Abstract
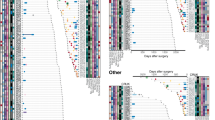
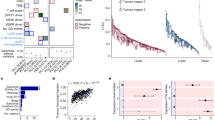
Lung cancer in never smokers (LCINS) is a common cause of cancer mortality but its genomic landscape is poorly characterized. Here high-coverage whole-genome sequencing of 232 LCINS showed 3 subtypes defined by copy number aberrations. The dominant subtype (piano), which is rare in lung cancer in smokers, features somatic UBA1 mutations, germline AR variants and stem cell-like properties, including low mutational burden, high intratumor heterogeneity, long telomeres, frequent KRAS mutations and slow growth, as suggested by the occurrence of cancer drivers’ progenitor cells many years before tumor diagnosis. The other subtypes are characterized by specific amplifications and EGFR mutations (mezzo-forte) and whole-genome doubling (forte). No strong tobacco smoking signatures were detected, even in cases with exposure to secondhand tobacco smoke. Genes within the receptor tyrosine kinase–Ras pathway had distinct impacts on survival; five genomic alterations independently doubled mortality. These findings create avenues for personalized treatment in LCINS.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The 232 normal and tumor-paired raw data (BAM files) of the WGS datasets have been deposited in the dbGaP under accession no. phs001697.v1.p1. Researchers will need to obtain authorization from the dbGaP to download these data. The RNA-seq raw data (FASTQ files) have been submitted to the Gene Expression Omnibus under accession no. GSE171415. The germline variant dataset from the EAGLE whole-exome sequencing study can be access at the dbGaP with accession no. phs002496.v1.p1. In addition, histological images of these tumors can be found at https://episphere.github.io/svs. Public datasets were used in this study including gnomAD v.2.1.1/ExAC v.0.3.1 (https://gnomad.broadinstitute.org/), 1000 Genomes (phase 3 v.5, https://www.internationalgenome.org/) and dbSNP (v.138, https://www.ncbi.nlm.nih.gov/snp/).
Code availability
The code for the WGS subclonal copy number caller can be found at https://github.com/Wedge-lab/battenberg (v.2.2.8). The code for somatic mutation filtering can be found at https://github.com/xtmgah/Sherlock-Lung. The code for the Dirichlet process-based methods for subclonal reconstruction of tumors can be found at https://github.com/Wedge-lab/dpclust (v.2.2.8). The code for the mutational signature analysis can be found at https://pypi.org/project/sigproextractor/ (SigProfilerExtractor v.0.0.5.77). The code for inferring the order of genomic events can be found at https://github.com/hturner/PlackettLuce (v.0.2-2). The code for the chronological timing analysis can be found at https://gerstung-lab.github.io/PCAWG-11/. The code for P-MACD can be found at https://github.com/NIEHS/P-MACD.
References
The Cancer Atlas: Lung Cancer (American Cancer Society, 2021); https://canceratlas.cancer.org/the-burden/lung-cancer/
Cho, J. et al. Proportion and clinical features of never-smokers with non-small cell lung cancer. Chin. J. Cancer 36, 20 (2017).
Campbell, J. D. et al. Distinct patterns of somatic genome alterations in lung adenocarcinomas and squamous cell carcinomas. Nat. Genet. 48, 607–616 (2016).
Collisson, E. A. et al. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Chen, J. et al. Genomic landscape of lung adenocarcinoma in East Asians. Nat. Genet. 52, 177–186 (2020).
Govindan, R. et al. Genomic landscape of non-small cell lung cancer in smokers and never-smokers. Cell 150, 1121–1134 (2012).
Imielinski, M. et al. Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell 150, 1107–1120 (2012).
Lee, J. J.-K. et al. Tracing oncogene rearrangements in the mutational history of lung adenocarcinoma. Cell 177, 1842–1857.e21 (2019).
Shi, J. et al. Somatic genomics and clinical features of lung adenocarcinoma: a retrospective study. PLoS Med. 13, e1002162 (2016).
Wang, C. et al. Whole-genome sequencing reveals genomic signatures associated with the inflammatory microenvironments in Chinese NSCLC patients. Nat. Commun. 9, 2054 (2018).
Fernandez-Cuesta, L. et al. Frequent mutations in chromatin-remodelling genes in pulmonary carcinoids. Nat. Commun. 5, 3518 (2014).
Campbell, P. J. et al. Pan-cancer analysis of whole genomes. Nature 578, 82–93 (2020).
Wu, K. et al. Frequent alterations in cytoskeleton remodelling genes in primary and metastatic lung adenocarcinomas. Nat. Commun. 6, 10131 (2015).
Carrot-Zhang, J. et al. Whole-genome characterization of lung adenocarcinomas lacking the RTK/RAS/RAF pathway. Cell Rep. 34, 108707 (2021).
Landi, M. T. et al. Tracing lung cancer risk factors through mutational signatures in never smokers: the Sherlock-Lung study. Am. J. Epidemiol. 190, 962–976 (2021).
Skoulidis, F. et al. Co-occurring genomic alterations define major subsets of KRAS-mutant lung adenocarcinoma with distinct biology, immune profiles, and therapeutic vulnerabilities. Cancer Discov. 5, 860–877 (2015).
Moll, U. M. & Petrenko, O. The MDM2-p53 interaction. Mol. Cancer Res. 1, 1001–1008 (2003).
Wala, J. A. et al. Selective and mechanistic sources of recurrent rearrangements across the cancer genome. Preprint at bioRxiv https://doi.org/10.1101/187609 (2017).
Reznik, E. et al. Mitochondrial DNA copy number variation across human cancers. eLife 5, e10769 (2016).
McGranahan, N. et al. Allele-specific HLA loss and immune escape in lung cancer evolution. Cell 171, 1259–1271.e11 (2017).
Bailey, M. H. et al. Comprehensive characterization of cancer driver genes and mutations. Cell 173, 371–385.e18 (2018).
Moudry, P. et al. Ubiquitin-activating enzyme UBA1 is required for cellular response to DNA damage. Cell Cycle 11, 1573–1582 (2012).
Martínez-Jiménez, F. A compendium of mutational cancer driver genes. Nat. Rev. Cancer 20, 555–572 (2020).
Huang, K.-L. et al. Pathogenic germline variants in 10,389 adult cancers. Cell 173, 355–370.e14 (2018).
Staaf, J. et al. Whole-genome sequencing of triple-negative breast cancers in a population-based clinical study. Nat. Med. 25, 1526–1533 (2019).
Alexandrov, L. B. et al. The repertoire of mutational signatures in human cancer. Nature 578, 94–101 (2020).
Bergstrom, E. N. et al. SigProfilerMatrixGenerator: a tool for visualizing and exploring patterns of small mutational events. BMC Genomics 20, 685 (2019).
Petljak, M. et al. Characterizing mutational signatures in human cancer cell lines reveals episodic APOBEC mutagenesis. Cell 176, 1282–1294.e20 (2019).
Jager, M. et al. Deficiency of nucleotide excision repair is associated with mutational signature observed in cancer. Genome Res. 29, 1067–1077 (2019).
Singh, V. K., Rastogi, A., Hu, X., Wang, Y. & De, S. Mutational signature SBS8 predominantly arises due to late replication errors in cancer. Commun. Biol. 3, 421 (2020).
Roberts, S. A. et al. An APOBEC cytidine deaminase mutagenesis pattern is widespread in human cancers. Nat. Genet. 45, 970–976 (2013).
Kucab, J. E. et al. A compendium of mutational signatures of environmental agents. Cell 177, 821–836.e16 (2019).
Tokiwa, H. & Sera, N. Contribution of nitrated polycyclic aromatic hydrocarbons in diesel particles to human lung cancer induction. Polycycl. Aromat. Compd. 21, 231–245 (2000).
Saini, N. et al. Mutation signatures specific to DNA alkylating agents in yeast and cancers. Nucleic Acids Res. 48, 3692–3707 (2020).
Chan, K. et al. An APOBEC3A hypermutation signature is distinguishable from the signature of background mutagenesis by APOBEC3B in human cancers. Nat. Genet. 47, 1067–1072 (2015).
Barthel, F. P. et al. Systematic analysis of telomere length and somatic alterations in 31 cancer types. Nat. Genet. 49, 349–357 (2017).
Feuerbach, L. et al. TelomereHunter—in silico estimation of telomere content and composition from cancer genomes. BMC Bioinformatics 20, 272 (2019).
Davies, H. et al. HRDetect is a predictor of BRCA1 and BRCA2 deficiency based on mutational signatures. Nat. Med. 23, 517–525 (2017).
Zhao, E. Y. et al. Homologous recombination deficiency and platinum-based therapy outcomes in advanced breast cancer. Clin. Cancer Res. 23, 7521–7530 (2017).
Letouzé, E. et al. Mutational signatures reveal the dynamic interplay of risk factors and cellular processes during liver tumorigenesis. Nat. Commun. 8, 1315 (2017).
Shinde, J. et al. Palimpsest: an R package for studying mutational and structural variant signatures along clonal evolution in cancer. Bioinformatics 34, 3380–3381 (2018).
Gerstung, M. et al. The evolutionary history of 2,658 cancers. Nature 578, 122–128 (2020).
Halvorsen, A. R. et al. TP53 mutation spectrum in smokers and never smoking lung cancer patients. Front. Genet. 7, 85 (2016).
Gu, J. et al. TP53 mutation is associated with a poor clinical outcome for non-small cell lung cancer: evidence from a meta-analysis. Mol. Clin. Oncol. 5, 705–713 (2016).
López, S. et al. Interplay between whole-genome doubling and the accumulation of deleterious alterations in cancer evolution. Nat. Genet. 52, 283–293 (2020).
Bielski, C. M. et al. Genome doubling shapes the evolution and prognosis of advanced cancers. Nat. Genet. 50, 1189–1195 (2018).
Jamal-Hanjani, M. et al. Tracking the evolution of non-small-cell lung cancer. N. Engl. J. Med. 376, 2109–2121 (2017).
IARC Working Group on the Evaluation of Carcinogenic Risks to Humans, International Agency for Research on Cancer. A Review of Human Carcinogens: Personal Habits and Indoor Combustions (International Agency for Research on Cancer, 2012).
United States Public Health Service. Office of the Surgeon General. The Health Consequences of Involuntary Exposure to Tobacco Smoke: A Report of the Surgeon General (US Department of Health and Human Services, Public Health Service, Office of the Surgeon General, 2006).
Lopez-Bigas, N. & Gonzalez-Perez, A. Are carcinogens direct mutagens? Nat. Genet. 52, 1137–1138 (2020).
Cho, I. J. et al. Mechanisms, hallmarks, and implications of stem cell quiescence. Stem Cell Reports 12, 1190–1200 (2019).
Fukada, S.-I., Ma, Y. & Uezumi, A. Adult stem cell and mesenchymal progenitor theories of aging. Front. Cell Dev. Biol. 2, 10 (2014).
Li, L. & Clevers, H. Coexistence of quiescent and active adult stem cells in mammals. Science 327, 542–545 (2010).
Kim, C. F. B. et al. Identification of bronchioalveolar stem cells in normal lung and lung cancer. Cell 121, 823–835 (2005).
Van Meter, M. E. M. et al. K-RasG12D expression induces hyperproliferation and aberrant signaling in primary hematopoietic stem/progenitor cells. Blood 109, 3945–3952 (2007).
Kubara, K. et al. Status of KRAS in iPSCs impacts upon self-renewal and differentiation propensity. Stem Cell Reports 11, 380–394 (2018).
Bax, M. et al. The ubiquitin proteasome system is a key regulator of pluripotent stem cell survival and motor neuron differentiation. Cells 8, 581 (2019).
Leon, T. Y. Y. et al. Transcriptional regulation of RET by Nkx2-1, Phox2b, Sox10, and Pax3. J. Pediatr. Surg. 44, 1904–1912 (2009).
Grey, W. et al. Activation of the receptor tyrosine kinase, RET, improves long-term hematopoietic stem cell outgrowth and potency. Blood 136, 2535–2547 (2020).
Fonseca-Pereira, D. et al. The neurotrophic factor receptor RET drives haematopoietic stem cell survival and function. Nature 514, 98–101 (2014).
Zhao, B. et al. ARID1A promotes genomic stability through protecting telomere cohesion. Nat. Commun. 10, 4067 (2019).
Sun, X. et al. Suppression of the SWI/SNF component Arid1a promotes mammalian regeneration. Cell Stem Cell 18, 456–466 (2016).
van der Vaart, A. & van den Heuvel, S. Switching on regeneration. Stem Cell Investig. 3, 41 (2016).
Wu, S., Zhang, R. & Bitler, B. G. Arid1a controls tissue regeneration. Stem Cell Investig. 3, 35 (2016).
Nagl, N. G. Jr, Wang, X., Patsialou, A., Van Scoy, M. & Moran, E. Distinct mammalian SWI/SNF chromatin remodeling complexes with opposing roles in cell-cycle control. EMBO J. 26, 752–763 (2007).
Chiba, S. Notch signaling in stem cell systems. Stem Cells 24, 2437–2447 (2006).
Yoshida, K. et al. Tobacco smoking and somatic mutations in human bronchial epithelium. Nature 578, 266–272 (2020).
Maeda, Y., Davé, V. & Whitsett, J. A. Transcriptional control of lung morphogenesis. Physiol. Rev. 87, 219–244 (2007).
Alanis, D. M., Chang, D. R., Akiyama, H., Krasnow, M. A. & Chen, J. Two nested developmental waves demarcate a compartment boundary in the mouse lung. Nat. Commun. 5, 3923 (2014).
Singh, I. et al. Hmga2 is required for canonical WNT signaling during lung development. BMC Biol. 12, 21 (2014).
Laughney, A. M. et al. Regenerative lineages and immune-mediated pruning in lung cancer metastasis. Nat. Med. 26, 259–269 (2020).
Duffy, M. J. et al. p53 as a target for the treatment of cancer. Cancer Treat. Rev. 40, 1153–1160 (2014).
Shaikh, M. F. et al. Emerging role of MDM2 as target for anti-cancer therapy: a review. Ann. Clin. Lab. Sci. 46, 627–634 (2016).
Chuang, J. C. et al. ERBB2-mutated metastatic non-small cell lung cancer: response and resistance to targeted therapies. J. Thorac. Oncol. 12, 833–842 (2017).
Harvey, R. D., Adams, V. R., Beardslee, T. & Medina, P. Afatinib for the treatment of EGFR mutation-positive NSCLC: a review of clinical findings. J. Oncol. Pharm. Pract. 26, 1461–1474 (2020).
Park, K. et al. Afatinib versus gefitinib as first-line treatment of patients with EGFR mutation-positive non-small-cell lung cancer (LUX-Lung 7): a phase 2B, open-label, randomised controlled trial. Lancet Oncol. 17, 577–589 (2016).
Shen, X. et al. A systematic analysis of the resistance and sensitivity of HER2YVMA receptor tyrosine kinase mutant to tyrosine kinase inhibitors in HER2-positive lung cancer. J. Recept. Signal Transduct. Res. 36, 89–97 (2016).
Miyazaki, M. et al. The p53 activator overcomes resistance to ALK inhibitors by regulating p53-target selectivity in ALK-driven neuroblastomas. Cell Death Discov. 4, 56 (2018).
Dey, P. et al. Genomic deletion of malic enzyme 2 confers collateral lethality in pancreatic cancer. Nature 542, 119–123 (2017).
Muller, F. L., Aquilanti, E. A. & DePinho, R. A. Collateral lethality: a new therapeutic strategy in oncology. Trends Cancer 1, 161–173 (2015).
Hsiehchen, D. et al. DNA repair gene mutations as predictors of immune checkpoint inhibitor response beyond tumor mutation burden. Cell Rep. Med. 1, 100034 (2020).
Rizvi, N. A. et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Hellmann, M. D. et al. Nivolumab plus ipilimumab in lung cancer with a high tumor mutational burden. N. Engl. J. Med. 378, 2093–2104 (2018).
Ready, N. et al. First-line nivolumab plus ipilimumab in advanced non-small-cell lung cancer (CheckMate 568): outcomes by programmed death ligand 1 and tumor mutational burden as biomarkers. J. Clin. Oncol. 37, 992–1000 (2019).
Canon, J. et al. The clinical KRAS(G12C) inhibitor AMG 510 drives anti-tumour immunity. Nature 575, 217–223 (2019).
Yang, L. et al. Targeting cancer stem cell pathways for cancer therapy. Signal Transduct. Target. Ther. 5, 8 (2020).
Medema, J. P. & Vermeulen, L. Microenvironmental regulation of stem cells in intestinal homeostasis and cancer. Nature 474, 318–326 (2011).
Jørsboe, E., Hanghøj, K. & Albrechtsen, A. fastNGSadmix: admixture proportions and principal component analysis of a single NGS sample. Bioinformatics 33, 3148–3150 (2017).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Kim, S. et al. Strelka2: fast and accurate calling of germline and somatic variants. Nat. Methods 15, 591–594 (2018).
Freed, D., Pan, R. & Aldana, R. TNscope: accurate detection of somatic mutations with haplotype-based variant candidate detection and machine learning filtering. Preprint at bioRxiv https://doi.org/10.1101/250647 (2018).
Zhu, B. et al. The genomic and epigenomic evolutionary history of papillary renal cell carcinomas. Nat. Commun. 11, 3096 (2020).
Karczewski, K. J. et al. The mutational constraints spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Ramos, A. H. et al. Oncotator: cancer variant annotation tool. Hum. Mutat. 36, E2423–E2429 (2015).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Hasan, M. S., Wu, X., Watson, L. T. & Zhang, L. UPS-indel: a universal positioning system for indels. Sci. Rep. 7, 14106 (2017).
Dentro, S. C., Wedge, D. C. & Van Loo, P. Principles of reconstructing the subclonal architecture of cancers. Cold Spring Harb. Perspect. Med. 7, a026625 (2017).
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012).
Scott, A. D. et al. CharGer: clinical Characterization of Germline variants. Bioinformatics 35, 865–867 (2019).
Landrum, M. J. et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 44, D862–D868 (2016).
Martincorena, I. et al. Universal patterns of selection in cancer and somatic tissues. Cell 171, 1029–1041.e21 (2017).
Muiños, F., Martínez-Jiménez, F., Pich, O., Gonzalez-Perez, A. & Lopez-Bigas, N. In silico saturation mutagenesis of cancer genes. Nature 596, 428–432 (2021).
Mermel, C. H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41 (2011).
Dewhurst, S. M. et al. Tolerance of whole-genome doubling propagates chromosomal instability and accelerates cancer genome evolution. Cancer Discov. 4, 175–185 (2014).
Yang, L. et al. Diverse mechanisms of somatic structural variations in human cancer genomes. Cell 153, 919–929 (2013).
Li, Y. et al. Patterns of somatic structural variation in human cancer genomes. Nature 578, 112–121 (2020).
Ding, Z. et al. Estimating telomere length from whole genome sequence data. Nucleic Acids Res. 42, e75 (2014).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Shukla, S. A. et al. Comprehensive analysis of cancer-associated somatic mutations in class I HLA genes. Nat. Biotechnol. 33, 1152–1158 (2015).
Bolli, N. et al. Heterogeneity of genomic evolution and mutational profiles in multiple myeloma. Nat. Commun. 5, 2997 (2014).
Luce, R. D. Individual Choice Behavior: a Theoretical Analysis (Wiley, 1959).
Plackett, R. L. The analysis of permutations. Appl. Stat. 24, 193 (1975).
Acknowledgements
This work has been supported by the Intramural Research Program of the Division of Cancer Epidemiology and Genetics, National Cancer Institute, and the Intramural Research Program of the National Institute of Environmental Health Sciences (project nos. Z01 ES050159 to S.H.W. and Z1AES103266 to D.A.G.), National Institutes of Health (NIH). This project was funded in whole or in part with federal funds from the National Cancer Institute, NIH, under contract nos. 75N91019D00024 and HHSN261201800001I. The content of this publication does not necessarily reflect the views or policies of the U.S. Department of Health and Human Services, nor does mention of trade names, commercial products or organizations imply endorsement by the U.S. Government. The research was supported by the Wellcome Trust Core Award, grant no. 203141/Z/16/Z with funding from the National Institute for Health Research Oxford Biomedical Research Centre. L.B.A. is an Abeloff V scholar and he is personally supported by an Alfred P. Sloan Research Fellowship and a Packard Fellowship for Science and Engineering. Research at the L.B.A. laboratory was supported by a National Institute of Environmental Health Sciences grant no. R01ES032547. The views expressed are those of the authors and not necessarily those of the National Health Service, National Institute for Health Research or Department of Health. The collection of samples from the Institut Universitaire de Cardiologie et de Pneumologie de Québec (IUCPQ), Université Laval was supported by the IUCPQ Foundation. The GR Program 2010-2316264 supported L.A.M. for sample collection by the Istituto di Ricovero e Cura a Carattere Scientifico Fondazione Casa Sollievo della Sofferenza. A.L.M. is supported by a Damon Runyon Cancer Research Foundation postdoctoral fellowship (no. DRG:2368-19) and a Postdoctoral Enrichment Program Award from the Burroughs Wellcome Fund (no. 1019903). C.F.K. is supported in part by grant no. R35HL150876-01, the Thoracic Foundation, Ellison Foundation, American Lung Association (no. LCD-619492) and the Harvard Stem Cell Institute. N.L-B. acknowledges funding from the European Research Council (consolidator grant no. 682398). P.H. is supported in part by the Association pour la Recherche contre le Cancer (CANC’AIR GENExposomics project). This work has been supported in part by the Tissue Core at the H. Lee Moffitt Cancer Center & Research Institute, a comprehensive cancer center designated by the National Cancer Institute and funded in part by a Moffitt Cancer Center Support Grant (no. P30-CA076292). B.E.G.R. is supported by NIH grant nos. 1P50 CA196530-01 and NIH 1K08 CA151645-01. We thank the Sherlock-Lung study scientific advisory board (M. Meyerson, J. Samet, M. Spitz, R. Summers, M. Thun and W. Travis) for their support. We also thank Y. Rubanova from Toronto University for her help with the TrackSig analysis. We thank the staff at the IUCPQ Université Laval Biobank, Nice Biobank Centre de Ressources Biologiques, Yale University and Moffitt Cancer Center & Research Institute for their valuable assistance in collecting samples and corresponding clinical data. This work utilized the computational resources of the NIH high-performance computational capabilities Biowulf cluster (http://hpc.nih.gov).
Author information
Authors and Affiliations
Contributions
M.T.L. and T.Z. conceptualized the study. T.Z., D.C.W., J. Shi., B.Z., N.A-P., N.L-B., B.Z., S.H.W., Y.P., H.C., T.R., D.R.S., D.A.G., L.B.A. and M.T.L. devised the methodology. T.Z., N.A-P., W.Z., P.H.H., R.L., K.H.-H., A.G.-P., F.M.-J., A.C., I.P., J. Sang, J. Shi, J.K., N.S., L.J.K., S.M.A.I., B.O., A.K., A.L.M. and C.F.K. carried out the formal analysis. A.H., N.C., J.C., D.H. and K.M.B. carried out the laboratory work. M.O., S.M.L., M.D., P.L., P.M.S.B. and J.S.A. carried out the pathology work. P.J., Y.B., P.H., D.C., A.C.P., L.A.M., B.E.G.R., M.L.P., M.C., M.B.S., N.E.C., M.L. and S.J.C. managed the resources. M.K., L.M. and J.R. curated the data. T.Z. and M.T.L. wrote the original draft. D.C.W., S.J.C., Y.B., Q.L., N.R., M.G-C., D.A.G., L.B.A., N.L-B., B.Z., J. Sang, J. Shi, T.Z., P.H.H. and M.T.L. reviewed and edited the draft. All authors carried out the data visualization. M.T.L. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
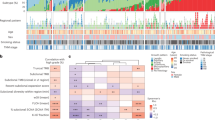
Extended Data Fig. 1 Genomic alterations of RTK-RAS pathway in Sherlock-Lung.
a, Oncoplot showing mutual exclusivity of genes within the RTK-RAS pathway, which were used to define the RTK-RAS status. The bottom bar shows tumor histological types. b, Comparison of genomic features between RTK-RAS negative and positive tumors. Left four panels: tumor mutational burden, percentage of genome with SCNAs, SV burden and T/N TL ratio. P-values are calculated using the two-sided Mann-Whitney U test; Middle three panels: enrichments for Kataegis events, WGD events, and BRCA2 LOH. P-values and OR are calculated using Fisher’s exact test (two-sided); Right panel: Contributions of each SBS signature.
Extended Data Fig. 2 Genomic alterations of TP53 pathway in Sherlock-Lung.
a, Oncoplot showing the mutual exclusivity between TP53 mutations and MDM2 amplification, which was used to define the TP53 proficient and deficient groups. The bottom bar shows tumor histological types. b, Comparison of genomic features between TP53-proficient and TP53-deficient tumors. Left three panels: tumor mutation burden, percentage of genome with SCNA and SV burden. P-values are calculated using the two-sided Mann-Whitney U test. Middle four panels: enrichments for BRCA1 LOH, Kataegis events, WGD events, and HLA LOH. P-values and OR are calculated using Fisher’s exact test (two-sided). Right panel: Contributions of each SBS signature.
Extended Data Fig. 3 Recurrence of SV breakpoints in Sherlock-Lung.
The frequencies of chromosomal breakpoints are calculated using 5 Mb as a window across the whole genome.
Extended Data Fig. 4 Summary of genomic features in LCINS based on different SCNA clusters.
Panels from top to bottom describe: 1) most frequently mutated or potential driver genes; 2) oncogenic fusions; 3) somatic mutations in surfactant associated genes; 4) significant focal SCNAs; 5) significant arm-level SCNAs; 6) genes with rare germline mutations; 7) and 8) other genomic features. The numbers on the right panel show the overall frequency (1-7) or median values (8). NRPCC: the number of reads per clonal copy.
Extended Data Fig. 5 Genes with signals of positive selection in Sherlock-Lung.
a, The scatter plot showing significantly mutated genes according to IntOGen q-value <0.05 (y-axis) and mutational frequency in the cohort (x-axis). Genes are colored according to their inferred mode of action in tumorigenesis. b, Recurrent non-synonymous driver mutations (in ≥2 patients).
Extended Data Fig. 6 Dominant endogenous processes in Sherlock-Lung.
a, Density plot of cosine similarity between original mutational profile and reconstructed mutational profile using reference signatures from (top to bottom): 65 COSMIC SBS signatures, 22 COSMIC SBS signatures for endogenous processes, 53 MutaGene SBS signatures of environmental exposures, and a combined set of signatures including the 22 endogenous and 53 environmental exposure signatures. b, Comparison of the cosine similarity between the original mutational profiles and reconstructed mutational profiles using endogenous and exogenous signatures (similar to a). Each dot represents one sample. The size and color represent the total number of mutations and tumor histological type, respectively.
Extended Data Fig. 7 Association between T/N TL ratio and somatic alterations in Sherlock-Lung.
a, Distribution of mean telomere lengths (TL) in Sherlock-Lung (dark blue, overall and by histological type), TCGA LUAD (green, overall and by smoking status) and TCGA other cancer types (Grey). Total sample numbers for each type are shown at the top. Error bars, 95% CIs from linear mixed model. b, Scatterplot showing association between T/N TL ratio and somatic alterations. Association P-values (two-sided t-test; FDR adjusted using Benjamini-Hochberg method) are shown on the y-axis. Genomic alterations with FDR <=0.1 or T/N TL ratio >1.1 or <0.9 are labeled and further highlighted in red when significant (FDR=0.05; horizontal dashed line). c, The proportion of each SCNA cluster among the group of tumors with somatic alterations significantly associated with shorten T/N TL including Chr22q Loss, Chr9p/q Loss or HLA LOH.
Extended Data Fig. 8 Homologous recombination deficiency (HRD) in Sherlock-Lung.
a, HRDetect scores of Sherlock-Lung samples. HRD-high: >0.7, HRD-low: < 0.005. b, Comparison of the number of total indels, microhomology deletions, SVs, and SNVs between samples with HRDetect score below 0.7 (group N) and above 0.7 (group Y). P-values are calculated using the two-sided Mann-Whitney U test. For box plots, center lines show the medians; box limits indicate the 25th and 75th percentiles; whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles.
Extended Data Fig. 9 Genomic alterations in HRD associated genes in Sherlock-Lung.
a, Oncoplot of genomic alterations in HRD associated genes, including germline mutations, somatic mutations and LOH. Samples with biallelic alterations are represented by bars with two different colors. The bottom bar shows tumor histological types. b, Boxplots of HRDetect scores (top) and SBS mutation loads (bottom) in tumors with and without LOH of six HR associated genes. FDR are calculated using the two-sided Mann-Whitney U test with multiple testing correction based on the Benjamini & Hochberg method. For box plots, center lines show the medians; box limits indicate the 25th and 75th percentiles; whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles.
Supplementary information
Supplementary Information
Supplementary Notes, Methods and Figs. 1–38.
Supplementary Tables
Supplementary Tables 1–8
Rights and permissions
About this article
Cite this article
Zhang, T., Joubert, P., Ansari-Pour, N. et al. Genomic and evolutionary classification of lung cancer in never smokers. Nat Genet 53, 1348–1359 (2021). https://doi.org/10.1038/s41588-021-00920-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-021-00920-0
This article is cited by
-
Shared and distinct mechanisms of UBA1 inactivation across different diseases
The EMBO Journal (2024)
-
Lung cancer in patients who have never smoked — an emerging disease
Nature Reviews Clinical Oncology (2024)
-
Distribution and prognostic impact of EGFR and KRAS mutations according to histological subtype and tumor invasion status in pTis-3N0M0 lung adenocarcinoma
BMC Cancer (2023)
-
PSMD3-ILF3 signaling cascade drives lung cancer cell proliferation and migration
Biology Direct (2023)
-
New insights into the biology and development of lung cancer in never smokers—implications for early detection and treatment
Journal of Translational Medicine (2023)