Abstract
Epigenetic inheritance of gene expression states enables a single genome to maintain distinct cellular identities. How histone modifications contribute to this process remains unclear. Using global chromatin perturbations and local, time-controlled modulation of transcription, we establish the existence of epigenetic memory of transcriptional activation for genes that can be silenced by the Polycomb group. This property emerges during cell differentiation and allows genes to be stably switched after a transient transcriptional stimulus. This transcriptional memory state at Polycomb targets operates in cis; however, rather than relying solely on read-and-write propagation of histone modifications, the memory is also linked to the strength of activating inputs opposing Polycomb proteins, and therefore varies with the cellular context. Our data and computational simulations suggest a model whereby transcriptional memory arises from double-negative feedback between Polycomb-mediated silencing and active transcription. Transcriptional memory at Polycomb targets thus depends not only on histone modifications but also on the gene-regulatory network and underlying identity of a cell.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The MS PRM data that support the findings of the present study have been deposited in the ProteomeXchange Consortium via the PRIDE partner repository with the dataset accession no. PXD023966. The NGS data that support the findings of the present study have been deposited in the Gene Expression Omnibus under accession no. GSE147568. Source data are provided with this paper.
Code availability
Customized code used for computational simulations is available on GitHub: https://github.com/ceclov/PRC2-transcription-model
References
Bonasio, R., Tu, S. & Reinberg, D. Molecular signals of epigenetic states. Science 330, 612–616 (2010).
D’Urso, A. & Brickner, J. H. Mechanisms of epigenetic memory. Trends Genet. 30, 230–236 (2014).
Moazed, D. Mechanisms for the inheritance of chromatin States. Cell 146, 510–518 (2011).
Pisco, A. O. & d’Herouel, A. F. Conceptual confusion: the case of epigenetics. Preprint at bioRxiv https://doi.org/10.13140/RG.2.1.3726.4249 (2016).
Bostick, M. et al. UHRF1 plays a role in maintaining DNA methylation in mammalian cells. Science 317, 1760–1764 (2007).
Catania, S. et al. Evolutionary persistence of DNA methylation for millions of years after ancient loss of a de novo methyltransferase. Cell 180, 263–277.e20 (2020).
Sharif, J. et al. The SRA protein Np95 mediates epigenetic inheritance by recruiting Dnmt1 to methylated DNA. Nature 450, 908–912 (2007).
Audergon, P. N. C. B. et al. Epigenetics. Restricted epigenetic inheritance of H3K9 methylation. Science 348, 132–135 (2015).
Coleman, R. T. & Struhl, G. Causal role for inheritance of H3K27me3 in maintaining the OFF state of a Drosophila HOX gene. Science 356, eaai8236 (2017).
Kowalik, K. M. et al. The Paf1 complex represses small-RNA-mediated epigenetic gene silencing. Nature 520, 248–252 (2015).
Laprell, F., Finkl, K. & Müller, J. Propagation of Polycomb-repressed chromatin requires sequence-specific recruitment to DNA. Science 356, 85–88 (2017).
Ragunathan, K., Jih, G. & Moazed, D. Epigenetics. Epigenetic inheritance uncoupled from sequence-specific recruitment. Science 348, 1258699 (2015).
Wang, X. & Moazed, D. DNA sequence-dependent epigenetic inheritance of gene silencing and histone H3K9 methylation. Science 356, 88–91 (2017).
Yu, R., Wang, X. & Moazed, D. Epigenetic inheritance mediated by coupling of RNAi and histone H3K9 methylation. Nature 558, 615–619 (2018).
Kuroda, M. I., Kang, H., De, S. & Kassis, J. A. Dynamic competition of Polycomb and Trithorax in transcriptional programming. Annu. Rev. Biochem. 89, 235–253 (2020).
Steffen, P. A. & Ringrose, L. What are memories made of? How Polycomb and Trithorax proteins mediate epigenetic memory. Nat. Rev. Mol. Cell Biol. 15, 340–356 (2014).
Berry, S., Hartley, M., Olsson, T. S. G., Dean, C. & Howard, M. Local chromatin environment of a Polycomb target gene instructs its own epigenetic inheritance. eLife 4, 105 (2015).
Beuchle, D., Struhl, G. & Müller, J. Polycomb group proteins and heritable silencing of Drosophila Hox genes. Development 128, 993–1004 (2001).
Jones, R. S. & Gelbart, W. M. Genetic analysis of the enhancer of zeste locus and its role in gene regulation in Drosophila melanogaster. Genetics 126, 185–199 (1990).
Simon, J., Chiang, A. & Bender, W. Ten different Polycomb group genes are required for spatial control of the abdA and AbdB homeotic products. Development 114, 493–505 (1992).
Struhl, G. & Akam, M. Altered distributions of Ultrabithorax transcripts in extra sex combs mutant embryos of Drosophila. EMBO J. 4, 3259–3264 (1985).
Reverón-Gómez, N. et al. Accurate recycling of parental histones reproduces the histone modification landscape during DNA replication. Mol. Cell 72, 239–249.e5 (2018).
Margueron, R. et al. Role of the polycomb protein EED in the propagation of repressive histone marks. Nature 461, 762–767 (2009).
Xu, C. et al. Binding of different histone marks differentially regulates the activity and specificity of polycomb repressive complex 2 (PRC2). Proc. Natl Acad. Sci. USA 107, 19266–19271 (2010).
Gaydos, L. J., Wang, W. & Strome, S. Gene repression. H3K27me and PRC2 transmit a memory of repression across generations and during development. Science 345, 1515–1518 (2014).
Højfeldt, J. W. et al. Accurate H3K27 methylation can be established de novo by SUZ12-directed PRC2. Nat. Struct. Mol. Biol. 20, 1123–1232 (2018).
Lavarone, E., Barbieri, C. M. & Pasini, D. Dissecting the role of H3K27 acetylation and methylation in PRC2 mediated control of cellular identity. Nat. Commun. 10, 1679 (2019).
Oksuz, O. et al. Capturing the onset of PRC2-mediated repressive domain formation. Mol. Cell 70, 1149–1162.e5 (2018).
Sarma, K., Margueron, R., Ivanov, A., Pirrotta, V. & Reinberg, D. Ezh2 requires PHF1 to efficiently catalyze H3 lysine 27 trimethylation in vivo. Mol. Cell Biol. 28, 2718–2731 (2008).
He, Y. et al. The EED protein–protein interaction inhibitor A-395 inactivates the PRC2 complex. Nat. Chem. Biol. 13, 389–395 (2017).
Konze, K. D. et al. An orally bioavailable chemical probe of the lysine methyltransferases EZH2 and EZH1. ACS Chem. Biol. 8, 1324–1334 (2013).
Ezhkova, E. et al. Ezh2 orchestrates gene expression for the stepwise differentiation of tissue-specific stem cells. Cell 136, 1122–1135 (2009).
Ezhkova, E. et al. EZH1 and EZH2 cogovern histone H3K27 trimethylation and are essential for hair follicle homeostasis and wound repair. Genes Dev. 25, 485–498 (2011).
Wassef, M. et al. Impaired PRC2 activity promotes transcriptional instability and favors breast tumorigenesis. Genes Dev. 29, 2547–2562 (2015).
Woodhouse, S., Pugazhendhi, D., Brien, P. & Pell, J. M. Ezh2 maintains a key phase of muscle satellite cell expansion but does not regulate terminal differentiation. J. Cell Sci. 126, 565–579 (2013).
Xu, J. et al. Landscape of monoallelic DNA accessibility in mouse embryonic stem cells and neural progenitor cells. Nat. Genet. 49, 377–386 (2017).
Chang, H. Y. et al. Diversity, topographic differentiation, and positional memory in human fibroblasts. Proc. Natl Acad. Sci. USA 99, 12877–12882 (2002).
Baubec, T. et al. Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation. Nature 520, 243–247 (2015).
Morselli, M. et al. In vivo targeting of de novo DNA methylation by histone modifications in yeast and mouse. eLife 4, e06205 (2015).
Hosogane, M., Funayama, R., Shirota, M. & Nakayama, K. Lack of transcription triggers H3K27me3 accumulation in the gene body. Cell Rep. 16, 696–706 (2016).
Riising, E. M. et al. Gene silencing triggers polycomb repressive complex 2 recruitment to CpG islands genome wide. Mol. Cell 55, 347–360 (2014).
Chen, S. et al. The histone H3 Lys 27 demethylase JMJD3 regulates gene expression by impacting transcriptional elongation. Genes Dev. 26, 1364–1375 (2012).
Lee, M. G. et al. Demethylation of H3K27 regulates polycomb recruitment and H2A ubiquitination. Science 318, 447–450 (2007).
Pasini, D. et al. Characterization of an antagonistic switch between histone H3 lysine 27 methylation and acetylation in the transcriptional regulation of Polycomb group target genes. Nucleic Acids Res. 38, 4958–4969 (2010).
Tie, F. et al. CBP-mediated acetylation of histone H3 lysine 27 antagonizes Drosophila Polycomb silencing. Development 136, 3131–3141 (2009).
Deaton, A. M. et al. Enhancer regions show high histone H3.3 turnover that changes during differentiation. eLife 5, e1002358 (2016).
Kraushaar, D. C. et al. Genome-wide incorporation dynamics reveal distinct categories of turnover for the histone variant H3.3. Genome Biol. 14, R121 (2013).
Beltran, M. et al. G-tract RNA removes Polycomb repressive complex 2 from genes. Nat. Struct. Mol. Biol. 26, 899–909 (2019).
Beltran, M. et al. The interaction of PRC2 with RNA or chromatin is mutually antagonistic. Genome Res. 26, 896–907 (2016).
Schmitges, F. W. et al. Histone methylation by PRC2 is inhibited by active chromatin marks. Mol. Cell 42, 330–341 (2011).
Yuan, W. et al. H3K36 methylation antagonizes PRC2-mediated H3K27 methylation. J. Biol. Chem. 286, 7983–7989 (2011).
Berry, S., Dean, C. & Howard, M. Slow chromatin dynamics allow polycomb target genes to filter fluctuations in transcription factor activity. Cell Syst. 4, 445–457.e8 (2017).
Lövkvist, C. & Howard, M. Using computational modelling to reveal mechanisms of epigenetic Polycomb control. Biochem Soc. Trans. 49, 71–77 (2021).
Jermann, P., Hoerner, L., Burger, L. & Schübeler, D. Short sequences can efficiently recruit histone H3 lysine 27 trimethylation in the absence of enhancer activity and DNA methylation. Proc. Natl Acad. Sci. USA 111, E3415–E3421 (2014).
Li, H. et al. Polycomb-like proteins link the PRC2 complex to CpG islands. Nature 549, 287–291 (2017).
Mendenhall, E. M. et al. GC-rich sequence elements recruit PRC2 in mammalian ES cells. PLoS Genet. 6, e1001244 (2010).
Chavez, A. et al. Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328 (2015).
Banaszynski, L. A., Chen, L.-C., Maynard-Smith, L. A., Ooi, A. G. L. & Wandless, T. J. A rapid, reversible, and tunable method to regulate protein function in living cells using synthetic small molecules. Cell 126, 995–1004 (2006).
Senturk, S. et al. Rapid and tunable method to temporally control gene editing based on conditional Cas9 stabilization. Nat. Commun. 8, 14370 (2017).
Deng, C. et al. HoxBlinc RNA recruits Set1/MLL complexes to activate hox gene expression patterns and mesoderm lineage development. Cell Rep. 14, 103–114 (2016).
Hu, M. et al. Histone H3 lysine 36 methyltransferase Hypb/Setd2 is required for embryonic vascular remodeling. Proc. Natl Acad. Sci. USA 107, 2956–2961 (2010).
Xu, Q. et al. SETD2 regulates the maternal epigenome, genomic imprinting and embryonic development. Nat. Genet. 51, 844–856 (2019).
Agger, K. et al. UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature 449, 731–734 (2007).
De Santa, F. et al. The histone H3 lysine-27 demethylase Jmjd3 links inflammation to inhibition of polycomb-mediated gene silencing. Cell 130, 1083–1094 (2007).
Lan, F. et al. A histone H3 lysine 27 demethylase regulates animal posterior development. Nature 449, 689–694 (2007).
Ptashne, M. On the use of the word ‘epigenetic’. Curr. Biol. 17, R233–R236 (2007).
Oktaba, K. et al. Dynamic regulation by polycomb group protein complexes controls pattern formation and the cell cycle in Drosophila. Dev. Cell 15, 877–889 (2008).
Erokhin, M. et al. Transcriptional read-through is not sufficient to induce an epigenetic switch in the silencing activity of Polycomb response elements. Proc. Natl Acad. Sci. USA 112, 14930–14935 (2015).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Wassef, M. et al. Versatile and precise gene-targeting strategies for functional studies in mammalian cell lines. Methods 121-122, 45–54 (2017).
Yusa, K., Zhou, L., Li, M. A., Bradley, A. & Craig, N. L. A hyperactive piggyBac transposase for mammalian applications. Proc. Natl Acad. Sci. USA 108, 1531–1536 (2011).
Kearns, N. A. et al. Cas9 effector-mediated regulation of transcription and differentiation in human pluripotent stem cells. Development 141, 219–223 (2014).
Vidigal, J. A. & Ventura, A. Rapid and efficient one-step generation of paired gRNA CRISPR–Cas9 libraries. Nat. Commun. 6, 8083–8087 (2015).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Chatterjee, P. et al. A Cas9 with PAM recognition for adenine dinucleotides. Nat. Commun. 11, 2474 (2020).
Koblan, L. W. et al. Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction. Nat. Biotechnol. 36, 843–846 (2018).
Suzuki, K. et al. In vivo genome editing via CRISPR/Cas9 mediated homology-independent targeted integration. Nature 540, 144–149 (2016).
Conti, L. et al. Niche-independent symmetrical self-renewal of a mammalian tissue stem cell. PLoS Biol. 3, e283 (2005).
Gendrel, A.-V. et al. Developmental dynamics and disease potential of random monoallelic gene expression. Dev. Cell 28, 366–380 (2014).
Wassef, M., Michaud, A. & Margueron, R. Association between EZH2 expression, silencing of tumor suppressors and disease outcome in solid tumors. Cell Cycle 15, 2256–2262 (2016).
Agger, K. et al. The H3K27me3 demethylase JMJD3 contributes to the activation of the INK4A-ARF locus in response to oncogene- and stress-induced senescence. Genes Dev. 23, 1171–1176 (2009).
Molden, R. C. & Garcia, B. A. Middle-down and top-down mass spectrometric analysis of co-occurring histone modifications. Curr. Protoc. Protein Sci. 77, 23.7.1–28 (2014).
Poullet, P., Carpentier, S. & Barillot, E. myProMS, a web server for management and validation of mass spectrometry-based proteomic data. Proteomics 7, 2553–2556 (2007).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinforma. 10, 48 (2009).
Supek, F., Bošnjak, M., Škunca, N. & Šmuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 6, e21800 (2011).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Castro-Mondragon, J. A., Jaeger, S., Thieffry, D., Thomas-Chollier, M. & van Helden, J. RSAT matrix-clustering: dynamic exploration and redundancy reduction of transcription factor binding motif collections. Nucleic Acids Res. 45, e119 (2017).
Gillespie, D. T. Exact stochastic simulation of coupled chemical reactions. J. Phys. Chem. 81, 2340–2361 (1977).
Karimi, M. et al. LUMA (LUminometric Methylation Assay)—a high throughput method to the analysis of genomic DNA methylation. Exp. Cell Res. 312, 1989–1995 (2006).
Acknowledgements
We thank J.-P. Concordet, O. Cuvier, D. Delpierre, M. Greenberg, D. Moazed, D. Reinberg, R. Schneider and M.-E. Torres-Padilla for valuable comments on the manuscript, M. Schulz for help with the LUMA assay, and members of the Margueron laboratory for discussions, help and advice. Work in the Margueron laboratory was supported by the FRM (Fondation pour la Recherche Médicale), the ARC (Fondation pour la Recherche sur le Cancer), the ANR (AMetHist) and the Labex DEEP. D.H. was supported by a postdoctoral fellowship from the FRM (no. SPF20150934266). The Cell & Tissue Imaging platform of Institut Curie provided training and access to microscopes. High-throughput sequencing was performed by the NGS platform of Institut Curie, with S. Lameiras, P. Legoix and V. Reynal providing valuable advice on experimental design. The platform is supported by grants (nos. ANR-10-EQPX-03 and ANR-10-INBS-09-08) from Agence Nationale de la Recherche (investissements d’avenir) and by Cancéropôle Île-de-France.
Author information
Authors and Affiliations
Contributions
M.W. made the initial observation that certain target genes remained stably de-repressed after transient disruption of PRC2. D.H., M.W. and R.M. conceived the study and designed the experiments with input from all the authors. D.H. and M.W. performed most of the experiments. C.L. and M.H. designed, and C.L. performed, all the mathematical modeling. D.Z. performed the bioinformatic analysis. S.A. performed experiments. B.L. carried out the MS experimental work and D.L. supervised MS and data analysis. T.H. assisted in the generation of cell lines. R.M. supervised the study. D.H., M.W. and R.M. prepared the manuscript with input from all the authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work. Peer review reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1
Transient disruption of PRC2 triggers permanent activation of a large subset of repressed target genes in differentiated somatic cells. a, Proportions of non-responsive and responsive genes among the indicated totals of H3K27me3-positive genes, in NPCs, ESCs and iMEF line B after treatment with PRC2 inhibitors and washout, and in iMEF lines A and B after genetic deletion of Ezh2 followed by rescue. H3K27me3 profiles in ESCs were determined from a published data set26. NPC, mouse neural progenitor cells. ESC, mouse embryonic stem cells. iMEF, immortalized mouse embryonic fibroblasts. b, Metaplots of average ATAC-seq density over the interval TSS -/ + 3 kb for the full sets of H3K27me3-positive and transcriptionally responsive PRC2 targets in NPC (ref. 36) and iMEF A (this study). NPC H3K27me3-positive and responsive: n = 5169 and n = 412 genes respectively; iMEF A H3K27me3-positive and responsive: n = 8344 and n = 326 genes respectively. p-values refer to two-sided Mann–Whitney tests comparing read counts for H3K27me3-positive and responsive genes over the interval. c, Western blot of nuclear extracts from the indicated conditions (n = 2 biologically independent samples). PRC2i, PRC2 inhibition with 2 µM UNC1999 and 4 µM A-395. d, MA plots comparing RNA-seq levels of H3K27me3-positive genes in ESCs cultured with PRC2 inhibitors, or after washout of the inhibitors, to mock-treated cells. CPM, counts per million. FC, fold change. Significantly differentially expressed genes (Benjamini-Hochberg-adjusted p < 0.05 for two-sided likelihood ratio test on negative binomial generalized linear model) are indicated in red. e, Western blot of nuclear extracts from the indicated conditions. Representative example of n = 2 independent samples. f, Heatmap of RNA-seq levels (log2 fold change versus the mean of the two mock replicates) in the indicated conditions for the full sets of reversible and irreversible PRC2 targets in iMEF B. g, Left, heatmap of RNA-seq levels (log2 fold change versus the mean of the two wild-type replicates) in the indicated conditions for the full sets of reversible and irreversible PRC2 targets in iMEF A. Right, metaplots of average H3K27me3 ChIP-seq density over the interval TSS – 3 kb to TES + 3 kb for the full sets of reversible and irreversible PRC2 targets in iMEF A in the indicated conditions. TSS, transcription start site. TES, transcription end site. h, As in g but in iMEF B.
Extended Data Fig. 2
Reversible and irreversible PRC2 targets are similarly enriched in gene ontology terms related to development. a, Left, top 10 most highly enriched gene ontology (GO) terms among genes whose transcription start site overlaps an H3K27me3 peak in NPCs, using all mouse genes as background. Right, top 10 most highly enriched GO terms among reversible and irreversible genes in NPCs, respectively, as indicated, using all mouse genes as background. p-values were calculated using a one-sided hypergeometric test and adjusted for multiple comparisons using the false discovery rate (FDR) method. b, As in a but for iMEF B subjected to transient PRC2i treatment (middle panels) or Ezh2 deletion and rescue (right panel, no significantly enriched GO term was found for reversible genes). PRC2i, PRC2 inhibition with 2 µM UNC1999 and 4 µM A-395. c, As in a, but for iMEF A.
Extended Data Fig. 3
Disruption of PRC2 activity in iMEFs triggers widespread genomic binding of de-repressed homeotic transcription factors. a, Tracks of ATAC-seq performed in the indicated genotypes (chromosome 5: 98,320,001-98,400,000). Red box highlights a peak irreversibly gained following temporary deletion of Ezh2. ATAC, Assay for Transposase-Accessible Chromatin. b, Results of mammalian Hox factors among total known motif search using HOMER software, on the indicated peaks called using SEACR, and with all Ezh2 KO peaks as background. p-values were calculated using a one-sided cumulative binomial distribution test. q-values represent p-values corrected for multiple testing using the Benjamini-Hochberg procedure (a blue bar denotes a q-value > 0.05; a red bar and * denote a q-value < 0.05; and a red bar and ** denote a q-value < 0.005). See also Supplementary Table 2. NS, not significant. c, Normalized RNA-seq counts in iMEF A were averaged among members of each Hox paralogous group 1, 2, 4, 9 and 13, and the fold derepression in Ezh2 rescue versus wild-type was plotted against the expression level in Ezh2 rescue. Hox group 9 and 13 are colored in red to highlight the combination of their high fold upregulation and high expression; other Hox groups are colored in blue. TMM-RPKM, trimmed mean of M-values-reads per kilobase per million. FC, fold change.
Extended Data Fig. 4
Stable epigenetic switching of PRC2 targets is not a gene-intrinsic property. a, Left, schematic illustrating the heterogeneous identity of distinct MEF clones. Right, tracks of H3K27me3 ChIP-seq and RNA-seq for the indicated conditions over individual genes (A830082K12Rik and Nr2f1, chromosome 13: 78,170,001-78,264,000; Smarca2, chromosome 19: 26,589,001-26,795,000; Pax9, chromosome 12: 56,690,001-56,714,000) comparing iMEF A and iMEF B. b, Tracks of H3K27me3 ChIP-seq and RNA-seq for the indicated conditions over individual genes or regions (Hoxb cluster, chromosome 11: 96,249,001-96,395,000; Axin2, chromosome 11: 108,913,001-108,955,000; Lin28b, chromosome 10: 45,369,001-45,495,000) comparing iMEF A and iMEF B.
Extended Data Fig. 5
Stable epigenetic switching of target genes is correlated to level of transcriptional activity upon disruption of PRC2. a, Box plots (median, lower and upper quartiles, lowest and highest values) of average nascent RNA-seq read densities, ChIP-seq read densities for the indicated features, ATAC-seq densities and fraction of CpG methylation. n = 124 reversible and n = 154 irreversible genes. For nascent RNA-seq, ChIP-seq and ATAC-seq, p-values refer to two-sided Mann-Whitney tests comparing irreversible and reversible genes. Difference in nascent RNA (Hodges-Lehmann): 0.15; 95% confidence interval: 0.05 to 0.27. Difference in H3K36me3 (Hodges-Lehmann): 0.56; 95% confidence interval: 0.39-0.76. For DNA methylation, p-values refer to a Kruskal-Wallis test corrected for multiple comparisons (Dunn’s test). Mean rank difference for reversible v. irreversible in Ezh2 KO: −88.90. TSS = transcription start site. NS, not significant. TMM-RPKM, trimmed mean of M-values-reads per kilobase per million. b, Genomic tracks for the indicated features at representative reversible (Casz1, chromosome 4: 148,719,001-149,041,000) and irreversible (Cpa6, chromosome 1: 10,086,001-10,929,000) PRC2 targets in iMEF A Ezh2 KO cells. c, Scatter plot of all reversible and irreversible genes according to fraction of methylated CpG and log2 TMM-RPKM H3K36me3 ChIP-seq read density. Spearman’s rank correlation coefficient and corresponding two-sided t-test p-value are indicated. 95% confidence interval: 0.24 to 0.46. d, Graph depicting levels of transcription (fraction of level reached in case with 20 cell cycles of transient PRC2 disruption), and H3K27me3 levels, predicted at the conclusion of a simulation similar to that conducted for Fig. 3d (top) for the indicated α, β values. Transient disruption of PRC2 was simulated for a fixed duration (corresponding to that of Fig. 3d) with varying cell cycle lengths, followed by 20 cell cycles of normal duration during which PRC2 activity was restored, and levels of transcription and H3K27me3 at 20th cell cycle were plotted. e, Left, box plots (median, lower and upper quartiles, lowest and highest values) comparing H3K27me3 levels of all simulated genes predicted to exhibit reversible or irreversible de-repression, respectively. The levels are from the wild-type simulations in Fig. 3b (left), with parameter values sampled logarithmically over parameter space (see Supplementary Methods). n = 903 reversible and n = 417 irreversible simulated genes. The p-value refers to a two-sided Mann–Whitney test comparing irreversible and reversible genes for each set of simulated values. Difference (Hodges-Lehmann): 0.10; 95% confidence interval: 0.01-0.12. Right, box plots (median, lower and upper quartiles, lowest and highest values) of observed H3K27me3 levels over all reversible and irreversible genes, respectively, in the indicated cell lines under wild-type untreated conditions. NPC PRC2i reversible and irreversible: n = 218 and n = 87 genes respectively; iMEFB PRC2i reversible and irreversible: n = 227 and n = 96 genes respectively; iMEFA Ezh2 KO reversible and irreversible: n = 124 and n = 154 genes respectively; iMEFB Ezh2 KO reversible and irreversible: n = 58 and n = 66 genes respectively. p-values refer to two-sided Mann–Whitney tests comparing irreversible and reversible genes for each cell line. Difference in H3K27me3 for iMEF A (Hodges-Lehmann): 0.45; 95% confidence interval: 0.14-0.58. PRC2i, PRC2 inhibition with 2 µM UNC1999 and 4 µM A-395.
Extended Data Fig. 6
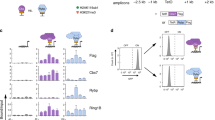
Transient activation of individual PRC2 target genes can trigger permanent switching of transcriptional states. a, H3K27me3 levels measured by CUT&RUN over indicated target genes in cells bearing the DD-dCas9-VPR construct and either lacking (wild-type, Ezh2 KO) or constitutively expressing (VPR OFF, washout) sgRNAs specific to the target gene. Values within each experiment are normalized to the wild-type condition. Horizontal lines represent mean values. b, Western blot analysis of nuclear extracts from the indicated conditions. Cells having undergone a transient induction experiment, and a subclone thereof in which DD-dCas9-VPR was genetically deleted, were subjected to a second Dox+Shield1 treatment to confirm the deletion and check the persistence of target gene product expression. Representative example of n = 2 independent samples. c, Messenger RNA levels of indicated target genes in cells constitutively expressing sgRNAs specific to the target gene and either bearing the DD-dCas9-VPR construct or deleted for DD-dCas9-VPR after the conclusion of a transient induction experiment. Values within each experiment are normalized to the VPR washout condition. n.d., not detected.
Extended Data Fig. 7 Hoxb cluster genes exhibit interdependent switching of transcriptional states in response to transient activation.
a, Messenger RNA levels of indicated target genes in cells bearing the DD-dCas9-VPR construct and either lacking (wild-type, Ezh2 KO) or constitutively expressing (VPR OFF, ON, washout) sgRNAs specific to the target gene. Values within each experiment are normalized to the Ezh2 KO condition. Horizontal lines represent mean values. b, Left, RNA-seq and nascent-RNA-seq over the Hoxb cluster (chromosome 11: 96,182,001-96,373,000) for the indicated conditions in iMEF B, with the position of the long-Hoxb transcript shown below. Right, as in a, but for long-Hoxb. c, Messenger RNA levels of indicated target genes in cells bearing the DD-dCas9-VPR construct and either lacking (wild-type, Ezh2 KO) or constitutively expressing (VPR OFF, ON, washout) sgRNAs specific to long-Hoxb.
Extended Data Fig. 8
Short pulses of transient activation can suffice to produce a switch in transcriptional states. a, Messenger RNA levels of long-Hoxb in cells bearing the DD-dCas9-VPR construct and either lacking (wild-type, Ezh2 KO) or constitutively expressing (VPR ON, washout) sgRNAs specific to long-Hoxb, after induction for the indicated duration followed or not by 9-day washout. Values are normalized to the Ezh2 KO condition. b, Messenger RNA levels of indicated target genes in cells bearing the DD-dCas9-VPR construct and either lacking (wild-type, Ezh2 KO) or constitutively expressing (VPR OFF, ON, washout) sgRNAs specific to the target gene. Values within each experiment are normalized to the Ezh2 KO condition. Horizontal lines represent mean values. n.d., not detected. c, H3K27me3 levels measured by CUT&RUN over Nr2f1 in cells bearing the DD-dCas9-VPR construct, constitutively expressing sgRNAs specific to Nr2f1, and subjected to the indicated conditions. VPR washout condition follows a 6-day induction. Values within each experiment are normalized to the VPR OFF condition. d, Graphs depicting the levels of transcription (fraction of the maximum level reached for the indicated α, β values), and H3K27me3 levels, predicted at the conclusion of a simulation similar to that conducted for Fig. 3d. A transient pulse of transcriptional activation with α = αmax = 100 was simulated for the indicated number of cell cycles, followed by 20 cell cycles at the indicated α, β values, and levels of transcription and H3K27me3 at 20th cell cycle were plotted.
Extended Data Fig. 9
Stable switching of transcriptional states can occur independently of Setd2, of Utx and Jmjd3, and of Crebbp, and global loss of DNA methylation does not restore PRC2-dependent silencing. a, b, c, Western blot analysis of nuclear extracts from iMEF B cells of the indicated genotypes. Representative examples of n = 2 independent samples per analysis. d, Global levels of CpG methylation measured by LUminometric Methylation Assay in iMEF B cells of the indicated genotypes after treatment for three days with different concentrations of 5-aza-2’-deoxycytidine as indicated. e, Messenger RNA levels of indicated target genes in cells of the indicated genotypes after treatment for three days with different concentrations of 5-aza-2’-deoxycytidine as indicated. The values of 2 biologically independent replicates are shown.
Extended Data Fig. 10
Switching of transcriptional states following transient activation can be gene-autonomous. Top, MA plots comparing RNA-seq levels in cells in which the indicated gene was subjected to transient activation as depicted in Fig. 4a (VPR washout) to levels in control untreated cells (VPR OFF). CPM, counts per million. FC, fold change. Significantly differentially expressed genes (Benjamini-Hochberg-adjusted p < 0.05) are indicated in red. A830082K12Rik is transcribed divergently from a common promoter region shared with Nr2f1. Bottom, RNA-seq tracks for the indicated conditions at transiently activated genes (Nr2f1, chromosome 13: 78,179,001-78,248,000; Foxa1, chromosome 12: 57,539,001-57,549,000).
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2, Supplementary Methods and Supplementary Source data.
Supplementary Tables
Supplementary Tables 1–5.
Source data
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 9
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Holoch, D., Wassef, M., Lövkvist, C. et al. A cis-acting mechanism mediates transcriptional memory at Polycomb target genes in mammals. Nat Genet 53, 1686–1697 (2021). https://doi.org/10.1038/s41588-021-00964-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-021-00964-2
This article is cited by
-
Novel epigenetic molecular therapies for imprinting disorders
Molecular Psychiatry (2023)
-
Impaired histone inheritance promotes tumor progression
Nature Communications (2023)
-
Remembering through the genome: the role of chromatin states in brain functions and diseases
Translational Psychiatry (2023)