Abstract
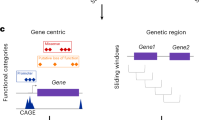
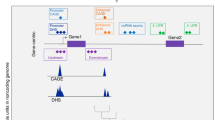
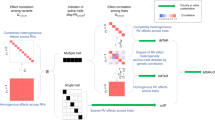
Large-scale whole-genome sequencing studies have enabled the analysis of rare variants (RVs) associated with complex phenotypes. Commonly used RV association tests have limited scope to leverage variant functions. We propose STAAR (variant-set test for association using annotation information), a scalable and powerful RV association test method that effectively incorporates both variant categories and multiple complementary annotations using a dynamic weighting scheme. For the latter, we introduce ‘annotation principal components’, multidimensional summaries of in silico variant annotations. STAAR accounts for population structure and relatedness and is scalable for analyzing very large cohort and biobank whole-genome sequencing studies of continuous and dichotomous traits. We applied STAAR to identify RVs associated with four lipid traits in 12,316 discovery and 17,822 replication samples from the Trans-Omics for Precision Medicine Program. We discovered and replicated new RV associations, including disruptive missense RVs of NPC1L1 and an intergenic region near APOC1P1 associated with low-density lipoprotein cholesterol.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
This paper used the TOPMed Freeze 5 WGS data and lipids phenotype data. Genotype and phenotype data are both available in database of Genotypes and Phenotypes. The discovery phase used data from the following four study cohorts (accession numbers provided in parentheses): Framingham Heart Study (phs000974.v1.p1); Genetics of Cardiometabolic Health in the Amish (phs000956.v1.p1); Jackson Heart Study (phs000964.v1.p1); and Multi-Ethnic Study of Atherosclerosis (phs001416.v1.p1). The replication phase used data from the following ten study cohorts: Atherosclerosis Risk in Communities Study (phs001211); Cleveland Family Study (phs000954); Cardiovascular Health Study (phs001368); Diabetes Heart Study (phs001412); Genetic Study of Atherosclerosis Risk (phs001218); Genetic Epidemiology Network of Arteriopathy (phs001345); Genetics of Lipid Lowering Drugs and Diet Network (phs001359); San Antonio Family Heart Study (phs001215); Genome-Wide Association Study of Adiposity in Samoans (phs000972); and Women’s Health Initiative (phs001237). The sample sizes, ancestry and phenotype summary statistics of these cohorts are given in Supplementary Table 3.
The functional annotation data are publicly available and were downloaded from the following links: GRCh38 CADD v.1.4 (https://cadd.gs.washington.edu/download); ANNOVAR dbNSFP v.3.3a (https://annovar.openbioinformatics.org/en/latest/user-guide/download); LINSIGHT (https://github.com/CshlSiepelLab/LINSIGHT); FATHMM-XF (http://fathmm.biocompute.org.uk/fathmm-xf); FANTOM5 CAGE (https://fantom.gsc.riken.jp/5/data); GeneCards (https://www.genecards.org; v.4.7 for hg38); and Umap/Bismap (https://bismap.hoffmanlab.org; ‘before March 2020’ version). In addition, recombination rate and nucleotide diversity were obtained from Gazal et al90. The whole-genome individual functional annotation data assembled from a variety of sources and the computed annotation principal components are available at the Functional Annotation of Variant–Online Resource (FAVOR) site (http://favor.genohub.org). The tissue-specific functional annotations were downloaded from ENCODE (https://www.encodeproject.org/report/?type=Experiment).
Code availability
STAAR is implemented as an open source R package available at https://github.com/xihaoli/STAAR and https://content.sph.harvard.edu/xlin/software.html.
References
Bansal, V., Libiger, O., Torkamani, A. & Schork, N. J. Statistical analysis strategies for association studies involving rare variants. Nat. Rev. Genet. 11, 773–785 (2010).
Kiezun, A. et al. Exome sequencing and the genetic basis of complex traits. Nat. Genet. 44, 623–630 (2012).
Lee, S., Abecasis, G. R., Boehnke, M. & Lin, X. Rare-variant association analysis: study designs and statistical tests. Am. J. Hum. Genet. 95, 5–23 (2014).
Morgenthaler, S. & Thilly, W. G. A strategy to discover genes that carry multi-allelic or mono-allelic risk for common diseases: a cohort allelic sums test (CAST). Mutat. Res. 615, 28–56 (2007).
Li, B. & Leal, S. M. Methods for detecting associations with rare variants for common diseases: application to analysis of sequence data. Am. J. Hum. Genet. 83, 311–321 (2008).
Madsen, B. E. & Browning, S. R. A groupwise association test for rare mutations using a weighted sum statistic. PLoS Genet. 5, e1000384 (2009).
Morris, A. P. & Zeggini, E. An evaluation of statistical approaches to rare variant analysis in genetic association studies. Genet. Epidemiol. 34, 188–193 (2010).
Wu, M. C. et al. Rare-variant association testing for sequencing data with the sequence kernel association test. Am. J. Hum. Genet. 89, 82–93 (2011).
Liu, Y. et al. ACAT: a fast and powerful p value combination method for rare-variant analysis in sequencing studies. Am. J. Hum. Genet. 104, 410–421 (2019).
Lee, S., Wu, M. C. & Lin, X. Optimal tests for rare variant effects in sequencing association studies. Biostatistics 13, 762–775 (2012).
Sun, J., Zheng, Y. & Hsu, L. A unified mixed-effects model for rare-variant association in sequencing studies. Genet. Epidemiol. 37, 334–344 (2013).
Pan, W., Kim, J., Zhang, Y., Shen, X. & Wei, P. A powerful and adaptive association test for rare variants. Genetics 197, 1081–1095 (2014).
Kichaev, G. et al. Integrating functional data to prioritize causal variants in statistical fine-mapping studies. PLoS Genet. 10, e1004722 (2014).
Kichaev, G. et al. Improved methods for multi-trait fine mapping of pleiotropic risk loci. Bioinformatics 33, 248–255 (2017).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Hu, Y. et al. Leveraging functional annotations in genetic risk prediction for human complex diseases. PLoS Comp. Biol. 13, e1005589 (2017).
Morrison, A. C. et al. Practical approaches for whole-genome sequence analysis of heart-and blood-related traits. Am. J. Hum. Genet. 100, 205–215 (2017).
Schaid, D. J., Chen, W. & Larson, N. B. From genome-wide associations to candidate causal variants by statistical fine-mapping. Nat. Rev. Genet. 19, 491–504 (2018).
Claussnitzer, M. et al. A brief history of human disease genetics. Nature 577, 179–189 (2020).
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 22, 1760–1774 (2012).
Frankish, A. et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 47, D766–D773 (2019).
Ng, P. C. & Henikoff, S. SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res. 31, 3812–3814 (2003).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005).
Pollard, K. S., Hubisz, M. J., Rosenbloom, K. R. & Siepel, A. Detection of nonneutral substitution rates on mammalian phylogenies. Genome Res. 20, 110–121 (2010).
Dunham, I. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Tang, H. & Thomas, P. D. Tools for predicting the functional impact of nonsynonymous genetic variation. Genetics 203, 635–647 (2016).
Lee, P. H. et al. Principles and methods of in-silico prioritization of non-coding regulatory variants. Hum. Genet. 137, 15–30 (2018).
Kellis, M. et al. Defining functional DNA elements in the human genome. Proc. Natl Acad. Sci. USA 111, 6131–6138 (2014).
Zuk, O. et al. Searching for missing heritability: designing rare variant association studies. Proc. Natl Acad. Sci. USA 111, E455–E464 (2014).
Hao, X., Zeng, P., Zhang, S. & Zhou, X. Identifying and exploiting trait-relevant tissues with multiple functional annotations in genome-wide association studies. PLoS Genet. 14, e1007186 (2018).
He, Z., Xu, B., Lee, S. & Ionita-Laza, I. Unified sequence-based association tests allowing for multiple functional annotations and meta-analysis of noncoding variation in Metabochip data. Am. J. Hum. Genet. 101, 340–352 (2017).
Ma, Y. & Wei, P. FunSPU: a versatile and adaptive multiple functional annotation-based association test of whole-genome sequencing data. PLoS Genet. 15, e1008081 (2019).
Breslow, N. E. & Clayton, D. G. Approximate inference in generalized linear mixed models. J. Am. Stat. Assoc. 88, 9–25 (1993).
Chen, H. et al. Control for population structure and relatedness for binary traits in genetic association studies via logistic mixed models. Am. J. Hum. Genet. 98, 653–666 (2016).
Chen, H. et al. Efficient variant set mixed model association tests for continuous and binary traits in large-scale whole-genome sequencing studies. Am. J. Hum. Genet. 104, 260–274 (2019).
Gogarten, S. M. et al. Genetic association testing using the GENESIS R/Bioconductor package. Bioinformatics 35, 5346–5348 (2019).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Rentzsch, P., Witten, D., Cooper, G. M., Shendure, J. & Kircher, M. CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res. 47, D886–D894 (2019).
Liu, X., Wu, C., Li, C. & Boerwinkle, E. dbNSFP v3.0: a one-stop database of functional predictions and annotations for human nonsynonymous and splice-site SNVs. Hum. Mutat. 37, 235–241 (2016).
Liu, Y. & Xie, J. Cauchy combination test: a powerful test with analytic p-value calculation under arbitrary dependency structures. J. Am. Stat. Assoc. 115, 393–402 (2020).
Schaffner, S. F. et al. Calibrating a coalescent simulation of human genome sequence variation. Genome Res. 15, 1576–1583 (2005).
Natarajan, P. et al. Deep-coverage whole genome sequences and blood lipids among 16,324 individuals. Nat. Commun. 9, 3391 (2018).
Taliun, D. et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Preprint at bioRxiv https://doi.org/10.1101/563866 (2019).
Huang, Y.-F., Gulko, B. & Siepel, A. Fast, scalable prediction of deleterious noncoding variants from functional and population genomic data. Nat. Genet. 49, 618–624 (2017).
Rogers, M. F. et al. FATHMM-XF: accurate prediction of pathogenic point mutations via extended features. Bioinformatics 34, 511–513 (2018).
Forrest, A. R. R. et al. A promoter-level mammalian expression atlas. Nature 507, 462–470 (2014).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Fishilevich, S. et al. GeneHancer: genome-wide integration of enhancers and target genes in GeneCards. Database (Oxford) 2017, bax028 (2017).
Dong, C. et al. Comparison and integration of deleteriousness prediction methods for nonsynonymous SNVs in whole exome sequencing studies. Hum. Mol. Genet. 24, 2125–2137 (2015).
Sabatti, C. et al. Genome-wide association analysis of metabolic traits in a birth cohort from a founder population. Nat. Genet. 41, 35–46 (2009).
Kathiresan, S. et al. Six new loci associated with blood low-density lipoprotein cholesterol, high-density lipoprotein cholesterol or triglycerides in humans. Nat. Genet. 40, 189–197 (2008).
Huang, C.-C. et al. Longitudinal association of PCSK9 sequence variations with low-density lipoprotein cholesterol levels: the Coronary Artery Risk Development in Young Adults Study. Circ. Cardiovasc. Genet. 2, 354–361 (2009).
Lange, L. A. et al. Whole-exome sequencing identifies rare and low-frequency coding variants associated with LDL cholesterol. Am. J. Hum. Genet. 94, 233–245 (2014).
Bomba, L., Walter, K. & Soranzo, N. The impact of rare and low-frequency genetic variants in common disease. Genome Biol. 18, 77 (2017).
Ference, B. A., Majeed, F., Penumetcha, R., Flack, J. M. & Brook, R. D. Effect of naturally random allocation to lower low-density lipoprotein cholesterol on the risk of coronary heart disease mediated by polymorphisms in NPC1L1, HMGCR, or both: a 2 × 2 factorial Mendelian randomization study. J. Am. Coll. Cardiol. 65, 1552–1561 (2015).
Teslovich, T. M. et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature 466, 707–713 (2010).
Surakka, I. et al. The impact of low-frequency and rare variants on lipid levels. Nat. Genet. 47, 589–597 (2015).
Kathiresan, S. et al. Common variants at 30 loci contribute to polygenic dyslipidemia. Nat. Genet. 41, 56–65 (2009).
Kamatani, Y. et al. Genome-wide association study of hematological and biochemical traits in a Japanese population. Nat. Genet. 42, 210–215 (2010).
Nagy, R. et al. Exploration of haplotype research consortium imputation for genome-wide association studies in 20,032 Generation Scotland participants. Genome Med. 9, 23 (2017).
Aulchenko, Y. S. et al. Loci influencing lipid levels and coronary heart disease risk in 16 European population cohorts. Nat. Genet. 41, 47–55 (2009).
Deelen, J. et al. Genome-wide association study identifies a single major locus contributing to survival into old age; the APOE locus revisited. Aging Cell 10, 686–698 (2011).
Klarin, D. et al. Genetics of blood lipids among ~300,000 multi-ethnic participants of the Million Veteran Program. Nat. Genet. 50, 1514–1523 (2018).
Hoffmann, T. J. et al. A large electronic-health-record-based genome-wide study of serum lipids. Nat. Genet. 50, 401–413 (2018).
Willer, C. J. et al. Discovery and refinement of loci associated with lipid levels. Nat. Genet. 45, 1274–1283 (2013).
Cohen, J. C. et al. Multiple rare variants in NPC1L1 associated with reduced sterol absorption and plasma low-density lipoprotein levels. Proc. Natl Acad. Sci. USA 103, 1810–1815 (2006).
Stitziel, N. O. et al. Inactivating mutations in NPC1L1 and protection from coronary heart disease. N. Engl. J. Med. 371, 2072–2082 (2014).
Cooper, G. M. et al. Single-nucleotide evolutionary constraint scores highlight disease-causing mutations. Nat. Methods 7, 250–251 (2010).
Cooper, G. M. & Shendure, J. Needles in stacks of needles: finding disease-causal variants in a wealth of genomic data. Nat. Rev. Genet. 12, 628–640 (2011).
Van Hout, C. V. et al. Whole exome sequencing and characterization of coding variation in 49,960 individuals in the UK Biobank. Preprint at bioRxiv https://doi.org/10.1101/572347 (2019).
Crosby, J. et al. Loss-of-function mutations in APOC3, triglycerides, and coronary disease. N. Engl. J. Med. 371, 22–31 (2014).
Myers, R. M. et al. A user’s guide to the Encyclopedia of DNA Elements (ENCODE). PLoS Biol. 9, e1001046 (2011).
Landrum, M. J. et al. ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res. 46, D1062–D1067 (2018).
Davis, H. R. & Veltri, E. P. Zetia: inhibition of Niemann-Pick C1 Like 1 (NPC1L1) to reduce intestinal cholesterol absorption and treat hyperlipidemia. J. Atheroscler. Thromb. 14, 99–108 (2007).
Klos, K. et al. APOE/C1/C4/C2 hepatic control region polymorphism influences plasma apoE and LDL cholesterol levels. Hum. Mol. Genet. 17, 2039–2046 (2008).
Lu, Q., Powles, R. L., Wang, Q., He, B. J. & Zhao, H. Integrative tissue-specific functional annotations in the human genome provide novel insights on many complex traits and improve signal prioritization in genome wide association studies. PLoS Genet. 12, e1005947 (2016).
Backenroth, D. et al. FUN-LDA: a latent Dirichlet allocation model for predicting tissue-specific functional effects of noncoding variation: methods and applications. Am. J. Hum. Genet. 102, 920–942 (2018).
Bodea, C. A. et al. PINES: phenotype-informed tissue weighting improves prediction of pathogenic noncoding variants. Genome Biol. 19, 173 (2018).
Park, J.-H. et al. Estimation of effect size distribution from genome-wide association studies and implications for future discoveries. Nat. Genet. 42, 570–575 (2010).
Derkach, A., Zhang, H. & Chatterjee, N. Power Analysis for Genetic Association Test (PAGEANT) provides insights to challenges for rare variant association studies. Bioinformatics 34, 1506–1513 (2018).
Li, Z. et al. Dynamic scan procedure for detecting rare-variant association regions in whole-genome sequencing studies. Am. J. Hum. Genet. 104, 802–814 (2019).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Conomos, M. P., Reiner, A. P., Weir, B. S. & Thornton, T. A. Model-free estimation of recent genetic relatedness. Am. J. Hum. Genet. 98, 127–148 (2016).
Dey, R., Schmidt, E. M., Abecasis, G. R. & Lee, S. A fast and accurate algorithm to test for binary phenotypes and its application to PheWAS. Am. J. Hum. Genet. 101, 37–49 (2017).
Zhou, W. et al. Efficiently controlling for case-control imbalance and sample relatedness in large-scale genetic association studies. Nat. Genet. 50, 1335–1341 (2018).
Karimzadeh, M., Ernst, C., Kundaje, A. & Hoffman, M. M. Umap and Bismap: quantifying genome and methylome mappability. Nucleic Acids Res. 46, e120 (2018).
Regier, A. A. et al. Functional equivalence of genome sequencing analysis pipelines enables harmonized variant calling across human genetics projects. Nat. Commun. 9, 4038 (2018).
Gazal, S. et al. Linkage disequilibrium-dependent architecture of human complex traits shows action of negative selection. Nat. Genet. 49, 1421–1427 (2017).
Acknowledgements
This work was supported by grant nos. R35-CA197449, P01-CA134294, U19-CA203654 and R01-HL113338 (to X. Lin), U01-HG009088 (to X. Lin, S.R.S. and B.M.N.), R01-HL142711 (to P.N. and G.M.P.), K01-HL125751 and R03-HL141439 (to G.M.P.), R35-HL135824 (to C.J.W.), 75N92020D00001, HHSN268201500003I, N01-HC-95159, 75N92020D00005, N01-HC-95160, 75N92020D00002, N01-HC-95161, 75N92020D00003, N01-HC-95162, 75N92020D00006, N01-HC-95163, 75N92020D00004, N01-HC-95164, 75N92020D00007, N01-HC-95165, N01-HC-95166, N01-HC-95167, N01-HC-95168, N01-HC-95169, UL1-TR-000040, UL1-TR-001079, UL1-TR-001420, UL1TR001881 and DK063491 (to J.I.R. and X.G.), HHSN268201800002I (to G.R.A.), R35-GM127131 and R01-MH101244 (to S.R.S.), U01-HL72518, HL087698, HL49762, HL59684, HL58625, HL071025, HL112064, NR0224103 and M01-RR000052 (to the Johns Hopkins General Clinical Research Center), R01-HL093093, R01-HL133040 (to D.E.W.), NO1-HC-25195, HHSN268201500001I, 75N92019D00031 and R01-HL092577-06S1 (to R.S.V. and L.A.C.), the Evans Medical Foundation and the Jay and Louis Coffman Endowment from the Department of Medicine, Boston University School of Medicine (to R.S.V.), HHSN268201800001I (to K.M.R., A.T.K., M.P.C. and J.G.B.), U01-HL137162 (to K.M.R. and M.P.C.), R35-HL135818 and R01-HL113338 (to S.R.), R01-HL113323, U01-DK085524, R01-HL045522, R01-MH078143, R01-MH078111 and R01-MH083824 (to J.M.P., M.C.M., J.E.C. and J.B.), R01-HL92301, R01-HL67348, R01-NS058700, R01-AR48797 and R01-AG058921 (to N.D.P. and D.W.B.), R01-DK071891 (to N.D.P., B.I.F. and D.W.B.), M01-RR07122 and F32-HL085989 (to the General Clinical Research Center of the Wake Forest University School of Medicine), the American Diabetes Association, P60-AG10484 (to the Claude Pepper Older Americans Independence Center of Wake Forest University Health Sciences), U01-HL137181 (to J.R.O.), R01-HL093093 (to S.T.M.), 1U24CA237617 and 5U24HG009446 (to X.S.L.), HHSN268201600018C, HHSN268201600001C, HHSN268201600002C, HHSN268201600003C and HHSN268201600004C (to C.L.K.), U01-HL072524, R01-HL104135-04S1, U01-HL054472, U01-HL054473, U01-HL054495, U01-HL054509 and R01-HL055673-18S1 (to M.R.I., S.A. and D.K.A.), Swedish Research Council grant no. 201606830 (to G.H.), grant nos. HHSN268201800010I, HHSN268201800011I, HHSN268201800012I, HHSN268201800013I, HHSN268201800014I and HHSN268201800015I (to A.C.), HHSN268201700001I, HHSN268201700002I, HHSN268201700003I, HHSN268201700005I and HHSN268201700004I (to E.B.), and R01-HL134320 (to C.M.B.). WGS for the TOPMed program was supported by the NHLBI. Centralized read mapping and genotype calling, along with variant quality metrics and filtering were provided by the TOPMed Informatics Research Center (no. 3R01HL-117626-02S1; contract no. HHSN268201800002I). Phenotype harmonization, data management, sample identity quality control and general study coordination were provided by the TOPMed Data Coordinating Center (no. 3R01HL-120393-02S1; contract no. HHSN268201800001I). We gratefully acknowledge the studies and participants who provided biological samples and data for TOPMed. The full study-specific acknowledgements are detailed in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
X. Li, Z.L., H.Z., G.R.A., J.I.R., C.J.W., G.M.P., P.N. and X. Lin designed the experiments. X. Li, Z.L., H.Z. and X. Lin performed the experiments. X. Li, Z.L., H.Z., S.M.G., Y.L., H.C., R.S., R.D., D.K.A., S.A., C.M.B., L.F.B., J.B., E.B., D.W.B., J.G.B., M.P.C., A.C., L.A.C., J.E.C., B.I.F., X.G., G.H., M.R.I., S.L.R.K., S.K., A.T.K., C.L.K., C.C.L., X.S.L., M.C.M., A.W.M., L.W.M., R.A.M., S.T.M., B.D.M., M.E.M., J.E.M., A.C.M., J.R.O., N.D.P., A.P., J.M.P., P.A.P., B.M.P., S.R., K.M.R., S.S.R., J.A.S., H.K.T., M.Y.T., R.S.V., F.F.W., D.E.W., Z.W., J.G.W., L.R.Y., B.M.N., S.R.S., G.R.A., J.I.R., C.J.W., G.M.P., P.N. and X. Lin acquired, analyzed or interpreted the data. G.M.P., P.N. and the NHLBI TOPMed Lipids Working Group provided administrative, technical or material support. X. Li, Z.L., S.M.G., J.I.R., G.M.P., P.N. and X. Lin drafted the manuscript and revised it according to suggestions by the coauthors. All authors critically reviewed the manuscript, suggested revisions as needed and approved the final version.
Corresponding author
Ethics declarations
Competing interests
S.A. reports equity and employment by 23andMe. L.A.C. spends part of her time consulting for the Dyslipidemia Foundation, a nonprofit company, as a statistical consultant. X.S.L. is cofounder, board member and scientific advisory board of GV20 Oncotherapy, board member of the scientific advisory board of 3DMedCare, consultant of Genentech and is a recipient of research grants from Sanofi and Takeda, all unrelated to the present work. For The Amish Research Program receives partial support from Regeneron Pharmaceuticals for B.D.M. M.E.M reports a grant from Regeneron Pharmaceuticals that is unrelated to the present work. B.M.P. serves on the steering committee of the Yale Open Data Access Project funded by Johnson & Johnson. S.R. reports interests in Jazz Pharmaceuticals, Eisai and Respircardia, all unrelated to the present work. Z.W. cofounded Rgenta Therapeutics and directs its scientific advisory board. B.M.N. is on the scientific advisory board of Deep Genomics, and is a consultant for CAMP4 Therapeutics, Takeda and Biogen. S.R.S. is a consultant to NGM Biopharmaceuticals and Inari Agriculture. He is also on the scientific advisory board of Veritas Genetics. G.R.A. is an employee of Regeneron Pharmaceuticals and owns stock and stock options for Regeneron Pharmaceuticals. The spouse of C.J.W. works at Regeneron Pharmaceuticals. P.N. reports grants from Amgen, Apple and Boston Scientific, and consulting income from Apple and Blackstone Life Sciences, all unrelated to the present work. X. Lin is a consultant to AbbVie Pharmaceuticals.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–17 and Note
Supplementary Tables
Supplementary Tables 1–14
Rights and permissions
About this article
Cite this article
Li, X., Li, Z., Zhou, H. et al. Dynamic incorporation of multiple in silico functional annotations empowers rare variant association analysis of large whole-genome sequencing studies at scale. Nat Genet 52, 969–983 (2020). https://doi.org/10.1038/s41588-020-0676-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-0676-4
This article is cited by
-
Enhancing prediction accuracy of coronary artery disease through machine learning-driven genomic variant selection
Journal of Translational Medicine (2024)
-
Genetic variation across and within individuals
Nature Reviews Genetics (2024)
-
BIGKnock: fine-mapping gene-based associations via knockoff analysis of biobank-scale data
Genome Biology (2023)
-
Unravelling the genetic architecture of human complex traits through whole genome sequencing
Nature Communications (2023)
-
Accurate and efficient estimation of local heritability using summary statistics and the linkage disequilibrium matrix
Nature Communications (2023)