Abstract
Transgenic papaya is widely publicized for controlling papaya ringspot virus. However, the impact of particle bombardment on the genome remains unknown. The transgenic SunUp and its progenitor Sunset genomes were assembled into 351.5 and 350.3 Mb in nine chromosomes, respectively. We identified a 1.64 Mb insertion containing three transgenic insertions in SunUp chromosome 5, consisting of 52 nuclear-plastid, 21 nuclear-mitochondrial and 1 nuclear genomic fragments. A 591.9 kb fragment in chromosome 5 was translocated into the 1.64 Mb insertion. We assembled a gapless 9.8 Mb hermaphrodite-specific region of the Yh chromosome and its 6.0 Mb X counterpart. Resequencing 86 genomes revealed three distinct groups, validating their geographic origin and breeding history. We identified 147 selective sweeps and defined the essential role of zeta-carotene desaturase in carotenoid accumulation during domestication. Our findings elucidated the impact of particle bombardment and improved our understanding of sex chromosomes and domestication to expedite papaya improvement.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Nanopore and PacBio whole-genome sequencing data, Hi-C, Illumina data and RNA-seq data have been deposited in the NCBI Sequence Read Archive (SRA) database as Bioproject PRJNA727683. The SunUp and Sunset genome assemblies were archived in the NCBI Genome database under the accession number JAIUCH000000000 for SunUp and JAIUCG000000000 for Sunset genome. The SunUp and Sunset genome assemblies and gene annotations have been also deposited in the Genome Warehouse (GWH) database in BIG data Center (https://ngdc.cncb.ac.cn/gwh/) under accession number GWHBFSC00000000 for SunUp genome and GWHBFSD00000000 for Sunset genome. VCF file that contains all clean SNPs was uploaded to the Mendeley database (https://data.mendeley.com/datasets/m5phbmw43c/1). The papaya sex-specific small RNA sequence data used can be obtained from NCBI’s Gene Expression Omnibus (GEO) under accession number GSE54097. Source data are provided with this paper.
Code availability
The Python scripts ‘filter_blast_result.sh’, ‘convert_bed_for_junction.py’, ‘extract_result.py’ and ‘plot_data_V8.py’ for Sunset and SunUp-specific nuclear organelle DNA junction sites identification were available at GitHub (https://github.com/sc-zhang/CUFscripts/tree/v1.0). These codes were also archived on Zenodo with the https://doi.org/10.5281/zenodo.6342427.
References
Liebman, B. Nutritional aspects of fruit. Nutrition Action Healthletter 1, 10–11 (1992).
Chandrika, U. G., Jansz, E. R., Wickramasinghe, S. M. D. N. & Warnasuriya, N. D. Carotenoids in yellow- and red-fleshed papaya (Carica papaya L). J. Sci. Food Agric. 83, 1279–1282 (2003).
Fuentes, G. & Santamaría, J. M. in Genetics and Genomics of Papaya (eds Ming, R. & Moore, P. H.) 3–15 (Springer, 2014).
Manshardt, R. in Genetics and Genomics of Papaya (eds Ming, R. & Moore, P. H.) 95–113 (Springer, 2014).
Liu, Z. et al. A primitive Y chromosome in papaya marks incipient sex chromosome evolution. Nature 427, 348–352 (2004).
Wang, J. et al. Sequencing papaya X and Yh chromosomes reveals molecular basis of incipient sex chromosome evolution. Proc. Natl Acad. Sci. USA 109, 13710–13715 (2012).
VanBuren, R. et al. Origin and domestication of papaya Yh chromosome. Genome Res. 25, 524–533 (2015).
Gonsalves, D. Control of papaya ringspot virus in papaya: a case study. Annu. Rev. Phytopathol. 36, 415–437 (1998).
Manshardt, R. UH Rainbow’ Papaya. Germplasm, G-1 (University of Hawaii College of Tropical Agriculture and Human Resources, 1998).
Fitch, M. M., Manshardt, R. M., Gonsalves, D., Slightom, J. L. & Sanford, J. C. Virus resistant papaya plants derived from tissues bombarded with the coat protein gene of papaya ringspot virus. Nat. Biotechnol. 10, 1466–1472 (1992).
Fitch, M. M., Manshardt, R. M., Gonsalves, D., Slightom, J. L. & Sanford, J. C. Stable transformation of papaya via microprojectile bombardment. Plant Cell Rep. 9, 189–194 (1990).
Kawakatsu, T., Kawahara, Y., Itoh, T. & Takaiwa, F. A whole-genome analysis of a transgenic rice seed-based edible vaccine against cedar pollen allergy. DNA Res. 20, 623–631 (2013).
Suzuki, J. Y. et al. Characterization of insertion sites in Rainbow papaya, the first commercialized transgenic fruit crop. Tropical Plant Biol. 1, 293–309 (2008).
Ming, R. et al. The draft genome of the transgenic tropical fruit tree papaya (Carica papaya Linnaeus). Nature 452, 991–996 (2008).
Jain, M. et al. Nanopore sequencing and assembly of a human genome with ultra-long reads. Nat. Biotechnol. 36, 338–345 (2018).
Zhang, X., Zhang, S., Zhao, Q., Ming, R. & Tang, H. Assembly of allele-aware, chromosomal-scale autopolyploid genomes based on Hi-C data. Nat. Plants 5, 833–845 (2019).
Na, J. K. et al. Construction of physical maps for the sex-specific regions of papaya sex chromosomes. BMC Genom. 13, 176 (2012).
Zhou, L., Christopher, D. A. & Paull, R. E. Defoliation and fruit removal effects on papaya fruit production, sugar accumulation, and sucrose metabolism. J. Am. Soc. Hortic. Sci. 125, 644–652 (2000).
Klein, T. M., Wolf, E. D., Wu, R. & Sanford, J. C. High-velocity microprojectiles for delivering nucleic acids into living cells. Biotechnology 24, 384–386 (1992).
Hirochika, H. Activation of tobacco retrotransposons during tissue culture. EMBO J. 12, 2521–2528 (1993).
Hirochika, H. Retrotransposons of rice: their regulation and use for genome analysis. Plant Mol. Biol. 35, 231–240 (1997).
Lisch, D. How important are transposons for plant evolution? Nat. Rev. Genet. 14, 49–61 (2012).
Miguel, C. & Marum, L. An epigenetic view of plant cells cultured in vitro: somaclonal variation and beyond. J. Exp. Bot. 62, 3713–3725 (2011).
Chen, S. et al. Distribution and characterization of over 1000 T-DNA tags in rice genome. Plant J. 36, 105–113 (2003).
Sawasaki, T., Takahashi, M., Goshima, N. & Morikawa, H. Structures of transgene loci in transgenic Arabidopsis plants obtained by particle bombardment: junction regions can bind to nuclear matrices. Gene 218, 27–35 (1998).
Stegemann, S., Hartmann, S., Ruf, S. & Bock, R. High-frequency gene transfer from the chloroplast genome to the nucleus. Proc. Natl Acad. Sci. USA 100, 8828–8833 (2003).
Ma, H. et al. High-density linkage mapping revealed suppression of recombination at the sex determination locus in papaya. Genetics 166, 419–436 (2004).
Lin, T. et al. Genomic analyses provide insights into the history of tomato breeding. Nat. Genet. 46, 1220–1226 (2014).
Sun, X. et al. Phased diploid genome assemblies and pan-genomes provide insights into the genetic history of apple domestication. Nat. Genet. 52, 1423–1432 (2020).
Wu, G. A. et al. Genomics of the origin and evolution of Citrus. Nature 554, 311–316 (2018).
Zerpa-Catanho, D., Zhang, X., Song, J., Hernandez, A. G. & Ming, R. Ultra-long DNA molecule isolation from plant nuclei for ultra-long read genome sequencing. STAR Protoc. 2, 100343 (2021).
Xie, T. et al. De novo plant genome assembly based on chromatin interactions: a case study of Arabidopsis thaliana. Mol. Plant 8, 489–492 (2015).
Zhang, X. et al. Genomes of the Banyan tree and pollinator wasp provide insights into fig–wasp coevolution. Cell 183, 875–889 (2020).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963 (2014).
Tamazian, G. et al. Chromosomer: a reference-based genome arrangement tool for producing draft chromosome sequences. Gigascience 5, 38 (2016).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95 (2017).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 21, 245 (2020).
Rhie, A. et al. Towards complete and error-free genome assemblies of all vertebrate species. Nature 592, 737–746 (2021).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Bao, Z. & Eddy, S. R. Automated de novo identification of repeat sequence families in sequenced genomes. Genome Res. 12, 1269–1276 (2002).
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics 21, i351–i358 (2005).
Abrusán, G., Grundmann, N., DeMester, L. & Makalowski, W. TEclass–a tool for automated classification of unknown eukaryotic transposable elements. Bioinformatics 25, 1329–1330 (2009).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Wu, T. D. & Watanabe, C. K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875 (2005).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Yang, X. & Li, L. miRDeep-P: a computational tool for analyzing the microRNA transcriptome in plants. Bioinformatics 27, 2614–2615 (2011).
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49 (2012).
Buels, R. et al. JBrowse: a dynamic web platform for genome visualization and analysis. Genome Biol. 17, 66 (2016).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Sedlazeck, F. J. et al. Accurate detection of complex structural variations using single-molecule sequencing. Nat. Methods 15, 461–468 (2018).
Marçais, G. et al. MUMmer4: A fast and versatile genome alignment system. PLoS Comput. Biol. 14, e1005944 (2018).
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Brown, J. E., Bauman, J. M., Lawrie, J. F., Rocha, O. J. & Moore, R. C. The structure of morphological and genetic diversity in natural populations of Carica papaya (Caricaceae) in Costa Rica. Biotropica 44, 179–188 (2012).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
VanBuren, R. et al. Extremely low nucleotide diversity in the X-linked region of papaya caused by a strong selective sweep. Genome Biol. 17, 230 (2016).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6, 80–92 (2012).
Lee, T.-H., Guo, H., Wang, X., Kim, C. & Paterson, A. H. SNPhylo: a pipeline to construct a phylogenetic tree from huge SNP data. BMC Genom. 15, 162 (2014).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Falush, D., Stephens, M. & Pritchard, J. K. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164, 1567–1587 (2003).
Chen, L. Y. et al. The bracteatus pineapple genome and domestication of clonally propagated crops. Nat. Genet. 51, 1549–1558 (2019).
Cockerham, C. C. & Weir, B. S. Covariances of relatives stemming from a population undergoing mixed self and random mating. Biometrics 40, 157–164 (1984).
Acknowledgements
This work was supported by US National Science Foundation (NSF) Plant Genome Research Program Award (DBI-1546890 to R.M.), National Natural Science Foundation of China grants (31701889 to J.Y.) and Natural Science Foundation of Fujian Province grants (2018J01601 to J.Y. and 2018J01604 to Xingtan Zhang). The Science and Technology Innovation Fund of Fujian Agriculture and Forestry University (CXZX2020091A to J.Y).
Author information
Authors and Affiliations
Contributions
R.M. conceived this genome project and coordinated the research activities. R.M., J.Y., R.V. and Q.Y. designed the experiments. J.Y., R.V., Juan Liu and J.F. carried out the experiments. Xingtan Zhang., X.X., H. Yan and Z. Liao assembled SunUp and Sunset genomes. J.Y., Z. Lin., B.D., Xiaobing Zhao., Xiaodan Zhang and F.D. processed genome sequencing and resequencing data. J.Y., Juan Liu., Q.Y., M.Y., Xingtan Zhang., Y.H., Z. Liao and J.F. analyzed transgenic insertions. J.Y., Jing Lin., X.X., S.Z., P.Z., Y.M., H. Yu., J.F., M.F., J.Z., H.T. and M.-L.W. analyzed SunUp and Sunset genome and RNA-seq data. R.V., C.M.W., S.C., N.J.C., R.E.P., E.M.-N., P.Q.-R., A.B., V.M.J., X.M. and D.Z.-C. maintained plant materials and analyzed resequenced genomes. J.Y., R.V. Juan Liu., J.F. and R.M. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Aureliano Bombarely, Jordi Garcia-Mas and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
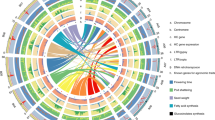
Extended Data Fig. 1 Genome-wide analysis of chromatin interactions in papaya SunUp and Sunset genomes.
Hi-C interactions heatmap diagram at 500Kb resolutions for the entire nine chromosomes and HSY region were showed from top left to bottom right in order (1–9, indicate Chr1-Chr9, HSY indicates hermaphrodite-specific Yh chromosome region). The color bars beside heat maps indicate strong interactions in red and weak interactions in yellow.
Extended Data Fig. 2 PCR validation of transgenic insertion sites in Sunset.
a, The schematic diagram for the 1.64 Mb insertion in SunUp. Primers were designed to amplify the insertion loci P3, and sequences of both sides were defined as P1 and P2 in SunUp and Sunset. b, The insertion loci P3 can be amplified in SunUp but not in Sunset. PCR products for band P3 were confirmed by sanger-sequencing, and the result was consistent with assembly sequence. The experiment was repeated three times and the same results were gotten. c-d, The dot plot for both border sequences in SunUp and Sunset which were marked as Sunset P1/P2 and SunUp P1/P2.
Extended Data Fig. 3 Validation of the nuclear sequences located on 1.64 Mb insertion by Nanopore reads in SunUp.
The nuclear DNA fragment located in Chr5: 37.29-37.31 Mb of SunUp was mapped to the Sunset genome with the corresponding region in Chr5: 37.04-37.06 Mb of Sunset. The assembly of the nuclear DNA fragment was verified by Nanopore reads. SunUp Nanopore reads were aligned against the SunUp reference genome using the BWA software, and the result was visualized by Jbrowse.
Extended Data Fig. 4 Pipeline of identification of SunUp-specific genomic integration of nuclear organelle DNA fragments.
a, Quality control of raw sequenced data. b, Search for SunUp nuclear organelle junction sites by BLASTN. c, Alignment between Sunset reads and SunUp reference genome. Unmapped reads were removed in further analysis. d, Nuclear organelle junction sites shared by SunUp and Sunset. A junction site was supposed to be shared by SunUp and Sunset genomes when there were mapped to reads and spanning its position in the SunUp reference genome. e, Extraction of shared reliable junction sites. f, Junction sites specific in SunUp.
Extended Data Fig. 5 Comparison of the 591.9 Kb deletion with SunUp, chloroplast and mitochondria genomic sequences by using BLASTN.
The 591.9 Kb deletion (Chr5: 22.0-22.6 Mb) in Sunset was aligned to SunUp Chr5, chloroplast and mitochondrial sequences. The plot indicates that the 591.9 Kb deletion has the corresponding region in SunUp Chr5: 36.4-38.1 Mb, chloroplast and mitochondrial sequences.
Extended Data Fig. 6 Model of integration of exogenous DNA into the papaya genome.
By comparison between SunUp and its host Sunset genome, our results supported a precisely regulated model that essential events may be involved in integration of exogenous DNA into the plant genome via particle bombardment. Three transgenic insertions were integrated into a single site locus. Particle bombardment mediated transformation brought foreign DNA fragments into the NUPT-rich region, accompanied by the integration of chloroplast and mitochondrial genome fragments into nuclear genome and the translocation and rearrangement of the original NUPTs and NUMTs. Organellar DNA fragments were transferred to nuclear genome by the driving force. This force may be penetration of cells with DNA-coated metal particles that elicits a wound response. This wound response would activate DNA repair and degradation enzymes on the introduced DNA, making transgenic sequences and NUPTs and NUMTs rearrangement.
Extended Data Fig. 7 PCR validation of transgenic insertions.
Three transgenic insertions, 9789 bp functional insertion including CP, 1533 bp tetA insertion and 290 bp nptII insertion were amplified in Sunset and SunUp. Three insertions can be amplified in SunUp but not in Sunset by using the 5 primer pairs listed in Supplementary table 66. The experiment was repeated three times and the same results were obtained. PCR products were confirmed by Sanger-sequencing, and the result was used to correct the assembly sequence.
Extended Data Fig. 8 Validation of the insertion sites by PacBio and Nanopore reads in SunUp.
Aligning the PacBio reads and Nanopore reads of SunUp and Sunset against the transgenic function fragment using BWA software, and the results were shown by Jbrowse. Part of SunUp reads were displayed on the region. No Sunset reads were spanned the functional insertion.
Extended Data Fig. 9 The position of PCR-amplified regions on the insertion.
We designed 65 pairs of primers which were randomly distributed on the 1.64 Mb insertion of SunUp (SunUp Chr5: 36.42-38.06 Mb) to amplify insertion sequences in SunUp and Sunset. The information of primers were shown in Supplementary table 66. Thick lines are shown on the insertion region, with blue and red lines indicating amplified bands. All blue and red bands can be amplified in SunUp genome, and blue lines indicate SunUp-specific fragments that can be only amplified in SunUp not in Sunset.
Supplementary information
Supplementary Information
Supplementary Tables 1–11, 17, 22–29, 33–35, 37–39, 41, 42, 44, 47, 50, 51, 57, 60, 62–65, 67 and 68, and Figs. 1–11.
Supplementary Tables
Supplementary Tables 12–16, 18–21, 30–32, 36–40, 43, 45, 46, 48, 49, 52–56, 58, 59, 61 and 66.
Source data
Source Data Fig. 1
Unprocessed gels.
Source Data Fig. 2
Unprocessed gels.
Rights and permissions
About this article
Cite this article
Yue, J., VanBuren, R., Liu, J. et al. SunUp and Sunset genomes revealed impact of particle bombardment mediated transformation and domestication history in papaya. Nat Genet 54, 715–724 (2022). https://doi.org/10.1038/s41588-022-01068-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-022-01068-1
This article is cited by
-
Comparative proteomic analysis of papaya bud flowers reveals metabolic signatures and pathways driving hermaphrodite development
Scientific Reports (2024)
-
The need for assessment of risks arising from interactions between NGT organisms from an EU perspective
Environmental Sciences Europe (2023)
-
Haplotype-resolved chromosomal-level genome assembly of Buzhaye (Microcos paniculata)
Scientific Data (2023)