Abstract
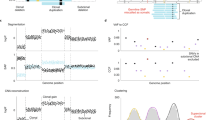
Most cancer genomic data are generated from bulk samples composed of mixtures of cancer subpopulations, as well as normal cells. Subclonal reconstruction methods based on machine learning aim to separate those subpopulations in a sample and infer their evolutionary history. However, current approaches are entirely data driven and agnostic to evolutionary theory. We demonstrate that systematic errors occur in the analysis if evolution is not accounted for, and this is exacerbated with multi-sampling of the same tumor. We present a novel approach for model-based tumor subclonal reconstruction, called MOBSTER, which combines machine learning with theoretical population genetics. Using public whole-genome sequencing data from 2,606 samples from different cohorts, new data and synthetic validation, we show that this method is more robust and accurate than current techniques in single-sample, multiregion and longitudinal data. This approach minimizes the confounding factors of nonevolutionary methods, thus leading to more accurate recovery of the evolutionary history of human cancers.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data in Fig. 3a were from Nik-Zainal et al.3. Data in Fig. 3b were from Griffith et al.20. Data in Fig. 3c–e were cases from Cross et al.21, here re-sequenced at higher sequencing depth. Sequence data from those colorectal cancer cases have been deposited at the European Genome-phenome Archive (EGA), which is hosted by the European Bioinformatics Institute and the Centre for Genomic Regulation, under accession no. EGAS00001003066. Further information about EGA can be found at https://ega-archive.org. Diploid SNVs and copy-number calls are available in the Supplementary Data. Data in Fig. 3f were from Lee et al.24. Data in Fig. 4 are available through the PCAWG consortium25. Whole-genome variant call data in Fig. 5, which were not available from the original publication, were provided upon email request by Korber et al.28.
Code availability
MOBSTER is available as an R package at https://github.com/sottorivalab/mobster; future updates, as well as all vignettes and manuals, are maintained at https://caravagn.github.io/mobster. A repository with all Supplementary Data are available at https://github.com/sottorivalab/mobster_supp_data. Supplementary Data contain vignettes that show the analysis of single-sample and multiregion simulated tumors, the whole analysis of multiregion colorectal samples and single-sample lung cancers, and summary results from the PCAWG and GBM cohorts. Somatic SNVs and copy-number calls used for the analysis of multiregion colorectal samples are also available as Supplementary Data. The implementation of all other R packages that we have developed are available at https://caravagn.github.io/.
References
Greaves, M. & Maley, C. C. Clonal evolution in cancer. Nature 481, 306–313 (2012).
Turajlic, S., Sottoriva, A., Graham, T. & Swanton, C. Resolving genetic heterogeneity in cancer. Nat. Rev. Genet. 20, 404–416 (2019).
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012).
Dentro, S. C., Wedge, D. C. & Van Loo, P. Principles of reconstructing the subclonal architecture of cancers. Cold Spring Harb. Perspect. Med. 7, a026625 (2017).
Roth, A. et al. PyClone: statistical inference of clonal population structure in cancer. Nat. Meth. 11, 396–398 (2014).
Deshwar, A. G. et al. PhyloWGS: reconstructing subclonal composition and evolution from whole-genome sequencing of tumors. Genome Biol. 16, 35 (2015).
Miller, C. A. et al. SciClone: inferring clonal architecture and tracking the spatial and temporal patterns of tumor evolution. PLoS Comput. Biol. 10, e1003665 (2014).
Lynch, M. et al. Genetic drift, selection and the evolution of the mutation rate. Nat. Rev. Genet. 17, 704–714 (2016).
Williams, M. J., Werner, B., Barnes, C. P., Graham, T. A. & Sottoriva, A. Identification of neutral tumor evolution across cancer types. Nat. Genet. 48, 238–244 (2016).
Kessler, D. A. & Levine, H. Large population solution of the stochastic Luria–Delbruck evolution model. Proc. Natl Acad. Sci. USA 110, 11682–11687 (2013).
Kessler, D. A. & Levine, H. Scaling solution in the large population limit of the general asymmetric stochastic Luria–Delbrück evolution process. J. Stat. Phys. 158, 783–805 (2015).
Durrett, R. Population genetics of neutral mutations in exponentially growing cancer cell populations. Ann. Appl. Probabil. 23, 230–250 (2013).
Nicholson, M. D. & Antal, T. Universal asymptotic clone size distribution for general population growth. Bull. Math. Biol. 78, 2243–2276 (2016).
Griffiths, R. C. & Tavaré, S. The age of a mutation in a general coalescent. Stoch. Models 14, 273–295 (1998).
Sun, R. et al. Between-region genetic divergence reflects the mode and tempo of tumor evolution. Nat. Genet. 49, 1015–1024 (2017).
Williams, M. J. et al. Quantification of subclonal selection in cancer from bulk sequencing data. Nat. Genet. 50, 895–903 (2018).
Hartl, D. L. & Clark, A. G. Principles of Population Genetics (Sinauer Associates, Inc., 2006).
Luria, S. E. & Delbrück, M. Mutations of bacteria from virus sensitivity to virus resistance. Genetics 28, 491–511 (1943).
Graham, T. A. & Sottoriva, A. Measuring cancer evolution from the genome. J. Pathol. 241, 183–191 (2017).
Griffith, M. et al. Optimizing cancer genome sequencing and analysis. Cell Systems 1, 210–223 (2015).
Cross, W. et al. The evolutionary landscape of colorectal tumorigenesis. Nat. Ecol. Evol. 2, 1661–1672 (2018).
Martincorena, I. et al. Universal patterns of selection in cancer and somatic tissues. Cell 171, 1–13 (2017).
Zapata, L. et al. Negative selection in tumor genome evolution acts on essential cellular functions and the immunopeptidome. Genome Biol. 19, 924 (2018).
Lee, J. J.-K. et al. Tracing oncogene rearrangements in the mutational history of lung adenocarcinoma. Cell 177, 1842–1857.e21 (2019).
The ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. Pan-cancer analysis of whole genomes. Nature 578, 82–93 (2020).
Gerstung, M. et al. The evolutionary history of 2,658 cancers. Nature 578, 122–128 (2020).
Williams, M. J. et al. Measuring the distribution of fitness effects in somatic evolution by combining clonal dynamics with dN/dS ratios. eLife Sci. 9, 612 (2020).
Körber, V. et al. Evolutionary trajectories of IDHWT glioblastomas reveal a common path of early tumorigenesis instigated years ahead of initial diagnosis. Cancer Cell 35, 692–704.e12 (2019).
Barthel, F. P. et al. Longitudinal molecular trajectories of diffuse glioma in adults. Nature 576, 112–120 (2019).
Shah, S. P. et al. The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature 486, 395–399 (2012).
Andor, N. et al. Pan-cancer analysis of the extent and consequences of intratumor heterogeneity. Nat. Med. 22, 105–113 (2016).
Morris, L. G. T. et al. Pan-cancer analysis of intratumor heterogeneity as a prognostic determinant of survival. Oncotarget 7, 10051–10063 (2016).
Jamal-Hanjani, M. et al. Tracking the evolution of non-small-cell lung cancer. N. Engl. J. Med. 376, 2109–2121 (2017).
Espiritu, S. M. G. et al. The evolutionary landscape of localized prostate cancers drives clinical aggression. Cell 173, 1003–1013.e15 (2018).
Salcedo, A. et al. A community effort to create standards for evaluating tumor subclonal reconstruction. Nat. Biotechnol. 38, 97–107 (2020).
Yang, L. et al. An enhanced genetic model of colorectal cancer progression history. Genome Biol. 20, 168 (2019).
Yates, L. R. et al. Genomic evolution of breast cancer metastasis and relapse. Cancer Cell 32, 169–184.e7 (2017).
Gundem, G. et al. The evolutionary history of lethal metastatic prostate cancer. Nature 520, 353–357 (2015).
Noorani, A. et al. Genomic evidence supports a clonal diaspora model for metastases of esophageal adenocarcinoma. Nat. Genet. 347, 1–10 (2020).
Navin, N. E. The first five years of single-cell cancer genomics and beyond. Genome Res. 25, 1499–1507 (2015).
Chkhaidze, K. et al. Spatially constrained tumour growth affects the patterns of clonal selection and neutral drift in cancer genomic data. PLoS Comput. Biol. 15, e1007243 (2019).
Fusco, D., Gralka, M., Kayser, J., Anderson, A. & Hallatschek, O. Excess of mutational jackpot events in expanding populations revealed by spatial Luria–Delbrück experiments. Nat. Commun. 7, 12760 (2016).
Teh, Y. W. Dirichlet processes. in Encyclopedia of Machine Learning (eds Sammut, C. & Webb, G.) 280–287 (Springer, 2011).
Ghahramani, Z., Jordan, M. I. & Adams, R. P. Tree-structured stick breaking for hierarchical data. in Advances in Neural Information Processing Systems (eds Lafferty, J. D. et al.) 2319–2327 (Neural Information Processing Systems, 2010).
Ma, Z. & Leijon, A. Bayesian estimation of beta mixture models with variational inference. IEEE Trans. Pattern Anal. Mach. Intell. 33, 2160–2173 (2011).
Clauset, A., Shalizi, C. R. & Newman, M. E. J. Power-law distributions in empirical data. SIAM Rev. 51, 661–703 (2009).
Schröder, C. & Rahmann, S. A hybrid parameter estimation algorithm for beta mixtures and applications to methylation state classification. Algorithms Mol. Biol. 12, 21 (2017).
Biernacki, C., Celeux, G. & Govaert, G. Assessing a mixture model for clustering with the integrated completed likelihood. IEEE Trans. Pattern Anal. Mach. Intell. 22, 719–725 (2000).
Acknowledgements
A.S. is supported by the Wellcome Trust (202778/B/16/Z) and Cancer Research UK (A22909). T.G. is supported by the Wellcome Trust (202778/Z/16/Z) and Cancer Research UK (A19771). We thank the Medical Research Council (MR/P000789/1) for funding A.S. and the National Institutes of Health (NCI U54 CA217376) for funding A.S and T.A.G. C.P.B. thanks the Wellcome Trust (209409/Z/17/Z) for funding. L.C. thanks Cancer Research UK (A24566) and Children with Cancer UK (17–235) for funding. This work was also supported by a Wellcome Trust award to the Centre for Evolution and Cancer (105104/Z/14/Z). L.Z. is supported by the European Union’s Horizon 2020 research and innovation program under the Marie Skłodowska–Curie Research Fellowship scheme (846614). We thank N. Matthews and the Tumour Profiling Unit at the ICR for their support with next-generation sequencing. We thank V. Körber and T. Höfer for sharing their data and for fruitful discussion around the glioblastoma cohort.
Author information
Authors and Affiliations
Contributions
G.C. conceived, designed and implemented the method. T.H. and K.C. developed the spatial tumor growth simulations. T.H. and M.W. generated the data for synthetic tests. G.C., T.H., M.W. and D.N. carried out and analyzed these tests. G.C., M.W. and L.Z. analyzed the data. W.C., G.D.C. and A.A. provided input and support with the analysis. G.S., C.B., T.A.G. and A.S. supervised the method design. L.C. contributed to study supervision. A.S. and T.A.G. conceived and supervised the study. All authors contributed to and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Synthetic tests with MOBSTER.
Example MOBSTER fit of synthetic single-sample tumors (details in Supplementary Note 1). All boxplots and violins show mean and inter quartile range (IQR), upper whisker is 3rd quartile +1.5 * IQR and lower whisker is 1st quartile - 1.5 * IQR. a,b, Subclonal reconstruction with MOBSTER, against standard methods (variational fit of a Dirichlet finite mixture, and a Markov Chain Monte Carlo sampling for a Dirichlet Process). These methodologies are at the basis of many approaches in the field. The test uses synthetic data from n = 150 simulated tumors (n = 120 with one subclone, and n = 30 without subclones), generated from a stochastic branching process. We report the logarithm of the ratio between the number of clones fit (kfit) and the true number (ktrue). Tests show different values of the concentration parameter α, which tunes the propensity to call clusters. Values (for example, α = 10−4) are point estimates, but we also test also a Dirichlet Process where α is learnt from the data using a Gamma prior. c, Proportion of mutations assigned to MOBSTER’s tail changes with coverage, at fixed 100% tumor purity. We span coverage from 40x to 200x, using a subset of n = 80 tumors from the test in panels (a, b). The red dashed line is the median tail size across the test set (obtained from simulated tumor); tests suggest the coverage required to fit a tail. d, As for coverage, we tested with n = 320 tumors (n = 80 per configuration) the ability of detecting tails as a function of purity, fixing a coverage of 120x. The average tail size is reported (number of SNVs assigned to the tail in the fit).
Extended Data Fig. 2 The hitchhiker mirage in multi-region sequencing data.
a,b, Evolutionary history of a tumor with one subclone. After the first cancer cell gives rise to the tumor (blue founder clone), the population evolves neutrally accumulating passenger mutations (orange), until eventually a subclonal driver occurs triggering a new subclonal expansion (green, with its own tail). The subclonal driver, together with its passenger hitchhikers (orange) will rise in frequency with the subclonal expansion, forming a subclonal cluster in the VAF distribution. However, some early hitchhikers will also be present elsewhere in the tumor as part of the tail of the founder clone. In the example of perfect cell doubling, we expect mutations in the first doubling to be in 50% of the cells of the tumor, mutations in the second doubling to be in 25% etc. We take monoclonal biopsies S1 and S2, and find the founder clone (S1) and a subclonal sweep (S2). c, The hitchhiker mirage (Supplementary Note 2) is a confounder determined by passengers that hitchhike to the subclonal driver in S2, but diffuse neutrally in S1 (orange). This can be seen in the S1 vs S2 VAF scatter, where the orange mutations do not travel together in the two samples, because cells in S1 do not harbor the subclonal driver (while those in S2 do). The VAF scatter shows that orange hitchhikers can generate an extra cluster with Binomial parameters 0.5/0.2 for S1/S2, on top of the green clone with different parameters (S1/S=0.5/0). Moreover, extra clusters are generated by fitting tail mutations with a Binomial mixture, further inflating the true number of clones (k = 2) and suggesting false clonal sweeps (from which the illusion of a non-existing clonal expansion). If we remove mutations assigned to a tail by MOBSTER we clean up the signal and retrieve the true clonal architecture.
Extended Data Fig. 3 The MRCA fallacy in multi-region sequencing data.
a, Every cell always has an ancestor, and the cell starting the tumor is the Most Recent Common Ancestor (MRCA) of the whole tumor. We never sequence that cell, we sequence some of its progeny. We can travers the phylogeny of cell divisions backward, and determine the MRCA of all biopsy cells (red and blue), or the MRCA of all biopsies (purple). b, The ancestor effect (Supplementary Note 2) is the MRCA of cells from a spatially-localized biopsy, compared to other biopsies. Hence, mutations that are observed at high frequency in one biopsy are not necessarily due to selection. We simulate the growth of a 2D neutral tumor, and sample two biopsies (100% purity, S1 and S2). Both samples contain truncal mutations; each biopsy also contains private mutations (green and orange) that are clonal within the sample but are not due to selection. When we generate a virtual staining of all cells that harbor the mutations in a cluster, we see the separation between cells in S1 and S2, and the branched evolutionary structure in the clone tree that is not due to selection, but to spatial sampling (Extended Data Figure 4). c, The admixing deception stems from spatial tumor intermixing, with cells that are close in space, but genetically distant in the phylogeny. In this example, whereas S1 is a bulk of closely related cells, suffering only from the ancestor fallacy, S2 contains a mixture of cell lineages from distinct parts of the tree (here split in right and left). Intermixing is bond to happen since distant parts of a phylogeny must mix somewhere in space; again, in this example, no selection is at play. From these biopsies, we find truncal (black) and private mutations in S1(green, ancestor fallacy). In S2 we find a mixture of lineages (orange and blue) peaked like subclonal clusters (here we omit neutral tails for simplicity). The orange and blue clusters deviate by an offset that is determined by the level of admixing, which is unknown a priori (see the VAF of S2 in Extended Data Figure 4).
Extended Data Fig. 4 Effects of the MRCA fallacy and the admixing deception in multi-region sequencing data.
a, Phylogenetic tree of cellular divisions in a neutral expansion, that is, inside a clonal expansion triggered by a driver hitting the grey cell. The tree shows the sampling of 2 biopsies (red and blue), and the MRCAs. For example, the mutational load present in the red MRCA will characterize cells in the red biopsy. b, Data distribution and the associated phylogenetic tree show how our estimate of the true evolution of this tumor is confounded by spatial sampling. Mutations that accrue in the lineages from which the MRCAs originate, will create clusters in the data. The corresponding phylogenetic model will also show an inflated number of clonal events (purple MRCA), and branches that do not represent real selection-driven branched evolution (red and blue MRCAs). c, Example admixing deception in the blue biopsy, where two independent lineages are represented. Admixing can be even or uneven, depending on the proportion of lineages (left versus right) in the biopsy. Remark that no subclonal selection is at play in this example. d, If we sequence the above biopsies, we find truncal mutations (gray and blue), and a number of clusters that look like genuine subclones. The admixing effect is observed on the vertical of the blue biopsy. In the even case (50% each), the admixing generates one 50% peak for both independent lineages. According to the relation between the frequencies of the observed ancestors, we can also fit two different trees to data; notice that the branching structure presented in both of them is the result of the confounders and does not reflect actual branched evolution. In the uneven case (60% versus 40%), the two admixing peaks separate, originating 2 peaks hitting at the frequencies of 40% and 60%. This shows the pervasive effect of admixing, with up to 8 clusters in this simple scenarios.
Extended Data Fig. 5 Interpreting clone trees as clonal evolution models.
Interpreting clone trees that contain spatial confounders as clonal evolution models can be difficult. We show an example consistent with the data shown in these Extended Data Figures 2–5. a, All the spatial confounders discussed in Supplementary Note 2 lead to additional nodes and branching structures in the estimated clone tree. These confounders need to be accounted for in a clonal deconvolution analysis, if we seek to identify waves of clonal expansions due to positive selection. The translation of clusters that originate from confounders, into clonal expansions due to selection is misleading, and the inferred clonal evolution is much more complex than the actual one. b, For clarification, a phylogenetic tree at the single cell level of the tumor, showing that clusters B, C, E and F are arbitrary ancestors identified by the specific spatial bias of the measurement.
Supplementary information
Supplementary Information
Supplementary Note and Figs. 1–23
Supplementary Data
Supplementary data released with the paper (with R vignettes, MOBSTER-installable R package).
Rights and permissions
About this article
Cite this article
Caravagna, G., Heide, T., Williams, M.J. et al. Subclonal reconstruction of tumors by using machine learning and population genetics. Nat Genet 52, 898–907 (2020). https://doi.org/10.1038/s41588-020-0675-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-0675-5
This article is cited by
-
Computational validation of clonal and subclonal copy number alterations from bulk tumor sequencing using CNAqc
Genome Biology (2024)
-
Tumor phylogeography reveals block-shaped spatial heterogeneity and the mode of evolution in Hepatocellular Carcinoma
Nature Communications (2024)
-
Clinical application of tumour-in-normal contamination assessment from whole genome sequencing
Nature Communications (2024)
-
State-dependent evolutionary models reveal modes of solid tumour growth
Nature Ecology & Evolution (2023)
-
Immune selection determines tumor antigenicity and influences response to checkpoint inhibitors
Nature Genetics (2023)