acknowledges support for this work from the National Institutes of Health (NIH)/National Institute of Diabetes and Digestive and Kidney Diseases (grants DK101478 and DK126194) and a Linda Pechenik Montague Investigator award. K.-M.C., S.M.D., J.M.G., C.J.O., L.S.P. and P.S.T. are supported by the VA Cooperative Studies Program. S.M.D. is supported by the VA (IK2 CX001780). Funding support is also acknowledged for M.S. (K23 DK115897), R.M.C. (R01 AA026302), D.E.K. (T32 HL007734), A.D.W. (R01 DK122586, R01 AI123539), J.B.M. (R01 HL151855, UM1 DK078616), W.R.W. (R01 HL137984 P41 EB029460), S.F.A.G. (R01 HD056465), L.B. (R01 LM010685), A.V.K. (K08 HG010155, U01 HG011719) and L.S.P. (VA awards CSP #2008, I01 CX001899, I01 CX001737 and I01 BX005831; NIH awards R01 DK127083, R03 AI133172, R21 AI156161, UL1 TR002378 and P30 DK111024; and Cystic Fibrosis Foundation award PHILLI12A0). The Rader lab was supported by NIH grants HL134853 (N.J.H. and D.J.R.) and DK114291-01A1 (K.T.C., N.J.H. and D.J.R.). We thank all study participants for their contribution. Support for imaging studies was provided by the University of Pennsylvania’s Institute for Translational Medicine and Therapeutics (NIH NCATS UL1 TR001878), the Penn Center for Precision Medicine Accelerator Fund.
Author information
Authors and Affiliations
Consortia
Contributions
M.V., S.R., K.-M.C., D.J.R., B.F.V. and P.S.T. were responsible for the concept and design. M.V., S.R., K.-M.C., D.J.R., B.F.V. and P.S.T. performed data acquisition, analysis or interpretation. M.V., S.R., K.-M.C. and B.F.V. drafted the manuscript. Critical revision of the manuscript for important intellectual content was carried out by M.V., K.M. Lorenz, K.M. Lee, M.S., D.E.K., X.Z., C.T., R.L.K., H.R.K., A.V., A.G., D.M.K., Y.V.S., J.E.H., D.R.M., P.D.R., L.S.P., S.M., S.L.D., J.S.L., T.L.A., S.P., K.C., T.L.E., S.M.D., P.W.W., J.M.G., C.J.O., P.S.T., J.B.M., J.A.L., B.F.V., K.-M.C., Q.M.A., R.D., H.J.C., A.K.D., L.A.L., J.B.N., A.E.L., M.B.J., N.V., A.B., S.R., X.G., J. He, Y.G., C.C., R.P.M., C.V.S., J.P., R.M.C., M.E.H., M.T.M., W.R.W., J. Huang, K.T.C., N.J.H., C.-T.L., M.T.L., J.Y., M.B., J.T., X.L., H.J.L., Y.-D.I.C., K.D.T., R.-K.C., R.M.K., S.V., J.B., K.R.R., B.A.N.-T., J.B.S., A.J.S., N.C., K.A.R., B.D.M., D.G., A.D.W., E.M., Y.S., N.M., A.V.K., S.F.A.G., C.D.B., D.S., L.B., J.I.R. and D.J.R. Finally, K.-M.C., D.J.R. and B.F.V. provided administrative, technical or material support. VA MVP: M.V., K.M. Lorenz, K.M. Lee, M.S., D.E.K., X.Z., C.T., R.L.K., H.R.K., A.V., A.G., D.M.K., Y.V.S., J.E.H., D.R.M., P.D.R., L.S.P., S.M., S.L.D., J.S.L., R.M.C., T.L.A., S.P., K.C., T.L.E., S.M.D., P.W.W., J.M.G., C.J.O., P.S.T., J.B.M., J.A.L., B.F.V. and K.-M.C. Geisinger-Regeneron DiscovEHR Collaboration/Regeneron Genetics Center: L.A.L., J.B.N., A.E.L., M.B.J., N.V. and A.B. EPoS Consortium: Q.M.A., R.D., H.J.C. and A.K.D.
Corresponding authors
Ethics declarations
Competing interests
H.R.K. is a scientific advisory board member for Dicerna Pharmaceuticals, Sophrosyne Pharmaceuticals and Enthion Pharmaceuticals; a consultant for Sobrera Pharmaceuticals; the recipient of research funding and medication supplies for an investigator-initiated study from Alkermes; a member of the American Society of Clinical Psychopharmacology’s Alcohol Clinical Trials Initiative, which during the past 3 years was supported by Alkermes, Amygdala Neurosciences, Arbor Pharmaceuticals, Dicerna, Ethypharm, Indivior, Lundbeck, Mitsubishi and Otsuka; and is named as an inventor on the Patent Cooperation Treaty patent application #15/878,640 entitled ‘Genotype-guided dosing of opioid agonists,’ filed 24 January 2018. D.G. is employed part-time by Novo Nordisk. A.V.K. is an employee and holds equity in Verve Therapeutics; has served as a scientific advisor to Amgen, Third Rock Ventures, Illumina, and Foresite Labs; received a sponsored research agreement from IBM Research; and is listed as a co-inventor on a patent application for use of imaging data in assessing body fat distribution and associated cardiometabolic risk. S.J.A. is President of Sanyal Bio; has stock options in Genfit, Galmed, Exhalenz, Durect, Tiziana, Algernon and Indalo; has served as a consultant to Intercept, Gilead, Bristol Myers Squibb, Novartis, Pfizer, Lilly, Novo Nordisk, AstraZeneca, Medimmune, Merck, Allergan, Albireo, Boehringer Ingelhiem, Celgene, NGM, Glympse, Conatus, Genentech, Tern, Takeda, Hemoshear, Immuron, Surrozen, Poxel, Path AI, Second Genome, Zydus, Chiasma, Surrozen, Poxel, Blade, Pliant, Liposcience, Cymabay, Salix, Ferring and Teva; and his institution has received grants from Intercept, Gilead, Novartis, Merck, AstraZeneca, Malinckrodt, Pfizer, Lilly, Salix and Bristol Myers Squibb. V.C.U. has ownership interests in Sanyal Bio. K.R.R. is on the NASH Advisory Board at Novo Nordisk and receives grant support from TARGET-NASH, Bristol Myers Squibb and Intercept Pharmaceuticals. J.B.N., A.E.L., M.B.J., N.V., A.B., M.E.H. and L.A.L. receive salary, stocks and/or stock options from Regeneron Pharmaceuticals. R.P.M. and C.C. are employees and shareholders of Gilead Sciences. Q.M.A. is coordinator of the EU IMI-2 LITMUS consortium, which is funded by the EU Horizon 2020 program and the European Federation of Pharmaceutical Industries and Associations. Q.M.A. reports research grant funding from Allergan/Tobira, AstraZeneca, Boehringer Ingelheim, GlaxoSmithKline, Glympse Bio, Intercept, Novartis Pharma and Pfizer; Consultancy for 89Bio, Abbvie/Allergan, Akero, Altimentiv, Altimmune, AstraZeneca, Axcella, Blade, BMS, BNN Cardio, Boehringer Ingelheim, Cirius, CymaBay, EcoR1, E3Bio, Eli Lilly & Company, Galmed, Genentech, Genfit, Gilead, Grunthal, HistoIndex, Indalo, Intercept Pharma, Inventiva, IQVIA, Janssen, Johnson & Johnson, Madrigal, MedImmune, Medpace, Merck, Metacrine, NGMBio, North Sea Therapeutics, Novartis, Novo Nordisk, PathAI, Pfizer, Poxel, ProSciento, Raptor Pharma, Roche, Servier, Shionogi, Terns, The Medicines Company and Viking Therapeutics; speaker fees from Abbott Laboratories, Allergan/Tobira, BMS, Clinical Care Options, Falk, Fishawack, Genfit, Gilead, Integritas Communications, Kenes and Medscape; and royalties from Elsevier. S.M.D. receives research support from RenalytixAI and personal consulting fees from Calico Labs outside the scope of the current research. S.L.D. reports grants from Alnylam Pharmaceuticals, Astellas Pharma, AstraZeneca Pharmaceuticals, Biodesix, Boehringer Ingelheim International, Celgene Corporation, Eli Lilly and Company, Genentech, Gilead Sciences, GlaxoSmithKline, Innocrin Pharmaceuticals, IQVIA, Janssen Pharmaceuticals, Kantar Health, MDxHealth, Merck & Co, Myriad Genetic Laboratories, Novartis International and Parexel International Corporation through the University of Utah or Western Institute for Veteran Research outside the submitted work. C.J.O. is an employee of Novartis Institute for Biomedical Research. S.F.A.G. is the Daniel B. Burke Endowed Chair for Diabetes Research. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–14 and note.
Supplementary Table 1
Supplementary Tables 1–33.
Source data
Source Data Fig. 4
The data are all the association of the lead SNPs with other traits, summarized into one file with directions.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Vujkovic, M., Ramdas, S., Lorenz, K.M. et al. A multiancestry genome-wide association study of unexplained chronic ALT elevation as a proxy for nonalcoholic fatty liver disease with histological and radiological validation. Nat Genet 54, 761–771 (2022). https://doi.org/10.1038/s41588-022-01078-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-022-01078-z
This article is cited by
-
Implications of the evolving knowledge of the genetic architecture of MASLD
Nature Reviews Gastroenterology & Hepatology (2024)
-
Metabolic links among milk, genes and gut
Nature Metabolism (2024)
-
Integrative common and rare variant analyses provide insights into the genetic architecture of liver cirrhosis
Nature Genetics (2024)
-
Engineered human hepatocyte organoids enable CRISPR-based target discovery and drug screening for steatosis
Nature Biotechnology (2023)
-
Personalisierte Therapie der metabolisch assoziierten Fettlebererkrankung
Die Gastroenterologie (2023)