Abstract
Animal chromosomes are partitioned into contact domains. Pathogenic domain disruptions can result from chromosomal rearrangements or perturbation of architectural factors. However, such broad-scale alterations are insufficient to define the minimal requirements for domain formation. Moreover, to what extent domains can be engineered is just beginning to be explored. In an attempt to create contact domains, we inserted a 2-kb DNA sequence underlying a tissue-invariant domain boundary—containing a CTCF-binding site (CBS) and a transcription start site (TSS)—into 16 ectopic loci across 11 chromosomes, and characterized its architectural impact. Depending on local constraints, this fragment variably formed new domains, partitioned existing ones, altered compartmentalization and initiated contacts reflecting chromatin loop extrusion. Deletions of the CBS or the TSS individually or in combination within inserts revealed its distinct contributions to genome folding. Altogether, short DNA insertions can suffice to shape the spatial genome in a manner influenced by chromatin context.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All main, extended data and supplementary figures include publicly available data. All Hi-C, Capture-C, RNA-seq, ChIP–seq, and other applicable next-generation sequencing raw data and processed data generated from the present study are available under accession no. GSE137376 (GEO database). Mouse CTCF ChIP–seq and mouse Hi-C domain boundaries (both asynchronous) shown in Fig. 6a–c are derived from Zhang et al.19 (https://doi.org/10.1038/s41586-019-1778-y), accession no. GSE129997 (GEO database). In Supplementary Fig. 1: Hi-C heatmaps from all cell lines, except for HAP1, are from GEO, accession no. GSE63525 by Rao et al.4 (https://doi.org/10.1016/j.cell.2014.11.021); K562 ChIP–seq data are from ENCODE, CTCF (DCC accession no. ENCSR000AKO), SMC3 (DCC accession no. ENCSR000EGW), RAD21 (DCC accession no. ENCSR000FAD) and Pol2 (DCC accession no. ENCSR000FAY). Source data are provided with this paper.
Code availability
Code used in the present study is available upon request as well as on GitHub (https://github.com/dizhmp/boundary-insertion).
References
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Rao, S. S. P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Phillips-Cremins, J. E. et al. Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell 153, 1281–1295 (2013).
Schwarzer, W. et al. Two independent modes of chromatin organization revealed by cohesin removal. Nature 551, 51–56 (2017).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320.e24 (2017).
Rowley, M. J. et al. Evolutionarily conserved principles predict 3D chromatin organization. Mol. Cell 67, 837–852.e7 (2017).
Nora, E. P. et al. Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169, 930–944.e22 (2017).
Hug, C. B., Grimaldi, A. G., Kruse, K. & Vaquerizas, J. M. Chromatin architecture emerges during zygotic genome activation independent of transcription. Cell 169, 216–228.e19 (2017).
Franke, M. et al. Formation of new chromatin domains determines pathogenicity of genomic duplications. Nature 538, 265–269 (2016).
Vietri Rudan, M. et al. Comparative Hi-C reveals that CTCF underlies evolution of chromosomal domain architecture. Cell Rep. 10, 1297–1309 (2015).
Fudenberg, G. & Pollard, K. S. Chromatin features constrain structural variation across evolutionary timescales. Proc. Natl Acad. Sci. USA 116, 2175–2180 (2019).
Symmons, O. et al. The shh topological domain facilitates the action of remote enhancers by reducing the effects of genomic distances. Dev. Cell 39, 529–543 (2016).
Lupiáñez, D. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015).
Narendra, V. et al. CTCF establishes discrete functional chromatin domains at the Hox clusters during differentiation. Science 347, 1017–1021 (2015).
Flavahan, W. A. et al. Insulator dysfunction and oncogene activation in IDH mutant gliomas. Nature 529, 110–114 (2016).
Hnisz, D. et al. Activation of proto-oncogenes by disruption of chromosome neighborhoods. Science 351, 1454–1458 (2016).
Zhang, Y. et al. Transcriptionally active HERV-H retrotransposons demarcate topologically associating domains in human pluripotent stem cells. Nat. Genet. 51, 1380–1388 (2019).
Barutcu, A. R., Maass, P. G., Lewandowski, J. P., Weiner, C. L. & Rinn, J. L. A TAD boundary is preserved upon deletion of the CTCF-rich Firre locus. Nat. Commun. 9, 1444 (2018).
Mátés, L. et al. Molecular evolution of a novel hyperactive Sleeping Beauty transposase enables robust stable gene transfer in vertebrates. Nat. Genet. 41, 753–761 (2009).
Carette, J. E. et al. Ebola virus entry requires the cholesterol transporter Niemann–Pick C1. Nature 477, 340–343 (2011).
Haarhuis, J. H. I. et al. The cohesin release factor WAPL restricts chromatin loop extension. Cell 169, 693–707.e14 (2017).
Van Bortle, K. et al. Insulator function and topological domain border strength scale with architectural protein occupancy. Genome Biol. 15, R82 (2014).
Mayer, A. et al. Native elongating transcript sequencing reveals human transcriptional activity at nucleotide resolution. Cell 161, 541–554 (2015).
Vian, L. et al. The energetics and physiological impact of cohesin extrusion. Cell 173, 1165–1178.e20 (2018).
Redolfi, J. et al. DamC reveals principles of chromatin folding in vivo without crosslinking and ligation. Nat. Struct. Mol. Biol. 26, 471–480 (2019).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, 6456 (2015).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Dixon, J. R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015).
Krijger, P. H. L. et al. Cell-of-origin-specific 3D genome structure acquired during somatic cell reprogramming. Cell Stem Cell 18, 597–610 (2016).
Ke, Y. et al. 3D chromatin structures of mature gametes and structural reprogramming during mammalian embryogenesis. Cell 170, 367–381.e20 (2017).
Du, Z. et al. Allelic reprogramming of 3D chromatin architecture during early mammalian development. Nature 547, 232–235 (2017).
Heinz, S. et al. Transcription elongation can affect genome 3D structure. Cell 174, 1522–1536.e22 (2018).
Gong, Y. et al. Stratification of TAD boundaries reveals preferential insulation of super-enhancers by strong boundaries. Nat. Commun. 9, 542 (2018).
Hughes, J. R. et al. Analysis of hundreds of cis-regulatory landscapes at high resolution in a single, high-throughput experiment. Nat. Genet. 46, 205–212 (2014).
Nuebler, J., Fudenberg, G., Imakaev, M., Abdennur, N. & Mirny, L. A. Chromatin organization by an interplay of loop extrusion and compartmental segregation. Proc. Natl Acad. Sci. USA 115, E6697–E6706 (2018).
Sun, L. et al. Mixed lineage kinase domain-like protein mediates necrosis signaling downstream of RIP3 kinase. Cell 148, 213–227 (2012).
Zhao, J. et al. Mixed lineage kinase domain-like is a key receptor interacting protein 3 downstream component of TNF-induced necrosis. Proc. Natl Acad. Sci. USA 109, 5322–5327 (2012).
Galluzzi, L., Buqué, A., Kepp, O., Zitvogel, L. & Kroemer, G. Immunogenic cell death in cancer and infectious disease. Nat. Rev. Immunol. 17, 97–111 (2017).
Shan, B., Pan, H., Najafov, A. & Yuan, J. Necroptosis in development and diseases. Genes Dev. 32, 327–340 (2018).
Yuan, J., Amin, P. & Ofengeim, D. Necroptosis and RIPK1-mediated neuroinflammation in CNS diseases. Nat. Rev. Neurosci. 20, 19–33 (2019).
Chung, C. C. et al. Meta-analysis identifies four new loci associated with testicular germ cell tumor. Nat. Genet. 45, 680–685 (2013).
Astle, W. J. et al. The allelic landscape of human blood cell trait variation and links to common complex disease. Cell 167, 1415–1429.e19 (2016).
Mitchell, J. S. et al. Genome-wide association study identifies multiple susceptibility loci for multiple myeloma. Nat. Commun. 7, 12050 (2016).
Hou, C., Zhao, H., Tanimoto, K. & Dean, A. CTCF-dependent enhancer-blocking by alternative chromatin loop formation. Proc. Natl Acad. Sci. USA 105, 20398–20403 (2008).
Rawat, P., Jalan, M., Sadhu, A., Kanaujia, A. & Srivastava, M. Chromatin domain organization of the TCRb locus and its perturbation by ectopic CTCF binding. Mol. Cell Biol. 37, e00557–16 (2017).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Busslinger, G. A. et al. Cohesin is positioned in mammalian genomes by transcription, CTCF and Wapl. Nature 544, 503–507 (2017).
Despang, A. et al. Functional dissection of the Sox9—Kcnj2 locus identifies nonessential and instructive roles of TAD architecture. Nat. Genet 51, 1263–1271 (2019).
Choudhary, M. N. et al. Co-opted transposons help perpetuate conserved higher-order chromosomal structures. Genome Biol. 21, 16 (2020).
Karijolich, J., Zhao, Y., Alla, R. & Glaunsinger, B. Genome-wide mapping of infection-induced SINE RNAs reveals a role in selective mRNA export. Nucleic Acids Res. 45, 6194–6208 (2017).
Zhang, H. et al. Chromatin structure dynamics during the mitosis-to-G1 phase transition. Nature 576, 158–162 (2019).
Sundaram, V. et al. Widespread contribution of transposable elements to the innovation of gene regulatory networks. Genome Res. 24, 1963–1976 (2014).
Schmidt, D. et al. Waves of retrotransposon expansion remodel genome organization and CTCF binding in multiple mammalian lineages. Cell 148, 335–348 (2012).
Bourque, G. et al. Evolution of the mammalian transcription factor binding repertoire via transposable elements. Genome Res. 18, 1752–1762 (2008).
Thybert, D. et al. Repeat associated mechanisms of genome evolution and function revealed by the Mus caroli and Mus pahari genomes. Genome Res. 28, 448–459 (2018).
Jin, F. et al. A high-resolution map of the three-dimensional chromatin interactome in human cells. Nature 503, 290–294 (2013).
Zhang, Y. et al. Chromatin connectivity maps reveal dynamic promoter-enhancer long-range associations. Nature 504, 306–310 (2013).
Kentepozidou, E. et al. Clustered CTCF binding is an evolutionary mechanism to maintain topologically associating domains. Genome Biol. 21, 5 (2020).
Rowley, M. J. & Corces, V. G. Organizational principles of 3D genome architecture. Nat. Rev. Genet. 19, 789–800 (2018).
Zhan, Y. et al. Reciprocal insulation analysis of Hi-C data shows that TADs represent a functionally but not structurally privileged scale in the hierarchical folding of chromosomes. Genome Res. 27, 479–490 (2017).
Hsieh, T. S. et al. Resolving the 3D landscape of transcription-linked mammalian chromatin folding. Mol. Cell 78, 539–553.e8 (2020).
Krietenstein, N. et al. Ultrastructural details of mammalian chromosome architecture. Mol. Cell 78, 554–565.e7 (2020).
Kurita, R. et al. Establishment of immortalized human erythroid progenitor cell lines able to produce enucleated red blood cells. PLoS ONE 8, e59890 (2013).
Zayed, H., Izsvák, Z., Walisko, O. & Ivics, Z. Development of hyperactive sleeping beauty transposon vectors by mutational analysis. Mol. Ther. 9, 292–304 (2004).
Huang, P. et al. Comparative analysis of three-dimensional chromosomal architecture identifies a novel fetal hemoglobin regulatory element. Genes Dev. 31, 1704–1713 (2017).
Davies, J. O. J. et al. Multiplexed analysis of chromosome conformation at vastly improved sensitivity. Nat. Methods 13, 74–80 (2016).
Hsiung, C. C.- et al. A hyperactive transcriptional state marks genome reactivation at the mitosis-G1 transition. Genes Dev. 30, 1423–1439 (2016).
Hsiau, T. et al. Inference of CRISPR edits from Sanger trace data. Preprint at bioRxiv https://doi.org/10.1101/251082 (2019).
Kim, S., Kim, D., Cho, S. W., Kim, J. & Kim, J. Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res. 24, 1012–1019 (2014).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Sloan, C. A. et al. ENCODE data at the ENCODE portal. Nucleic Acids Res. 44, 726–732 (2016).
Kerpedjiev, P. et al. HiGlass: web-based visual exploration and analysis of genome interaction maps. Genome Biol. 19, 125 (2018).
Forcato, M. et al. Comparison of computational methods for Hi-C data analysis. Nat. Methods 14, 679–685 (2017).
Crane, E. et al. Condensin-driven remodelling of X chromosome topology during dosage compensation. Nature 523, 240–244 (2015).
Filippova, D., Patro, R., Duggal, G. & Kingsford, C. Identification of alternative topological domains in chromatin. Algorithms Mol. Biol. 9, 14 (2014).
Eisenberg, E. & Levanon, E. Y. Human housekeeping genes, revisited. Trends Genet. 29, 569–574 (2013).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization. Nat. Methods 9, 999–1003 (2012).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Gilgenast, T. G. & Phillips-Cremins, J. E. Systematic evaluation of statistical methods for identifying looping interactions in 5C data. Cell Syst. 8, 197–211.e13 (2019).
Hunter, J. D. Matplotlib: a 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Ambrosini, G., Groux, R. & Bucher, P. PWMScan: a fast tool for scanning entire genomes with a position-specific weight matrix. Bioinformatics 34, 2483–2484 (2018).
Khan, A. et al. JASPAR 2018: update of the open-access database of transcription factor binding profiles and its web framework. Nucleic Acids Res. 46, D260–D266 (2018).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98 (2016).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011).
Magoč, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Langmead, B. Aligning short sequencing reads with Bowtie. Curr. Protoc. Bioinform. Chapter 11, Unit 11.7 (2010).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Xu, S., Grullon, S., Ge, K. & Peng, W. Spatial clustering for identification of ChIP-enriched regions (SICER) to map regions of histone methylation patterns in embryonic stem cells. Methods Mol. Biol. 1150, 97–111 (2014).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Ross-Innes, C. S. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon: fast and bias-aware quantification of transcript expression using dual-phase inference. Nat. Methods 14, 417–419 (2017).
Soneson, C., Love, M. I. & Robinson, M. D. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Research 4, 1521 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Weiss, M. J., Yu, C. & Orkin, S. H. Erythroid-cell-specific properties of transcription factor GATA-1 revealed by phenotypic rescue of a gene-targeted cell line. Mol. Cell. Biol. 17, 1642–1651 (1997).
Norton, H. K. et al. Detecting hierarchical genome folding with network modularity. Nat. Methods 15, 119–122 (2018).
Acknowledgements
We thank B. van Steensel (Netherlands Cancer Institute) for providing HAP1 cells; Z. Izsvák (Max Delbrück Center) and Z. Ivics (The Paul Ehrlich Institute) for providing the Sleeping Beauty transposon constructs; A. Raj, O. Symmons and F. Yue for helpful comments on the manuscript. We thank the Flow Cytometry Core at the Children’s Hospital of Philadelphia; J. Yano and P. Evans for assistance; and members of the Blobel laboratory for helpful discussions. This work was supported by grants (nos. R01DK054937 and U01HL129998A to G.A.B and R24DK106766 to R.C.H. and G.A.B.). This work was also supported by the Spatial and Functional Genomics program at the Children’s Hospital of Philadelphia.
Author information
Authors and Affiliations
Contributions
D.Z. and G.A.B. conceived the study and designed the experiments. D.Z. performed a large majority of the experiments, analyzed all datasets and interpreted the results. P.H. conducted Hi-C and Capture-C for half the replicates for transposon-edited and control cell lines, and helped with Hi-C and Capture-C analysis and interpretation. M.S. helped generate and characterize cell lines derived from CRISPR targeting the TSSs and the 2-kb elements. C.A.K., B.G. and R.C.H. prepared ChIP–seq and RNA-seq libraries, performed all sequencing, uploaded sequencing data, and conducted RNA-seq alignment and ChIP–seq peak calling. H.Z. generated mouse ChIP and Hi-C datasets used for recent mouse genome evolution analysis. T.G.G. and J.E.P.-C. helped with Hi-C data visualization and interpretation. D.Z. and G.A.B wrote the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
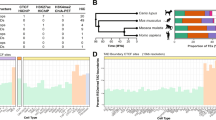
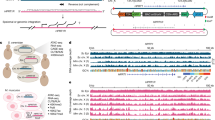
Extended Data Fig. 1 Generation and characterization of transposon genome-edited clones with multiple insertions.
a, Estimated insertion copy numbers using qPCR (see Methods) after transposon insertion in pooled cells and in single-cell-derived clones (numbered). N = 1 qPCR measurement. b, Insertion site mapping: fragmented gDNA containing insertions are captured by biotinylated oligos capturing the inversed repeats (green rectangles), which flank the 2 kb element (orange rectangles). Junction reads are mapped to identify insertion sites. c, Junction read coverage for Clone 21: horizontal axis denotes genomic coordinates (single nucleotide resolution) with > 25X coverage; vertical axis shows read coverage. The spike in the middle of each peak consists of two neighboring nucleotides between which an insertion is located. Data from N = 1 experiment. d, The locations and orientations of Clone 21 insertion sites. The CBS and TSS are in cis (Fig. 1a). “→” denotes that the CBS is on the plus strand and that the TSS transcribes from left to right, and vice versa for “←”. Each insertion site orientation was confirmed in (g). e, Junction read coverage for Clone 25, similar to (c). Data is from N = 1 experiment. f, The locations and orientations of Clone 25 insertion sites, similar to (d). g, Insertion-driven transcription in both directions/strands measured by quantitative PCR with reverse transcription (RT-qPCR). Transcript levels were normalized relative to the geometric mean of the Ct values of 11 housekeeping genes. N = 2 independent experiments for each genotype.
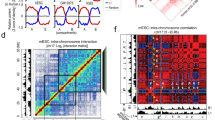
Extended Data Fig. 2 Insertion-driven new domains: detailed comparisons (an extension to Fig. 1).
Throughout, red arrow: insertion site; green arrow: upstream or downstream CBSs; blue/purple arrow: nearby boundaries; orange arrowhead in the browser tracks: site and orientation of the insertion. Green lines demarcate new domains. Yellow/green rectangles (squares) indicate regions with overall depleted (enriched) contacts upon insertion. (a, b): related to Fig. 1b, c. a, An extension to Fig. 1b showing Hi-C maps for both no-insertion controls (left and middle) and the insertion clone (right) at C21S4, each accompanied by corresponding data tracks. b, Log2 fold changes in interaction frequencies between two no-insertion controls (left), and between the insertion clone and no-insertion controls (middle and right) for the region in (a). Yellow/green rectangles: depleted interactions upon insertion; yellow/green squares: increased interactions between two B-compartment domains partitioned by the new domain with A compartment signature. (c, d): related to Fig. 1d, e. c, An extension to Fig. 1d showing both no-insertion controls at C21S2. d, Log2 fold changes in interaction frequencies between no-insertion controls and between insertion and no-insertion controls for the region in (c). (e, f): related to Fig. 1f, g. e, An extension to Fig. 1f showing both no-insertion controls at C21S5. f, Log2 fold changes between no-insertion controls and between insertion and no-insertion controls for the region in (e). Each Hi-C heatmap presents merged data from 2 independent experiments for each genotype. 2 CTCF & RAD21 ChIP-seq and 2 RNA-seq experiments were performed for each genotype, with 1 of each displayed.
Extended Data Fig. 3 Additional insertion loci with possible domain-level changes.
Throughout, red arrow: insertion site; green or blue arrow: nearby boundaries; orange/blue arrowhead in the browser tracks: site and orientation of insertion. Green lines demarcate (possible) new domains. Yellow/green rectangles indicate regions with overall depleted contacts upon insertion. a, De novo domain upon insertion at C21S1: Hi-C maps for both no-insertion controls (left and middle) and the insertion clone (right) at C21S1, each accompanied by corresponding data tracks. b, Insulation scores for the region in (a). c, Log2 fold changes in interaction frequencies between the two no-insertion controls (left) and between the insertion clone and no-insertion controls (middle and right) for the region in (a). d, A small subtle domain forms upon insertion at C25S3 locus. e, Insulation scores for the region in (d). f, Log2 fold changes in interaction frequencies for the region in (d). g, Modest strengthening of an existing boundary upon insertion at C25S4. h, Insulation scores for the region in (g). i, Log2 fold changes for the region in (g). j, Subtle strengthening of an existing boundary upon insertion at C25S1. The black arrowheads point at insertion-associated changes. k, Insulation scores for the region in (j). l, Log2 fold changes for the region in (j). Each Hi-C heatmap presents merged data from 2 independent experiments for each genotype. 2 CTCF & RAD21 ChIP-seq and 2 RNA-seq experiments were performed for each genotype, with 1 of each displayed.
Extended Data Fig. 4 An ectopic insertion can redirect its local chromatin from B to A compartment.
Throughout, left: compartment eigenvectors (cyan denotes B compartment; red denotes A compartment) for the ~14 Mb region marked by the green rectangle on the chromosome diagram; middle: Hi-C heatmaps for this ~14 Mb region surrounding C21S4; right: distal interactions between this ~14 Mb region and a ~40 Mb region downstream marked by the purple rectangle. Black arrows: compartment switch; orange arrowhead: location of the insertion; black arrowhead: corresponding locations in no-insertion controls. a, No-insertion control 1 (WT) at C21S4. b, No-insertion control 2 (Clone 25) at C21S4. c, Insertion clone (Clone 21) at C21S4: compartment eigenvectors demonstrate the insertion locus trending from a strong B compartment towards A as the largest change in the region. The Hi-C heatmap for the ~14 Mb with the insertion at the center shows a plaid like pattern, with gained interactions between the insertion locus and its nearby A compartment regions. Distal interactions (right) shows the insertion locus forming distal interactions with other A-compartment regions (black arrows), which are absent in (a, b). Each Hi-C result depicts merged data from 2 independent Hi-C experiments for each genotype.
Extended Data Fig. 5 Boundary-associated DNA insertions can strengthen pre-established boundaries: additional controls (an extension to Fig. 2).
Throughout, red arrow: insertion site; green or blue arrow: nearby boundaries; Blue/orange arrowhead in the browser tracks: site and orientation of the insertion. Yellow/green rectangles indicate regions with overall depleted contacts upon insertion. (a, b) are related to Fig. 2a–c. a, An extension to Fig. 2a showing both no-insertion controls (left and middle) and the insertion clone (right) at C25S5, each accompanied by corresponding data tracks. b, An extension to Fig. 2c: log2 fold changes in interaction frequencies between two no-insertion controls (left) and between the insertion clone and no-insertion controls (middle and right) for the region in (a). (c, d) are related to Fig. 2d–f. c, An extension to Fig. 2d showing both no-insertion controls at C21S7. d, An extension to Fig. 2f: log2 fold changes in interaction frequencies between two no-insertion controls and between the insertion clone and no-insertion controls for the region shown in (d). Each Hi-C heatmap represents merged data from 2 independent experiments for each genotype. 2 CTCF & RAD21 ChIP-seq and 2 RNA-seq experiments were conducted for each genotype, with 1 of each exhibited.
Extended Data Fig. 6 Insertion loci without apparent detectable domain-level changes.
Throughout, red arrow: insertion site; orange/blue arrowhead in the browser tracks: locus/orientation of the insertion. a, An insertion at C21S6: Hi-C maps for both no-insertion controls (left and middle) and the insertion clone (right) at C21S6, each accompanied by corresponding data tracks. b, Insulation scores for the region in (a). c, Log2 fold changes in interaction frequencies between two no-insertion controls (left) and between the insertion clone and no-insertion controls (middle and right) for the region in (a). d, Hi-C contact maps at C21S10. e, Insulation score profiles for the region in (d). f, Log2 fold changes in interaction frequencies between two no-insertion controls and between the insertion clone and no-insertion controls for the region in (d). g, Hi-C contact maps at C25S6. h, Insulation score profiles for the region in (g). i, Log2 fold changes in interaction frequencies for the region shown in (g). Each Hi-C heatmap presents merged data from 2 independent experiments performed for each genotype. 2 CTCF & RAD21 ChIP-seq and 2 RNA-seq experiments were performed for each genotype, with 1 of each displayed.
Extended Data Fig. 7 Transcription of insertion-proximal genes remains mostly stable, with MLKL as an exception.
a, An MA plot showing Clone 21 vs. non-Clone 21 transcriptomes. Each dot: a gene; red dots: differentially expressed (DE) genes at an FDR < 0.01; color-coded circles: insertion-proximal genes by distance ranges; red line: no-change line; two orange lines: +/− 1 log2 fold change. b, Clone 25 vs. non-Clone 25 transcriptomes. c, Clone 21 has ~95 DE genes transcriptome-wide (related to (a)). d, Clone 25 has ~160 DE genes transcriptome-wide (related to (b)). e, DE status of all insertion-proximal genes. The DE gene between 50 kb and 500 kb to an insertion, MLKL, is characterized in (f) and (h). In (a–e), 2 RNA-seq experiments were performed for each genotype. DE analysis was conducted with Clone 21 vs. non-Clone 21 (WT and Clone 25) and Clone 25 vs. non-Clone 25 (WT and Clone 21). f, RT-qPCR of MLKL and GLG1/RFWD3, two genes flanking the insertion (see (h)). N = 2 independent experiments for each genotype. g, GWAS significant variants near GLG1/RFWD3/MLKL insertion locus43,44,45. h, GLG1/RFWD3/MLKL locus (blue arrowhead: location/orientation of the insertion) using ChIP-seq/RNA-seq/Capture-C. The insertion coincides with reduced RAD21 binding at a peak immediately downstream. The insertion contacts the promoter of GLG1 (Capture-C: Probing the insert). MLKL promoter also interacts with GLG1 promoter (Capture-C: Probing MLKL promoter), albeit no apparent changes in interactions of MLKL promoter upon insertion. Capture-C presents merged data from 2 independent experiments for each genotype. 2 CTCF & RAD21 ChIP-seq, 1 H3K27ac ChIP-seq and 2 RNA-seq experiments were conducted for each genotype, with 1 of each shown.
Extended Data Fig. 8 CRISPR dissections of insertion, and CTCF/RAD21 at C21S4.
a, Left: sgRNAs within the insertion element (red lines: Pol2/CTCF peak centers). Right: TSS_sgRNA_2&4 and TSS_sgRNA_3&4 reduce transcription more effectively at C21S4. N = 1. b, CRISPR deletion of the inserted CBS spares transcription. c, Clone 21 ΔTSS: TSS_sgRNA_2&4-edited Clone 21 abrogates transcription, with the CBS intact. d, e, Clone 21 ΔCTCF/ΔTSS #1: Clone 21 with its CBS already disrupted (b) further edited with TSS_sgRNA_2&4 and TSS_sgRNA_3&4, respectively. In (b–e), N = 2 experiments for each genotype. In (f, g and i), red arrow: insertion site; green arrow: downstream CBSs; blue/purple arrow: strong boundary nearby; orange arrowhead: insertion location/orientation. f, Hi-C of Clone 21 ΔCTCF/ΔTSS #1 (d) at C21S4: a ~27 Mb heterozygous deletion (h) influences heatmap interpretation. g, Hi-C of Clone 21 ΔCTCF/ΔTSS #2 (e) at C21S4: domain configuration restored close to pre-insertion level (Fig. 4a). h, Virtual 4 C (black arrow: viewpoint; red star: C21S4; GRIK2: C21S5): Clone 21 ΔCTCF/ΔTSS #1 has both short-range contacts and strong >25-Mb distal contacts, suggesting a heterozygous deletion between C21S4 and C21S5 (grey bars: chromosomes; dashed line: deletion). i, ΔCBS/ΔTSS restores nearby chromatin folding pattern to pre-insertion levels. Differentially bound CTCF (C2, C4) and RAD21 peaks (R1-R5) upon insertion highlighted. Directionality Index of Clone 21 ΔCTCF/ΔTSS #1 Capture-C: Fig. 4j. In (f–i), each Hi-C/Capture-C describes merged data from at least 2 independent experiments for each genotype. 2 CTCF/RAD21 ChIP-seq and 1 H3K27ac ChIP-seq for each genotype, with 1 of each shown. j, Pairwise comparisons between genotypes of CTCF binding (C2 and C4: (i) and Fig. 4f–i). k, Pairwise comparisons between genotypes of RAD21 binding (R1-R5: (i) and Fig. 4f–i). In (j, k), non-Clone 21: 3 genotypes without Clone21 insertions, each with 2 ChIP-seq replicates. Clone21 CTCF/TSS and derived CRISPR clones: 1 genotype, each with 2 ChIP-seq replicates. P-values (not adjusted for multiple comparisons): from a two-sided Wald test.
Extended Data Fig. 9 CRISPR dissections of insertion, and RAD21 distribution at C21S2.
a, TSS_sgRNA_2&4 and TSS_sgRNA_3&4 (as in Extended Data Fig. 8a) reduce transcription more effectively at C21S2 in CRISPR-Cas9 RNP-transfected cells. N = 1 experiment. b, Deletion of the inserted CBS reduces but does not abolish transcription at C21S2. c, Clone 21 ΔTSS derived from TSS_sgRNA_2&4-edited Clone 21 abrogates transcription, with the CBS intact. d, Clone 21 ΔCTCF/ΔTSS #1: derived from CBS-disrupted Clone 21 (b) further edited with TSS_sgRNA_2&4. e, Clone 21 ΔCTCF/ΔTSS #2: derived from CBS-disrupted Clone 21 (b) further edited with TSS_sgRNA_3&4. In (b–e), N = 2 independent experiments for each genotype. In (f–h), red arrow: insertion site; green or blue arrow: downstream CBSs; orange arrowhead in the browser tracks: locus/orientation of the insertion. f, g, Hi-C maps of Clone 21 ΔCTCF/ΔTSS #1 (d) and of Clone 21 ΔCTCF/ΔTSS #2 (e), respectively, at C21S2: deletions of both the CBS and the TSS restore the domain configuration close to pre-insertion level (Fig. 5a). h, Capture-C and corresponding data tracks showing that ΔCTCF/ΔTSS rescues local chromatin contact pattern close to that of WT. Differentially bound RAD21 peaks (R6, R7) upon CBS-TSS insertion highlighted. Directionality Index of Clone 21 ΔCTCF/ΔTSS #1 Capture-C: Fig. 5j. In (f–h), each Hi-C/Capture-C depicts merged data from at least 2 independent experiments for each genotype. 2 CTCF/RAD21 ChIP-seq and 1 H3K27ac ChIP-seq for each genotype, with 1 of each shown. i, Pairwise comparisons between genotypes of RAD21 binding at two RAD21 peaks (R6 and R7, as in (h) and Fig. 5f–i). Non-Clone 21: 3 genotypes without Clone 21 insertions, each with 2 ChIP-seq replicates. All others: 1 genotype, each with 2 ChIP-seq replicates. P-values (not adjusted for multiple comparisons) are derived from a two-sided Wald test through DiffBind.
Extended Data Fig. 10 Deletion of the endogenous 2 kb element leads to a boundary shift, while local domain organization is stable.
a, Hi-C of no-deletion control showing the endogenous boundary where the 2 kb element (blue arrowhead) is derived, accompanied by corresponding data tracks. b, Deletion of the 2 kb (crossed-out blue arrowhead) leaves the overall domain configuration largely intact. The highlighted ~400 kb region is further examined in (c) and (f). c, Insulation scores show overall concordance, with a possible shift in boundary by ~60 kb to the left upon deletion. d, Genotyping confirms the desired deletion between sgRNAs flanking the 2 kb. e, ChIP-seq further verifies the deletion, as reflected in lack of signal (black arrows) within the 2 kb element (highlighted). f, Upon 2 kb deletion (highlighted in red), the point of local maximal insulation shifts ~60 kb to the left (c), coinciding with the distance between the TSSs of PARL and its nearest transcribed gene: MAP6D1 (highlighted in yellow). This shift (red line) also corresponds to the distance between the deleted CBS and its nearest CTCF peak to the left, which now has reduced CTCF/RAD21 binding. Each Hi-C result presents merged data from 2 independent experiments for each genotype. 2 CTCF & RAD21 ChIP-seq experiments for each genotype, with 1 of each shown.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2
Supplementary Tables
Supplementary Tables 1–4
Supplementary Data 1
Sequence map of the Sleeping Beauty transposon with the 2-kb insert.
Source data
Source Data Fig. 6
Statistical source data for Fig. 6c.
Source Data Extended Data Fig. 7
Statistical source data for Extended Data Fig. 7e.
Source Data Extended Data Fig. 8
Statistical source data for Extended Data Fig. 8j and k.
Source Data Extended Data Fig. 9
Statistical source data for Extended Data 9i.
Source Data Extended Data Fig. 10
Uncropped gel for Extended Data Fig. 10d.
Rights and permissions
About this article
Cite this article
Zhang, D., Huang, P., Sharma, M. et al. Alteration of genome folding via contact domain boundary insertion. Nat Genet 52, 1076–1087 (2020). https://doi.org/10.1038/s41588-020-0680-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-0680-8
This article is cited by
-
Capture-C: a modular and flexible approach for high-resolution chromosome conformation capture
Nature Protocols (2022)
-
Sequence-based modeling of three-dimensional genome architecture from kilobase to chromosome scale
Nature Genetics (2022)
-
A chromosomal loop anchor mediates bacterial genome organization
Nature Genetics (2022)
-
Cohesin-mediated loop anchors confine the locations of human replication origins
Nature (2022)
-
CTCF and transcription influence chromatin structure re-configuration after mitosis
Nature Communications (2021)