Abstract
Microglia, the tissue-resident macrophages of the central nervous system (CNS), play critical roles in immune defense, development and homeostasis. However, isolating microglia from humans in large numbers is challenging. Here, we profiled gene expression variation in primary human microglia isolated from 141 patients undergoing neurosurgery. Using single-cell and bulk RNA sequencing, we identify how age, sex and clinical pathology influence microglia gene expression and which genetic variants have microglia-specific functions using expression quantitative trait loci (eQTL) mapping. We follow up one of our findings using a human induced pluripotent stem cell-based macrophage model to fine-map a candidate causal variant for Alzheimer’s disease at the BIN1 locus. Our study provides a population-scale transcriptional map of a critically important cell for human CNS development and disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Patients were consented to share both expression and raw genotype data under managed access, and all data are available under managed access from the EGA, upon approval by the Wellcome Sanger Institute Data Access Committee. More details on how to access these data can be found at https://ega-archive.org/datasets/EGAD00001005736. Raw data (fastq files and CRAM files) of Smart-seq2 and bulk RNA-seq for the primary microglia samples, as well as the raw genotype data (Illumina Omni 2.5) and imputed genotype data by Beagle software, are available from the European Phenome-Genome Archive (EGA) (accession ID: EGAD00001005736). Summary statistics of eQTLs mapped by linear regression and RASQUAL for primary microglia are also available from EGA (accession ID: EGAD00001005736). The 1000 Genomes Phase III integrated variant set can be obtained from the project website (http://www.internationalgenome.org/data). GTEx v.7 summary statistics and brain DroNc-seq data with cell type annotation data can be obtained from the GTEx project website (https://www.gtexportal.org/home/datasets). PBMC 68,000 single-cell data are available from the project GitHub page (https://github.com/10XGenomics/single-cell-3prime-paper/tree/master/pbmc68k_analysis). The BLUEPRINT monocyte RNA-seq data are available from EGA (accession ID: EGAD00001002674). For details on how to access these data, please visit https://ega-archive.org/datasets/EGAD00001002674.
References
Schafer, D. P. & Stevens, B. Microglia function in central nervous system development and plasticity. Cold Spring Harb. Perspect. Biol. 7, a020545 (2015).
Li, Q. & Barres, B. A. Microglia and macrophages in brain homeostasis and disease. Nat. Rev. Immunol. 18, 225–242 (2018).
Salter, M. W. & Stevens, B. Microglia emerge as central players in brain disease. Nat. Med. 23, 1018–1027 (2017).
Guerreiro, R. et al. TREM2 variants in Alzheimer’s disease. N. Engl. J. Med. 368, 117–127 (2013).
Jonsson, T. et al. Variant of TREM2 associated with the risk of Alzheimer’s disease. N. Engl. J. Med. 368, 107–116 (2013).
Tansey, K. E., Cameron, D. & Hill, M. J. Genetic risk for Alzheimer’s disease is concentrated in specific macrophage and microglial transcriptional networks. Genome Med. 10, 14 (2018).
Gjoneska, E. et al. Conserved epigenomic signals in mice and humans reveal immune basis of Alzheimer’s disease. Nature 518, 365–369 (2015).
Olah, M. et al. A transcriptomic atlas of aged human microglia. Nat. Commun. 9, 539 (2018).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290.e17 (2017).
Hammond, T. R. et al. Single-cell RNA sequencing of microglia throughout the mouse lifespan and in the injured brain reveals complex cell-state changes. Immunity 50, 253–271.e6 (2019).
Masuda, T. et al. Spatial and temporal heterogeneity of mouse and human microglia at single-cell resolution. Nature 566, 388–392 (2019).
Mrdjen, D. et al. High-dimensional single-cell mapping of central nervous system immune cells reveals distinct myeloid subsets in health, aging, and disease. Immunity 48, 380–395.e6 (2018).
Mathys, H. et al. Temporal tracking of microglia activation in neurodegeneration at single-cell resolution. Cell Rep. 21, 366–380 (2017).
Kilpinen, H. et al. Common genetic variation drives molecular heterogeneity in human iPSCs. Nature 546, 370–375 (2017).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science 356, eaal3222 (2017).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Mathys, H. et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570, 332–337 (2019).
Sankowski, R. et al. Mapping microglia states in the human brain through the integration of high-dimensional techniques. Nat. Neurosci. 22, 2098–2110 (2019).
Kumasaka, N., Knights, A. J. & Gaffney, D. J. Fine-mapping cellular QTLs with RASQUAL and ATAC-seq. Nat. Genet. 48, 206–213 (2016).
Jansen, I. et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 51, 404–413 (2019).
Kunkle, B. W. et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat. Genet. 51, 414–430 (2019).
Lambert, J. C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
Marioni, R. E. et al. GWAS on family history of Alzheimer’s disease. Transl. Psychiatry 8, 99 (2018).
Schwartzentruber, J. et al. Genome-wide meta-analysis, fine-mapping and integrative prioritization implicate new Alzheimer’s disease risk genes. Nat. Genet. 53, 392–402 (2021).
Raj, T. et al. CD33: increased inclusion of exon 2 implicates the Ig V-set domain in Alzheimer’s disease susceptibility. Hum. Mol. Genet. 23, 2729–2736 (2014).
Alasoo, K. et al. Shared genetic effects on chromatin and gene expression indicate a role for enhancer priming in immune response. Nat. Genet. 50, 424–431 (2018).
Barbosa, A. C. et al. MEF2C, a transcription factor that facilitates learning and memory by negative regulation of synapse numbers and function. Proc. Natl Acad. Sci. USA 105, 9391–9396 (2008).
Nott, A. et al. Brain cell type-specific enhancer-promoter interactome maps and disease-risk association. Science 366, 1134–1139 (2019).
Vela, J. M., Yáñez, A., González, B. & Castellano, B. Time course of proliferation and elimination of microglia/macrophages in different neurodegenerative conditions. J. Neurotrauma 19, 1503–1520 (2002).
Abud, E. M. et al. iPSC-derived human microglia-like cells to study neurological diseases. Neuron 94, 278–293.e9 (2017).
Okbay, A. et al. Genetic variants associated with subjective well-being, depressive symptoms, and neuroticism identified through genome-wide analyses. Nat. Genet. 48, 624–633 (2016).
Luciano, M. et al. Association analysis in over 329,000 individuals identifies 116 independent variants influencing neuroticism. Nat. Genet. 50, 6–11 (2018).
Alasoo, K. et al. Transcriptional profiling of macrophages derived from monocytes and iPS cells identifies a conserved response to LPS and novel alternative transcription. Sci. Rep. 5, 12524 (2015).
Douvaras, P. et al. Directed differentiation of human pluripotent stem cells to microglia. Stem Cell Rep. 8, 1516–1524 (2017).
Muffat, J. et al. Efficient derivation of microglia-like cells from human pluripotent stem cells. Nat. Med. 22, 1358–1367 (2016).
Zhang, H. et al. Functional analysis and transcriptomic profiling of iPSC-derived macrophages and their application in modeling Mendelian disease. Circ. Res. 117, 17–28 (2015).
Chen, L. et al. Genetic drivers of epigenetic and transcriptional variation in human immune cells. Cell 167, 1398–1414.e24 (2016).
Jiang, H., Lei, R., Ding, S.-W. & Zhu, S. Skewer: a fast and accurate adapter trimmer for next-generation sequencing paired-end reads. BMC Bioinform. 15, 182 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Kumasaka, N., Knights, A. J. & Gaffney, D. J. High-resolution genetic mapping of putative causal interactions between regions of open chromatin. Nat. Genet. 51, 128–137 (2019).
Kang, H. M. et al. Multiplexed droplet single-cell RNA-sequencing using natural genetic variation. Nat. Biotechnol. 36, 89–94 (2018).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Haghverdi, L., Lun, A. T. L., Morgan, M. D. & Marioni, J. C. Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. https://doi.org/10.1038/nbt.4091 (2018)
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Veyrieras, J.-B. et al. High-resolution mapping of expression-QTLs yields insight into human gene regulation. PLoS Genet. 4, e1000214 (2008).
Acknowledgements
R.F. was supported by funding from the UK Multiple Sclerosis Society (MS50), the Adelson Medical Research Foundation and a core support grant from the Wellcome Trust and MRC to the Wellcome Trust-Medical Research Council Cambridge Stem Cell Institute (203151/Z/16/Z). A.Y. is supported by a Wellcome Trust Clinicians PhD Fellowship (RRZD/029). All data for this study were generated under Open targets project OTAR039. N.K. and D.J.G. were funded by the Wellcome Trust grant WT206194. We thank the staff in the Cellular Genetics and Phenotyping and Sequencing core facilities at the Wellcome Sanger Institute.
Author information
Authors and Affiliations
Contributions
The brain samples were obtained under neurosurgery by A.M.H.Y., H.B., J.C., K.P.B., A.J., M. R. Guilfoyle, R.T., R.K., R. Morris, M. R. Garnett, I.T., I.J., K.H., R. Mannion, R. Mair, C.W., S.J.P., P.J.K., T.S. and P.J.H. Cell isolation protocols were performed by A.M.H.Y., M.S., N.A.M. and C.E.M. Microglia isolation strategies were designed by A.M.H.Y., T.R.H., B.S. and R.J.M.F. Single-cell and bulk RNA-seq were performed by A.M.H.Y., A.K. and F.C. Cell culture experiments iPSDMacs were performed by A.K. and N.P. The main analyses and data preparations were performed by N.K. E.M. and M.G. processed GWAS summary statistics for colocalization analysis. J.S. and J.L. provided the summary statistics of Alzheimer’s disease. K.K. and N.S. provided the imputed BLUEPRINT genotype data. N.P. preprocessed the iPSDMac RNA-seq data. J.S.P. and O.A.B. performed the RNAscope and immunohistochemistry assay. A.M.H.Y., N.K., R.J.M.F. and D.J.G. wrote the manuscript. T.R.H. and B.S. assisted in editing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
D.J.G. and E.M. were employees of Genomics PLC at the time the manuscript was submitted.
Additional information
Peer review information Nature Genetics thanks Julie Williams and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Overview of bulk and single-cell RNA-seq data.
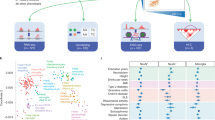
a, UMAP of bulk RNA-seq for myeloid cells. The ‘Primary microglia’ cluster contains samples collected in this study (pink dots) and previous studies (purple dots) (information on the source of previous study data can be found in Supplementary Table 7). ‘Cultured primary and IPS-derived cells’, includes IPS-derived macrophages and microglia (blue dots), cultured primary microglia and monocyte derived macrophages (orange dots). ‘Monocytes’ (green dots) denotes primary monocytes obtained from the BLUEPRINT project. b, Feature plots of three microglia marker genes (P2RY12, CX3CR1 and TMEM119) using the same UMAP coordinates as Fig. 1d. c, Age versus percentage of infiltrating cells. Red line shows the logistic regression line, the red transparent band shows the 95% confidence interval estimated using a generalised linear mixed model for the binary outcome (Materials and Methods). d, UMAP plot identical to Fig. 1d. e, UMAP plot from the first 12 principal components computed from the same input data for the linear mixed model without any batch correction. f, UMAP of the same 12 PCs where the batch effect was corrected by using Harmony47. g, UMAP of batch corrected data using the canonical correlation analysis method implemented in Seurat V348 with a default setting. We computed the 12 PCs from the integrated data for UMAP plot. h, UMAP of batch corrected data using MNN correct49. Note that points were coloured according to the cell types (same as Fig. 1d): glutamatergic neurons from the PFC (exPFC); pyramidal neurons from the hip CA region (exCA); GABAergic interneurons (GABA); granule neurons from the hip dentate gyrus region (exDG); astrocytes (ASC); oligodendrocytes (ODC); oligodendrocyte precursor cells (OPC); neuronal stem cells (NSC); endothelial cells (END); dendritic cell (DC); B cell (B); hematopoietic progenitor cell (CD34 + ); NK T cell (NK).
Extended Data Fig. 2 Microglia marker gene comparisons and validations.
a, Marker gene enrichment analysis with Alzheimer’s disease associated microglia20 and glioma associated microglia21. There are three different comparisons for Alzheimer’s disease associated microglia and 14 different populations for glioma associated microglia. Heatmap shows odds ratios and Benjamini-Hochberg (BH) Q-values of the Fisher exact tests between our marker genes and differentially expressed genes in other studies. b, Differentially expressed genes between microglia from different patient pathologies using single cell RNA-seq data. Heatmap shows averaged, normalised expression level (defined as the posterior mean of pathology random effect term, see Materials and Methods) of differentially expressed genes at local true sign rate (ltsr) greater than 0.9 ((Urbut et al. 2019); see Materials and Methods for details). Heatmap is divided into groups based on all possible pairwise groupings of the four cell populations, ordered by most transcriptionally distinct, such that the most different grouping, trauma versus all non-trauma, appears at the top. c, Differential expression of candidate marker genes for immunohistochemistry in fresh frozen patient tissue samples. d, Immunohistochemistry panel of each pathology to validate expression of a differentially expressed gene at the protein level; hydrocephalus (C3), tumour (CCL4), haemorrhage (CD63) and trauma (BIN-1) compared to control. Iba-1 (red) and protein of interest (green). e, RNAScope image of differentially expressed gene panel for cluster C; HAMP (yellow) and RAC2 (purple) with C1QC (green) used to identify microglia. f, RNAScope image of differentially expressed gene panel for cluster D; KLF (yellow) and CCL20 (purple). Scale bar 10 µM.
Extended Data Fig. 3 Differential expression analysis with bulk RNA-seq data.
a, Variance components analysis of log CPM values for the bulk RNA-seq data (N = 102) with biological and technical factors using the linear mixed model (Online methods). b, Heatmap shows the effect size of age for each gene (each row) estimated by the linear mixed model (Online methods). The genes with LTSR > 0.9 in single-cell data are shown. c, PADI2 normalised expression in bulk RNA-seq data against patients’ age. d, P2RY12 expression in bulk RNA-seq data against patients’ age. e, Heatmap shows the average expression of males and females for each gene (each row) estimated by the linear mixed model (Online methods). The genes with LTSR > 0.9 in single-cell data are shown. f, C1QA normalised expression in bulk RNA-seq data for males (M) and females (F). g, HLA-DQB1 normalised expression in bulk RNA-seq data for males (M) and females (F). h, Heatmap shows the average expression for 5 different brain regions estimated by the linear mixed model (Online methods). The genes differentially expressed between a combination of Occipital and Cerebellum and the 3 other regions (LTSR > 0.9) in single-cell data are shown.
Extended Data Fig. 4 Colocalisation of eQTLs with various GWAS traits.
a, eQTL effect size comparison for 502 eQTL genes at FDR 5% (linear regression) whose gene body contains at least one feature SNP with sufficient coverage (greater than 5% of average coverage across coding regions). The x-axis shows the eQTL effect size (beta) estimated from linear regression and the y-axis shows the eQTL effect size (pi value) from RASQUAL using only allele-specific count data. The red line shows the least square line crossing (0, 0.5). Note that, x = 0 is the null hypothesis for linear regression and y = 0.5 is the null hypothesis for RASQUAL. b, Examples of colocalised eQTLs in microglia. Colocalisation with Parkinson’s disease at KLHL7-AS1 eQTL (left column), colocalisation with Fed-up feelings at DAG1 eQTL (middle column) and colocalisation with Crohn’s disease at ERAP2 eQTL (right column). The y-axis of each panel shows log10 association Bayes factor for the eQTL or the GWAS trait. The colour of each point indicates LD index (r2 value) to the lead eQTL variant shown by the purple diamond. c, Heatmap of the posterior probability for colocalisation (PP4) between various GWAS traits and cell types/tissues. Each row corresponds to a specific combination of gene and a GWAS trait. Each column corresponds to eQTLs discovered in different cell types and tissues. The first column of the heatmap corresponds to microglia eQTLs, the second column corresponds to eQTLs in IPS cell derived macrophage (IPSDMac) from this study (Materials and Methods), the third column shows eQTLs in primary monocytes from the BLUEPRINT project (Materials and Methods) and the remaining 48 tissues are eQTLs from GTEx V7 (Materials and Methods). The colour of each grid shows the strength of PP4 (white: PP4 = 0.0 and red: PP4 = 1.0). Gray indicates that the gene was very weakly or not expressed, and therefore no eQTL summary statistics were available.
Extended Data Fig. 5 Fine mapping of microglia eQTLs.
a, Regional association plots at the CD33 locus. b, Coverage plot shows the normalised expression level around the CD33 gene stratified by genotype at the putative splice variant (rs12459419C > T). The zoom-in panel shows a coverage plot of expression level around the second exon (ENST0000262262.4). The coverage shows the first intron expression is negatively correlated with the second exon expression, suggesting the expression of non-coding isoform (ENST00000601785.5) is increased by the alternative allele (T) of the splicing QTL. c, Colocalisation between an association with risk for Alzheimer’s disease on chromosome 2 and an eQTL for the noncoding RNA gene EPHA1-AS1 in microglia, GTEx tissues and myeloid cell types. The x-axis shows the posterior probability of colocalisation (PP4) and y-axis shows the average expression level (log10 TPM) for each tissue or cell type. d, Colocalisation between AD risk and expression of the protein-coding EPHA1 gene. The x-axis shows the posterior probability of colocalisation (PP4) and y-axis shows the average expression level (log10 TPM) for each tissue or cell type. e, Boxplots show the relationship between expression at the PTK2B gene and genotype at the lead eQTL variant (rs28834970C > T) three myeloid cell types. The y-axis shows normalised expression levels (log TPM value). Each dot on the box shows the expression level of a single sample. f, Coverage plot shows chromatin accessibility in iPS cell derived macrophages stratified by three genotype groups of the lead AD GWAS/BIN1 eQTL variant g, Scatter plot of MEF2C (x-axis) and BIN1 (y-axis) expression in GTEx brain tissues and myeloid cell type.
Supplementary information
Supplementary Information
Supplementary Methods
Supplementary Tables
Supplementary Tables 1–7
Rights and permissions
About this article
Cite this article
Young, A.M.H., Kumasaka, N., Calvert, F. et al. A map of transcriptional heterogeneity and regulatory variation in human microglia. Nat Genet 53, 861–868 (2021). https://doi.org/10.1038/s41588-021-00875-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-021-00875-2
This article is cited by
-
Mapping the functional impact of non-coding regulatory elements in primary T cells through single-cell CRISPR screens
Genome Biology (2024)
-
Human skeletal muscle aging atlas
Nature Aging (2024)
-
Cell subtype-specific effects of genetic variation in the Alzheimer’s disease brain
Nature Genetics (2024)
-
Integrative transcriptomic analysis of the amyotrophic lateral sclerosis spinal cord implicates glial activation and suggests new risk genes
Nature Neuroscience (2023)
-
Systematic differences in discovery of genetic effects on gene expression and complex traits
Nature Genetics (2023)