Abstract
Ultraconserved enhancer sequences show perfect conservation between human and rodent genomes, suggesting that their functions are highly sensitive to mutation. However, current models of enhancer function do not sufficiently explain this extreme evolutionary constraint. We subjected 23 ultraconserved enhancers to different levels of mutagenesis, collectively introducing 1,547 mutations, and examined their activities in transgenic mouse reporter assays. Overall, we find that the regulatory properties of ultraconserved enhancers are robust to mutation. Upon mutagenesis, nearly all (19/23, 83%) still functioned as enhancers at one developmental stage, as did most of those tested again later in development (5/9, 56%). Replacement of endogenous enhancers with mutated alleles in mice corroborated results of transgenic assays, including the functional resilience of ultraconserved enhancers to mutation. Our findings show that the currently known activities of ultraconserved enhancers do not necessarily require the perfect conservation observed in evolution and suggest that additional regulatory or other functions contribute to their sequence constraint.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Images of all transgenic whole-mount-stained embryos are included in Supplementary Fig. 1. Images of brains sections from knock-in animals and wild-type littermates are in Extended Data Figs. 8, 9 and 10. Human single-nucleotide variants were obtained from TOPMed (https://bravo.sph.umich.edu/freeze5/hg38) in June 2020. JASPAR transcription factor motif data were downloaded from http://expdata.cmmt.ubc.ca/JASPAR/downloads/UCSC_tracks/2018/hg19. Public chromatin immunoprecipitation sequencing data were obtained from https://www.encodeproject.org (mouse embryonic H3K27ac and H3K27me3, mouse and human CTCF) and https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE124936 (DLX transcription factors). The cloning vector for the transgenic assay (PCR4-Shh::lacZ-H11) is available from Addgene (catalog no. 139098). All other vectors described in the present study are available from the authors upon request. Transgenic embryos and stable knock-in lines can also be made available upon request.
References
Bejerano, G. et al. Ultraconserved elements in the human genome. Science 304, 1321–1325 (2004).
Hecker, N. & Hiller, M. A genome alignment of 120 mammals highlights ultraconserved element variability and placenta-associated enhancers. Gigascience 9, giz159 (2020).
Katzman, S. et al. Human genome ultraconserved elements are ultraselected. Science 317, 915 (2007).
Drake, J. A. et al. Conserved noncoding sequences are selectively constrained and not mutation cold spots. Nat. Genet. 38, 223–227 (2006).
Ovcharenko, I. Widespread ultraconservation divergence in primates. Mol. Biol. Evol. 25, 1668–1676 (2008).
Habic, A. et al. Genetic variations of ultraconserved elements in the human genome. OMICS 23, 549–559 (2019).
Pennacchio, L. A. et al. In vivo enhancer analysis of human conserved non-coding sequences. Nature 444, 499–502 (2006).
Visel, A. et al. Ultraconservation identifies a small subset of extremely constrained developmental enhancers. Nat. Genet. 40, 158–160 (2008).
Dickel, D. E. et al. Ultraconserved enhancers are required for normal development. Cell 172, 491–499 e15 (2018).
Nolte, M. J. et al. Functional analysis of limb transcriptional enhancers in the mouse. Evol. Dev. 16, 207–223 (2014).
Ahituv, N. et al. Deletion of ultraconserved elements yields viable mice. PLoS Biol. 5, e234 (2007).
Gaynor, K. U. et al. Studies of mice deleted for Sox3 and uc482: relevance to X-linked hypoparathyroidism. Endocr. Connect. 9, 173–186 (2020).
Chen, C. T., Wang, J. C. & Cohen, B. A. The strength of selection on ultraconserved elements in the human genome. Am. J. Hum. Genet. 80, 692–704 (2007).
Kryukov, G. V., Schmidt, S. & Sunyaev, S. Small fitness effect of mutations in highly conserved non-coding regions. Hum. Mol. Genet. 14, 2221–2229 (2005).
Keightley, P. D., Kryukov, G. V., Sunyaev, S., Halligan, D. L. & Gaffney, D. J. Evolutionary constraints in conserved nongenic sequences of mammals. Genome Res. 15, 1373–1378 (2005).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005).
Harmston, N., Baresic, A. & Lenhard, B. The mystery of extreme non-coding conservation. Philos. Trans. R. Soc. Lond. B Biol. Sci. 368, 20130021 (2013).
Patwardhan, R. P. et al. Massively parallel functional dissection of mammalian enhancers in vivo. Nat. Biotechnol. 30, 265–270 (2012).
Melnikov, A. et al. Systematic dissection and optimization of inducible enhancers in human cells using a massively parallel reporter assay. Nat. Biotechnol. 30, 271–277 (2012).
Dickel, D. E., Visel, A. & Pennacchio, L. A. Functional anatomy of distant-acting mammalian enhancers. Philos. Trans. R. Soc. Lond. B Biol. Sci. 368, 20120359 (2013).
Kircher, M. et al. Saturation mutagenesis of twenty disease-associated regulatory elements at single base-pair resolution. Nat. Commun. 10, 3583 (2019).
Lettice, L. A., Devenney, P., De Angelis, C. & Hill, R. E. The conserved sonic hedgehog limb enhancer consists of discrete functional elements that regulate precise spatial expression. Cell Rep. 20, 1396–1408 (2017).
Canver, M. C. et al. BCL11A enhancer dissection by Cas9-mediated in situ saturating mutagenesis. Nature 527, 192–197 (2015).
Kvon, E. Z. et al. Comprehensive in vivo interrogation reveals phenotypic impact of human enhancer variants. Cell 180, 1262–1271 (2020).
Karolchik, D. et al. The UCSC table browser data retrieval tool. Nucleic Acids Res. 32, D493–D496 (2004).
Hinrichs, A. S. et al. The UCSC genome browser database: update 2006. Nucleic Acids Res. 34, D590–D598 (2006).
Chiang, C. W. et al. Ultraconserved elements: analyses of dosage sensitivity, motifs and boundaries. Genetics 180, 2277–2293 (2008).
Osterwalder, M. et al. Enhancer redundancy provides phenotypic robustness in mammalian development. Nature 554, 239–243 (2018).
Turner, T. N. et al. Genomic patterns of de novo mutation in simplex autism. Cell 171, 710–722.e12 (2017).
Fakhouri, W. D. et al. An etiologic regulatory mutation in IRF6 with loss- and gain-of-function effects. Hum. Mol. Genet. 23, 2711–2720 (2014).
Viturawong, T., Meissner, F., Butter, F. & Mann, M. A DNA-centric protein interaction map of ultraconserved elements reveals contribution of transcription factor binding hubs to conservation. Cell Rep. 5, 531–545 (2013).
McCole, R. B., Erceg, J., Saylor, W. & Wu, C. T. Ultraconserved elements occupy specific arenas of three-dimensional mammalian genome organization. Cell Rep. 24, 479–488 (2018).
Pollard, K. S., Hubisz, M. J., Rosenbloom, K. R. & Siepel, A. Detection of nonneutral substitution rates on mammalian phylogenies. Genome Res. 20, 110–121 (2010).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Kvon, E. Z. et al. Progressive loss of function in a limb enhancer during snake evolution. Cell 167, 633–642.e11 (2016).
Montague, T. G., Cruz, J. M., Gagnon, J. A., Church, G. M. & Valen, E. CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res. 42, W401–W407 (2014).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Khan, A. et al. JASPAR 2018: update of the open-access database of transcription factor binding profiles and its web framework. Nucleic Acids Res. 46, D1284 (2018).
Acknowledgements
This work was supported by the US National Institutes of Health (grant no. R01HG003988 to L.A.P.), the National Institute of Neurological Disorders and Stroke (grant no. R01NS034661 to J.L.R.R.) and the National Institute of Mental Health (grant no. R01MH049428 to J.L.R.R.). A.R.Y. was supported by a grant from Fondation Fyssen. Research was conducted at the E.O. Lawrence Berkeley National Laboratory and performed under US Department of Energy contract DE-AC02-05CH11231, University of California. We thank F. Darbellay and S. Rajderkar for help with embryo scoring. We also thank J. Hu (UCSF) for kindly providing the VIP riboprobe.
Author information
Authors and Affiliations
Contributions
V.S., A.R.Y., B.J.M., I.P.-F., C.S.N., A.N.H., Q.T.P., M.K., E.Z.K., Y.Z., M.S., R.D.H., E.M., J.G., J.A.A., S.T. and V.A. performed experiments, including making and/or characterizing all transgenic and knock-in mouse lines. V.S., D.E.D., A.V., L.A.P., A.R.Y. and J.L.R.R. planned the study and wrote the manuscript with input from the remaining authors.
Corresponding authors
Ethics declarations
Competing interests
J.L.R.R. is a cofounder and stockholder, and currently on the scientific board of Neurona, a company studying the potential therapeutic use of interneuron transplantation. The other authors declare no competing interests.
Additional information
Peer review information Nature Genetics thanks Menno Creyghton, Stephen Gisselbrecht, Katherine Pollard and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of base pairs selected for mutagenesis.
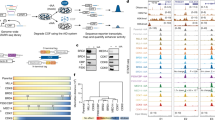
a, Detailed breakdown of conservation (phylop100way score) and position of base pair mutations for selected examples of ultraconserved enhancers mutated in this study. Base pairs selected to achieve various levels of mutagenesis: 2%, 5%, 20%, or all base pairs conserved among ~100 vertebrates are marked according to the legend. All coordinates are in hg19. Locations of conserved transcription factor motifs26 and human SNVs (TOPMed, https://bravo.sph.umich.edu/freeze5/hg38/) are also included (see legend). See Supplementary Information for similar plots of all 23 ultraconserved enhancers selected for mutagenesis. b, Percentage of mutations introduced into ultraconserved sequences that also overlapped human SNVs (all overlap rare human variants, that is found in <1% of human population. No introduced mutations overlap a SNV common in the human population).
Extended Data Fig. 2 Example of enhancer scoring.
Top: calibration images given for one ultraconserved enhancer (hs111) to show different categories of enhancer activity. Randomized embryo images were shown to five reviewers alongside the calibration images displayed at the top. The reviewers scored the strength of enhancer activity independently and blind to the identity of the enhancer allele. Bottom: breakdown of enhancer activity scoring results per embryo for four different alleles tested for the hs111 enhancer: reference allele, allele with 5% of bp mutated #1, allele with 5% of bp mutated #2, and allele with 20% of bp mutated. See Supplementary Information for a similar scoring breakdown of all 23 ultraconserved enhancers selected for mutagenesis.
Extended Data Fig. 3 Distribution of 23 ultraconserved enhancers among the three groups of enhancer mutagenesis outcomes.
a, Images of embryos and enhancer activity results for all 23 enhancers. Bar plots under the embryo images show the strength and reproducibility of LacZ staining (serving as a proxy for enhancer activity) among individual transgenic embryos from each enhancer allele. Enhancer activity was scored by five independent reviewers blind to the embryo genotype after the images were randomized (see Methods, Extended Data Fig. 2 and Supplementary Information for details). Embryo images for CRISPR-assisted transgenic experiments for the reference alleles of enhancers hs200 and hs215 were published previously24. All other transgenic assays were newly performed for this study. All of the mutation alleles tested harbored sequence changes at 2, 5, or 20% of ultraconserved bases, as indicated above, with the exception of three of the alleles grouped with the 20% variants (marked with an asterisk [*] for hs122, hs200, and hs267). For the hs122 allele, 25% of bp were actually mutated. For hs200 (16% mutation) and hs267 (12% mutation), fewer than 20% of base pairs are well-conserved among ~100 vertebrates, so lower levels of mutagenesis were used. CN, cranial nerve; DRG, dorsal root ganglia; EY, eye; FB, forebrain; HB, hindbrain; HT, heart; LB, limb; MB, midbrain; NT, neural tube. b, Length of ultraconserved sequences for all 23 enhancers arranged by enhancer mutagenesis outcomes with mean and median shown per group as solid and dashed lines, respectively. No statistically significant differences detected between groups (two-tailed t-tests). bp, base pairs.
Extended Data Fig. 4 Motif enrichment analysis.
a, Motifs that are significantly enriched in 370 noncoding ultraconserved sequences compared to the whole human genome. b, Motifs that are significantly enriched in 15 base pair k-mers centered on introduced mutations compared to the whole human genome. c, Left: Bar plot comparing the number of transcription factor motifs in the JASPAR database that overlap introduced mutations in alleles of the same enhancer that either decreased enhancer activity (grey bars) or not (blue bars). Each allele has 5% of ultraconserved base pairs mutated. Right: Boxplot summarizing data from all eight enhancers shown in the bar plot. Data is presented as median in the center, first (25th percentile) and third (75th percentile) quartile as bounds of the boxes, and ± 1.5 x ICR (interquartile range = third – first quartile) from bounds of the boxes as whiskers. d, Left: Bar plot showing the percentage of JASPAR transcription factor motifs overlapping 23 enhancers mutated in this study that occur in the ultraconserved sequence of an enhancer only once (unique, black) or multiple times (redundant, grey). On average, 48% of motifs were unique. Right: Bar plot comparing distribution of unique vs. redundant transcription factor motifs that overlap introduced mutations between mutant alleles that resulted in decreased enhancer activity (grey bars) and those that did not (blue bars). On average, ~52% of motifs that overlapped introduced mutations that did not decrease enhancer activity were unique, while the average for unique motifs that overlapped introduced mutations that decreased enhancer activity was ~44%.
Extended Data Fig. 5 Chromatin signatures of 23 ultraconserved enhancers mutated in this study in multiple tissues and developmental times points in mouse and human.
a, Histone modifications associated with active enhancers (H3K27ac) and inactive sequences (H3K27me3) sampled from multiple tissues throughout mouse development. ‘+‘ and ‘-‘ for ‘Tg+ e11.5’ indicate the presence or absence, respectively, of the activity by the reference enhancer allele in e11.5 (e12.5 for hs124) transgenic mouse embryos reported in this study. ChIP-seq data was obtained from Gorkin et al., 202040. b, Binding of Dlx transcription factors during embryonic development of mouse basal ganglia. ChIP-seq data was obtained from Lindtner et al., 201941. c, Binding of CTCF in embryonic and adult mouse tissues and adult human tissues. ChIP-seq data was obtained from Yue et al., 201442 for mouse samples and from Dunham et al., 201243 for human samples.
Extended Data Fig. 6 Knock-in strategy to introduce the mutations into the endogenous mouse alleles of hs122 (uc468+uc469) and hs121 (uc467).
a, e, The mutant enhancer alleles were knocked-in via homologous recombination of the reference allele via a CRISPR/Cas9 microinjection protocol. A sgRNA for hs122 was designed to target the reference allele sequence that overlaps mutations introduced into the mutant allele. The target sequence of a sgRNA for hs121 was 115 bp upstream of the ultraconserved sequence, thus hs121 repair templates carried a point mutation altering the PAM sequence for the sgRNA (schematically indicated by the pink lines) to prevent their cleavage by the CRISPR/Cas9 complex. b, f, The genomic region between the Arx and Pola1 genes contains hs122, and hs121 is located in the first intron of the Pola1 gene. Conservation tracks (green) show phastCons scores between mouse and other vertebrate genomes. c, g, Positions of the ultraconserved sequences within enhancers (uc468+uc469 OR uc467), the sequences tested in the transgenic assay, the homologous recombination templates, and genotyping primers are shown schematically in black. Position of the sgRNA target sites shown in pink. d, h, Males hemizygous for the mutant alleles of the hs122 and hs121 enhancers were genotyped with PCR amplification (positions of genotyping primers shown as black triangles in c and g) and Sanger sequencing. Top: alignment of reads from the knock-in alleles and wild-type alleles to the reference mouse genome (mm10) is shown in grey, with mismatches indicating the location of the introduced mutations represented by red lines. Bottom: Sanger sequencing traces for mutant knock-in and wild-type alleles are shown for the region within the dotted lines. Introduced mutations are highlighted in red. See Supplementary Tables 4, 5 for more information on coordinates and sequences.
Extended Data Fig. 7 Transgenic reporter assays with either human or mouse sequences surrounding the ultraconserved core show the same effect on the enhancer activity.
The sequences surrounding the ultraconserved parts of the (a) hs122 (uc468+uc469) and (b) hs121 (uc467) enhancers tested in transgenic assays differ between the human and mouse genomes (mismatches shown in green). To make sure that mouse regions flanking the ultraconserved sequences (shown in red) will not compensate for mutations introduced into the endogenous mouse alleles of the enhancers, we tested the activity of the mouse sequence in transgenic assays for both the reference alleles and the alleles with 5% of base pairs mutated. Regions to test were selected to span the hs122 and hs121 sequences deleted from the mouse genome previously9.
Extended Data Fig. 8 Arx expression in mice with mutated endogenous hs121 and hs122 enhancers.
In situ hybridization for Arx expression (purple staining) performed on coronal cross-sections of e12.5 forebrains from hemizygous knock-in males shown alongside wild-type littermate controls from three knock-in lines: (a) hs122mut (allele with 5% of base pairs mutated that eliminated enhancer activity in mouse transgenic assay), (b) hs121mut1 (allele with 5% of base pairs mutated that did not affect enhancer activity in mouse transgenic assay), and (c) hs121mut2 (allele with 5% of base pairs mutated that eliminated enhancer activity in mouse transgenic assay). Arrows point to a loss of Arx signal in the caudal cortical regions of hs122mut/Y sections. We did not detect changes in Arx expression in hemizygous knock-in male embryos for either of the hs121 enhancer mutant alleles (hs121mut1/Y and hs121mut2/Y). However, this is consistent with the data from hs121-null embryos9, and is likely due to tissue heterogeneity and the presence of another Arx enhancer (hs119) with a similar activity domain. Scale bars, 200 μm.
Extended Data Fig. 9 Mutagenesis of the endogenous hs122 enhancer results in dentate gyrus abnormalities (a smaller dentate gyrus with disorganized appearance).
Rostral (a) and caudal (b) coronal cross-sections of postnatal hippocampus from all phenotyped animals: three wild-type littermate male controls and four hemizygous knock-in males shown side-by-side for comparison. Numbers represent unique identifiers for each mouse examined (details in Supplementary Table 6). Scale bars, 500 μm. c, Bar graph showing quantification of dentate gyrus length for wild-type mice (n = 3) and knock-in mice (n = 4). Bars indicate mean lengths, with individual biological replicates represented by dots. Error bars, mean ± s.d. Difference in length between wild-type and knock-in mice assessed by a two-tailed paired t-test.
Extended Data Fig. 10 Mutagenesis of the endogenous hs121 enhancer results in abnormalities in the number of vasointestinal peptide (VIP+) interneurons in adult mice brains only for the allele that abolished hs121 enhancer activity in transgenic reporter assay.
Coronal cross-sections through cortex from two knock-in lines generated for mutant alleles of the hs121 enhancer: one that was active (a) in the transgenic reporter assay and one that was inactive (c). Representative images shown from all phenotyped animals: three wild-type littermate male controls and four hemizygous knock-in males for each allele, displayed side-by-side for comparison. Numbers represent unique identifiers for each mouse examined (details in Supplementary Table 7). Scale bars, 1 mm. b, d, Bar graphs showing quantification of VIP+ interneuron densities (cells/mm2) between wild-type (n = 3) and knock-in (n = 4) mice. Bars indicate mean densities, with individual biological replicates represented by dots. Error bars, mean ± s.d. Differences between wild-type and knock-in mice assessed by two-tailed paired t-tests.
Supplementary information
Supplementary Information
Supplementary Tables 3, 4 and 7 and Supplementary Figs. 1 and 2.
Supplementary Tables
Supplementary Table 1. Loss-of-function results from ultraconserved enhancer mutagenesis. P values and FASTA sequences of reference and mutated alleles. Supplementary Table 2. Gain-of-function results from ultraconserved enhancer mutagenesis. P values and FASTA sequences of reference and mutated alleles. Supplementary Table 5. Primers used to amplify enhancers to make transgenic enhancer–reporter mice. Supplementary Table 6. Primers used to generate and characterize hs122 and hs121 enhancer knock-in mice.
Rights and permissions
About this article
Cite this article
Snetkova, V., Ypsilanti, A.R., Akiyama, J.A. et al. Ultraconserved enhancer function does not require perfect sequence conservation. Nat Genet 53, 521–528 (2021). https://doi.org/10.1038/s41588-021-00812-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-021-00812-3
This article is cited by
-
Genetic effects of sequence-conserved enhancer-like elements on human complex traits
Genome Biology (2024)
-
Cis-regulatory interfaces reveal the molecular mechanisms underlying the notochord gene regulatory network of Ciona
Nature Communications (2024)
-
Multi-omics analysis in human retina uncovers ultraconserved cis-regulatory elements at rare eye disease loci
Nature Communications (2024)
-
STR mutations on chromosome 15q cause thyrotropin resistance by activating a primate-specific enhancer of MIR7-2/MIR1179
Nature Genetics (2024)
-
Decoding enhancer complexity with machine learning and high-throughput discovery
Genome Biology (2023)