Abstract
During aging, progenitor cells acquire mutations, which may generate clones that colonize the surrounding tissue. By middle age, normal human tissues, including the esophageal epithelium (EE), become a patchwork of mutant clones. Despite their relevance for understanding aging and cancer, the processes that underpin mutational selection in normal tissues remain poorly understood. Here, we investigated this issue in the esophageal epithelium of mutagen-treated mice. Deep sequencing identified numerous mutant clones with multiple genes under positive selection, including Notch1, Notch2 and Trp53, which are also selected in human esophageal epithelium. Transgenic lineage tracing revealed strong clonal competition that evolved over time. Clone dynamics were consistent with a simple model in which the proliferative advantage conferred by positively selected mutations depends on the nature of the neighboring cells. When clones with similar competitive fitness collide, mutant cell fate reverts towards homeostasis, a constraint that explains how selection operates in normal-appearing epithelium.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Code availability
The code developed in this study has been made publicly available and can be found at https://github.com/gp10/ClonalCOMMUTE and https://doi.org/10.5281/zenodo.3648706.
References
Fuster, J. J. et al. Clonal hematopoiesis associated with TET2 deficiency accelerates atherosclerosis development in mice. Science 355, 842–847 (2017).
Lee-Six, H. et al. Population dynamics of normal human blood inferred from somatic mutations. Nature 561, 473–478 (2018).
Suda, K. et al. Clonal expansion and diversification of cancer-associated mutations in endometriosis and normal endometrium. Cell Rep. 24, 1777–1789 (2018).
Martincorena, I. et al. Somatic mutant clones colonize the human esophagus with age. Science 362, 911–917 (2018).
Yokoyama, A. et al. Age-related remodelling of oesophageal epithelia by mutated cancer drivers. Nature 565, 312–317 (2019).
Alcolea, M. P. et al. Differentiation imbalance in single oesophageal progenitor cells causes clonal immortalization and field change. Nat. Cell Biol. 16, 615–622 (2014).
Alcolea, M. P. & Jones, P. H. Cell competition: winning out by losing notch. Cell Cycle 14, 9–17 (2015).
Murai, K. et al. Epidermal tissue adapts to restrain progenitors carrying clonal p53 mutations. Cell Stem Cell 23, e688 (2018).
Brown, S. et al. Correction of aberrant growth preserves tissue homeostasis. Nature 548, 334–337 (2017).
Ellis, S. J. et al. Distinct modes of cell competition shape mammalian tissue morphogenesis. Nature 569, 497–502 (2019).
Madan, E. et al. Flower isoforms promote competitive growth in cancer. Nature 572, 260–264 (2019).
Kon, S. et al. Cell competition with normal epithelial cells promotes apical extrusion of transformed cells through metabolic changes. Nat. Cell Biol. 19, 530–541 (2017).
Bowling, S., Lawlor, K. & Rodriguez, T. A. Cell competition: the winners and losers of fitness selection. Development 146, dev167486 (2019).
Morata, G. & Ripoll, P. Minutes: mutants of Drosophila autonomously affecting cell division rate. Dev. Biol. 42, 211–221 (1975).
Doupe, D. P. et al. A single progenitor population switches behavior to maintain and repair esophageal epithelium. Science 337, 1091–1093 (2012).
Piedrafita, G. et al. A single-progenitor model as the unifying paradigm of epidermal and esophageal epithelial maintenance in mice. Nat. Commun. 11, 1429 (2020).
Hall, M. W. J., Jones, P. H. & Hall, B. A. Relating evolutionary selection and mutant clonal dynamics in normal epithelia. J. R. Soc. Interface 16, 20190230 (2019).
Rubio, C. A., Liu, F. S., Chejfec, G. & Sveander, M. The induction of esophageal tumors in mice: dose and time dependency. In Vivo 1, 35–38 (1987).
Frede, J., Greulich, P., Nagy, T., Simons, B. D. & Jones, P. H. A single dividing cell population with imbalanced fate drives oesophageal tumour growth. Nat. Cell Biol. 18, 967–978 (2016).
Gerstung, M., Papaemmanuil, E. & Campbell, P. J. Subclonal variant calling with multiple samples and prior knowledge. Bioinformatics 30, 1198–1204 (2014).
Dow, M. et al. Integrative genomic analysis of mouse and human hepatocellular carcinoma. Proc. Natl Acad. Sci. USA 115, E9879–E9888 (2018).
Connor, F. et al. Mutational landscape of a chemically-induced mouse model of liver cancer. J. Hepatol. 69, 840–850 (2018).
Martincorena, I. et al. Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Science 348, 880–886 (2015).
Martincorena, I. et al. Universal patterns of selection in cancer and somatic tissues. Cell 171, 1029–1041.e21 (2017).
Jiang, M. et al. BMP-driven NRF2 activation in esophageal basal cell differentiation and eosinophilic esophagitis. J. Clin. Investig. 125, 1557–1568 (2015).
Lefort, K. et al. Notch1 is a p53 target gene involved in human keratinocyte tumor suppression through negative regulation of ROCK1/2 and MRCKalpha kinases. Genes Dev. 21, 562–577 (2007).
Fernandez-Antoran, D. et al. Outcompeting p53-mutant cells in the normal esophagus by redox manipulation. Cell Stem Cell 25, 329–341 (2019).
Gao, Y. B. et al. Genetic landscape of esophageal squamous cell carcinoma. Nat. Genet. 46, 1097–1102 (2014).
Wu, S. et al. ARID1A spatially partitions interphase chromosomes. Sci. Adv. 5, eaaw5294 (2019).
Mansour, A. A. et al. The H3K27 demethylase Utx regulates somatic and germ cell epigenetic reprogramming. Nature 488, 409–413 (2012).
Seegar, T. C. M. et al. Structural basis for regulated proteolysis by the alpha-Secretase ADAM10. Cell 171, 1638–1648.e37 (2017).
Bray, S. J. Notch signalling in context. Nat. Rev. Mol. Cell Biol. 17, 722–735 (2016).
Lee, P. et al. Phosphorylation of Pkp1 by RIPK4 regulates epidermal differentiation and skin tumorigenesis. EMBO J. 36, 1963–1980 (2017).
Huang, C. S. et al. Crystal structure of Ripk4 reveals dimerization-dependent kinase activity. Structure 26, 767–777.e5 (2018).
Oberbeck, N. et al. The RIPK4–IRF6 signalling axis safeguards epidermal differentiation and barrier function. Nature 574, 249–253 (2019).
Stephenson, N. L. & Avis, J. M. Direct observation of proteolytic cleavage at the S2 site upon forced unfolding of the Notch negative regulatory region. Proc. Natl Acad. Sci. USA 109, E2757–E2765 (2012).
Weber, S. et al. The disintegrin/metalloproteinase Adam10 is essential for epidermal integrity and Notch-mediated signaling. Development 138, 495–505 (2011).
Klein, A. M., Doupe, D. P., Jones, P. H. & Simons, B. D. Kinetics of cell division in epidermal maintenance. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 76, 021910 (2007).
Martincorena, I., Jones, P. H. & Campbell, P. J. Constrained positive selection on cancer mutations in normal skin. Proc. Natl Acad. Sci. USA 113, E1128–E1129 (2016).
Simons, B. D. Deep sequencing as a probe of normal stem cell fate and preneoplasia in human epidermis. Proc. Natl Acad. Sci. USA 113, 128–133 (2016).
Lynch, M. D. et al. Spatial constraints govern competition of mutant clones in human epidermis. Nat. Commun. 8, 1119 (2017).
Zhu, M. et al. Somatic mutations increase hepatic clonal fitness and regeneration in chronic liver disease. Cell 177, 608–621.e12 (2019).
Moreno, E. & Basler, K. dMyc transforms cells into super-competitors. Cell 117, 117–129 (2004).
Tanimura, N. & Fujita, Y. Epithelial defense against cancer (EDAC). Semin. Cancer Biol. https://doi.org/10.1016/j.semcancer.2019.05.011 (2019).
Lowell, S., Jones, P., Le Roux, I., Dunne, J. & Watt, F. M. Stimulation of human epidermal differentiation by delta-notch signalling at the boundaries of stem-cell clusters. Curr. Biol. 10, 491–500 (2000).
Bras-Pereira, C. & Moreno, E. Mechanical cell competition. Curr. Opin. Cell Biol. 51, 15–21 (2018).
Franco, J. J., Atieh, Y., Bryan, C. D., Kwan, K. M. & Eisenhoffer, G. T. Cellular crowding influences extrusion and proliferation to facilitate epithelial tissue repair. Mol. Biol. Cell 30, 1890–1899 (2019).
Eisenhoffer, G. T. et al. Crowding induces live cell extrusion to maintain homeostatic cell numbers in epithelia. Nature 484, 546–549 (2012).
Roshan, A. et al. Human keratinocytes have two interconvertible modes of proliferation. Nat. Cell Biol. 18, 145–156 (2016).
Watt, F. M., Jordan, P. W. & O’Neill, C. H. Cell shape controls terminal differentiation of human epidermal keratinocytes. Proc. Natl Acad. Sci. USA 85, 5576–5580 (1988).
Clayton, E. et al. A single type of progenitor cell maintains normal epidermis. Nature 446, 185–189 (2007).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997v2 (2013).
Acknowledgements
This work was supported by grants from the Wellcome Trust to the Wellcome Sanger Institute (098051 and 296194) and Cancer Research UK Programme Grants to P.H.J. (C609/A17257 and C609/A27326). G.P. is supported by a Talento program fellowship from Comunidad de Madrid. B.A.H. and M.W.J.H. are supported by the Medical Research Council (Grant-in-Aid to the MRC Cancer unit grant no. MC_UU_12022/9 and NIRG to B.A.H. grant no. MR/S000216/1). M.W.J.H. acknowledges support from the Harrison Watson Fund at Clare College, Cambridge. B.A.H. acknowledges support from the Royal Society (grant no. UF130039). I.M. is funded by Cancer Research UK (C57387/A21777). S.D. benefited from the award of an ESPOD fellowship, 2018-21, from the Wellcome Sanger Institute and the European Bioinformatics Institute EMBL-EBI. We thank E. Choolun and staff at the MRC ARES and Sanger RSF facilities for technical support.
Author information
Authors and Affiliations
Contributions
B.C., M.P.A., A.W. and J.C.F. designed experiments. B.C., M.P.A., A.W., A.H. and J.C.F. performed experiments. I.M. adapted Shearwater for mice. B.C., R.K.S., S.H.O., S.C.D., C.K. and M.W.J.H. analyzed sequence data. G.P., M.W.J.H. and B.A.H. performed clone simulations. B.C., M.W.J.H., G.P., B.A.H. and P.H.J. wrote the paper. B.A.H., M.G. and P.H.J. supervised the research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Mutational landscape of the mouse EE.
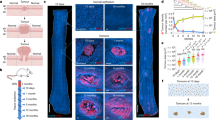
a, The single progenitor model: All progenitor cells in the EE basal layer are functionally equivalent and divisions result in producing either 2 progenitor cells that will persist in the tissue, 2 differentiating cells that will cease division and stratify or 1 cell of each type (Supplementary Note.). In homeostasis, the likelihood of the two progenitor and two differentiating cell outcomes is equal (A). Mutations may tip the balance towards a non-neutral behavior resulting in clonal growth if they favor proliferation of daughter cells (B) or clone loss if they favor differentiation (C). b, Representative H&E images of esophagus from DEN-treated (12 months post-DEN) and control mice (n = 3 mice). White dots delineate the basal cell layer (b). sb = suprabasal layers, str = stroma. Scale-bars: 20 µm (left panels) and 250 µm (right panels). c, d, Confocal images (Dapi staining; 12 month time-point) and number of basal cells per mm2 of control or DEN-treated EE. Dots indicate field of views (n numbers for control/DEN, respectively, are: 10d = 20/30, 1 m = 30/30, 3 m = 30/29, 6 m = 30/27, 12 m = 29/30; from 3 mice (except 10d control = 2 mice), see Supplementary Table 1. Mean ± s.e.m.; Two-sided Mann-Whitney test. Scale bars: 20 µm. e, List of the 192 mouse genes sequenced. Genes in purple are common between the mouse and human bait sets. f, Median sequencing coverage of the 192 genes (sample size: MD5690 = 41, MD5691 = 37, MD5692 = 43, MD5693 = 42, MD5694 = 36, MD5695 = 46; Mean ± s.e.m.). g, Grid maps of the 3 control and 3 DEN-treated EEs sequenced. Colour indicates number of mutations.
Extended Data Fig. 2 Genetic positive selection in control mouse EE and comparison of mutational landscapes in mutagen treated mouse and aging human EE.
a, Observed-to-expected ratios for Notch1 missense and nonsense + splice substitutions and indels in control aged mouse EE (66 mutations; R package dndscv (24)). See Supplementary Table 3. b, Number and type of Notch1 mutations in control aged mouse EE. c, Distribution of mutations within the Notch1 domains in the normal EE of control and DEN-treated aged mice. d, Age range and number of mouse and human esophagus used for the inter-species comparison. e, f Gene dN/dS ratios for missense (e) and nonsense + splice (f) substitutions in mouse and human esophagus. The 16 genes identified by dN/dS analysis as positively selected in human EE are shown (Supplementary Table 5). Green boxes indicate positively selected genes for each species. g, Percentage of EE carrying non-synonymous mutations for each gene. Range indicates upper and lower bound estimates. h, Distribution of mutations within the Notch1 gene in normal human and DEN-treated mouse EE.
Extended Data Fig. 3 Measurement of clone areas.
a, Confocal image of the whole EE of an AhcreERTRosa26flEYFP/wt control animal, 1 year post-induction. Scale bars: 2 mm (left image), 150 μm (insets) b, Images illustrate the relation between projected and basal tissue areas of a AhcreERTRosa26flEYFP/wt control animal, 1 year post-induction. c, Quantifications and correlation between projected vs basal cell clone areas (each dot represents a clone, n = 164). Scale bars: 1 mm (top image), 20 μm (bottom images).
Extended Data Fig. 4 Average cell division and stratification rates are not altered in DEN-treated EE.
a, Protocol: Rosa26M2rtTA/TetO-HGFP mice received DEN or vehicle for 2 months. 8 weeks later mice received doxycycline (Doxy) for 4 weeks and tissues were collected 0 and 7 days after stopping Doxy. HGFP dilution was used to infer cell division rate (b, Supplementary Note; Methods). c, Confocal images showing HGFP in basal cells of control and DEN-treated EE at 0 and 7 days post-Doxy withdrawal. Scale bars: 20 µm. d, Basal cell HGFP intensity in control and DEN-treated EE at the indicated time points post-Doxy withdrawal (number per cells: C0 = 4,296, C7 = 2,599, D0 = 4,766 and D7 = 4,177). Each box represents a mouse, median (central box line) 25th-75th percentiles (box), 1st-99th percentiles (whiskers) and outliers (dots). See Supplementary Table 7. e, Protocol: EdU lineage tracing. Mice received DEN or vehicle and tissues were collected 6 months later. 48 hours before collection mice received EdU to label proliferating basal cells in S phase and track the behavior of their daughters. f, Images showing EdU staining in suprabasal (differentiated) and basal layers of EE from control and DEN-treated mice. Scale bars: 20 µm. g, Percentage of total (basal and suprabasal) EdU-labelled cells 48 h after EdU injection in control and DEN-treated mice (Two-sided Student’s t-test; mean ± s.e.m., n = 6 mice, 1,873 and 2,080 EdU cells, respectively). h, Stratification rate measured as the ratio of suprabasal to total (basal and suprabasal) EdU+ cells 48 h after injection (Two-sided Student’s t-test; mean ± s.e.m., n = 6 mice). See Supplementary Table 8 for EdU data.
Extended Data Fig. 5 Whole exome sequencing of single clones isolated from the EE of DEN-treated mice.
a, Size distribution of sequenced confetti clones (right) vs all confetti clones present in the tissues (left). The number of clones for each category is reported in brackets. b, Cumulative sequencing coverage of the whole exome clone triplicate samples. c, Distribution of the variant allele fraction (VAF) for the mutations common within triplicates in each clone. Red line indicates the cut off for analysis, VAF > 0.3. d, e, Percentage of silent, missense, nonsense and splice mutations and indels identified from whole exome sequencing of single clones (d, % mutations per individual clone; e, % of mutations for all clones). f, Number of mutations observed at each trinucleotide in the isolated clones. g, Observed-to-expected (dN/dS) ratios for missense and truncating (nonsense + essential splice site) substitutions and indels for the genes under significant (q < 0.05) positive selection in the isolated clones (100,544 mutations; R package dndscv (24)). h, Number of silent, missense, nonsense and splice mutations and indels detected in each of the significantly positively selected genes identified by dN/dS analysis in the isolated clones.
Extended Data Fig. 6 Chromosomal alterations are rare in DEN-treated EE.
a, Summary of chromosomal alterations found in isolated clones from the EE of DEN-treated mice. 2n1 (top) is a representative example of a clone without chromosomal alterations. Four out of 250 clones (8141n11, 8141n16, 8141n5 and 8141n6) showed alterations (indicated by colour). Only alterations present in all 3 triplicates (b) were considered valid calls.
Extended Data Fig. 7 Simulations for range of parameters and modelling assumptions.
a, Model used for lattice-based simulations of mutant clone dynamics. Grid consists of wild-type (N) and mutant (M) cells (mutation rate χmut). N and M cells show equal division potential and once a cell divides, it replaces a neighbor (replacement rate Λ). M and N populations differ in their resilience to be displaced (differentiation) by other cells (δN vs δM). b, Number of neighboring cells per basal cell in 1 year post-DEN treated or control mouse EE, quantified from confocal images of tissues stained with Dapi and Keratin 14 (left image depicts control tissue). n = 4 mice, 2,360 cells. Scale-bar 20 µm. c. Weeks taken until 100% of cells in a simulation are mutant (in gray those not reaching 100% within 52 weeks). L = 0.8 for all simulations shown. d, Clone sizes for sweeps of Λ, δM, or χmut (non-varying parameters are fixed at Λ = 0.8; δM = 0.5; χmut = 0.02). e, Clone sizes for χmut. Time is scaled by the characteristic time, Tc (Supplementary Note.). f, Clone sizes for simulations with 4-, 6- or 8-cell neighborhoods. Λ = 0.8, δM = 0.5, χmut = 0.02. g, Clone sizes for δM values drawn from \({\cal{F}} = \left( {1 - Gamma\left( {\kappa ,1/\kappa } \right)} \right)\) for a range of κ values. Λ = 0.8, χmut = 0.1. h, Clone sizes for off-lattice simulations using the HAL simulation framework. Time points defined by the number of simulation steps (st) taken. Mutations are either neutral, reduce the cell death rate, reduce the division-inhibiting effect of neighboring cells, or increase a bias towards division compared to the WT cells.
Extended Data Fig. 8 Cell fate imbalance at clone borders drives clonal dynamics in the DEN-mutated EE.
a, Cartoon illustrating in vivo genetic lineage tracing using AhcreERTRosa26wt/DNM-GFP reporter mice. Upon injection of the drugs tamoxifen (TAM) and ß-napthoflavone (BNF), Cre-mediated recombination results in the heritable expression of the highly competitive dominant negative allele of Maml-1 fused to GFP fluorescent protein (DN-Maml1), which will then be expressed in all the progeny of the single marked cells, generating clusters of labelled mutant clones. b, Protocol: AhcreERTRosa26flEYFP/wt (YFP-Cre) and AhcreERTRosa26wt/DNM-GFP (MAML-Cre) mice were treated with DEN for 2 months and clonal labelling was induced after the last dosage. Tissues were harvested at the indicated time points. c, Violin plots depicting the distributions of individual YFP or DN-Maml1 clones (projected areas) from YFP-Cre or MAML-Cre DEN-treated mice collected at the indicated time points following mutagen treatment. Lines show median and quartiles. Number of mice (and clones) in MAML-Cre/YFP-Cre: 10d = 3/3 (786/15,092), 1 m = 5/3 (522/5,682), 3 m = 3/3 (249/539), 6 m = 4/4 (204/281), 12 m = 3/3 (101/188). See Supplementary Table 14. Two-sided Two-sample Kolmogorov-Smirnov test. d, Schematic of the AhcreERTRosa26flConfetti/DNM-GFP (Confetti-MAML-Cre) and possible outcomes following induction with BNF and TAM.
Supplementary information
Supplementary Information
Supplementary Note
Supplementary Tables 1–15
Supplementary Table 1. Number of basal cells per mm2 of control and DEN-treated mouse EE. Supplementary Table 2. Targeted sequencing data from control and DEN-treated mouse EE. Supplementary Table 3. dN/dS results from DEN-treated and control mouse EE. Supplementary Table 4. Targeted sequencing data from human normal EE. Supplementary Table 5. dN/dS results from human normal EE. Supplementary Table 6. YFP clone areas in control and DEN-treated mouse EE. Supplementary Table 7. HGFP basal cell intensity in control and DEN-treated mouse EE. Supplementary Table 8. Percentage of EdU-positive cells in control and DEN-treated mouse EE. Supplementary Table 9. Area of confetti clones from DEN-treated EE. Supplementary Table 10. Code IDs for the clone triplicates and their matching controls. Supplementary Table 11. WES results of isolated clones from DEN-treated mouse EE. Supplementary Table 12. dN/dS results of isolated clones from DEN-treated mouse EE. Supplementary Table 13. Area of EE covered by DN-Maml1 clones in control and DEN-treated mice. Supplementary Table 14. YFP or DN-Maml1 clone sizes in DEN-treated mice. Supplementary Table 15. Size of confetti clones located at the edge or enclosed within DN-Maml1 areas.
Supplementary Video 1
3D structures of the highly mutated regions of the ligand-binding domain of NOTCH1 (light gray) bound to JAGGED1 (dark gray) (PDB 5UK5). Colored circles represent recurrently mutated codons within the NOTCH1 ligand-binding domain as follows: cysteine residues in disulfide bonds (blue), leucine to proline in β-sheets (orange), mutations affecting D469 (cyan), mutations of calcium-binding residues (red) and mutations on the ligand-binding interface (green).
Supplementary Video 2
3D structures of the NRR domain and cleavage site for NOTCH1 after ligand binding (PDB 3ETO). Blue colored circles represent recurrently mutated cysteine residues in disulfide bonds within the NOTCH1 NRR domains.
Supplementary Video 3
In silico simulations of competing wild-type (top) and mixed wild-type and mutant (bottom) clones over time, mimicking in vivo control and DEN treatment scenarios, respectively. Simulations and labeling of clones (middle column) start right after the treatment. In the first column clones are color coded according to their fitness, which is randomly assigned to each generated mutant clone and ranges from dark blue (lower) to dark red (higher fitness than wild-type cells that are in green). Note that only clones with the highest fitness prevail until the simulation ends, by outcompeting their surrounding ‘weaker’ clones for the limited space. The middle column shows the time course of randomly ‘labeled’ clones (yellow), simulating the in vivo lineage tracing experiments in Fig. 3c. Columns on the right show all the clones within the simulation, with colors assigned randomly. The videos are representative from at least five independent simulations run on a 300 × 300 lattice under the same parameter conditions (see Supplementary Note).
Supplementary Video 4
Supplementary Video 4 In silico simulations of the expansion of highly competitive single mutant clones (Mml+ cells) induced within a wild-type environment (top) or within a mutated landscape (bottom), equivalent to that generated following in vivo DEN treatment. Labeling of Mml+ mutant clones (green, middle column) starts after the treatments. In the first column clones are color coded according to fitness values, which were randomly assigned for mutagen-induced mutant clones (blue: low fitness; red: high); intermediate (green) and high (dark red) values were assigned to wild-type and Mml+ clones, respectively. The middle column show only the Mml+ ‘labeled’ clones (green), simulating the in vivo lineage tracing experiments in Fig. 6d. Columns on the right show all the clones within the simulation, with colors assigned randomly. The videos are representative from at least five independent simulations run on a 300 × 300 lattice under the same parameter conditions (see Supplementary Note).
Supplementary Video 5
Supplementary Video 5 In silico simulation to analyze the expanding behavior of subclones enclosed within or at the edges of mutant clones (see Fig. 7b,c). The movie on the left shows the expansion of highly competitive single mutant clones (Mml+, green) induced within a wild-type environment. The movie in the middle shows the random labeling of subclones (in red or yellow) in the same setup. The movie on the right shows the combination of the previous two. The video is representative from at least five independent simulations run on a 300 × 300 lattice under the same parameter conditions (see Supplementary Note).
Supplementary Video 6
In silico simulation using NetLogo. The movie shows a different on-lattice model of competition between neighbors. Following the induction of a subset of clones with a fitness advantage, wild-type cells (pink) are outcompeted by the fit (non-pink) population. Fit cells are initially colored green, and after a short delay a subset of these are colored cyan or yellow depending on whether they neighbor wild-type cells or not (‘boundary’ versus ‘core’ cells, respectively). The clone size distribution starts as a straight line, kinks over time, before becoming straight after the fitter clones take over the tissue. The population of yellow cells stays roughly stable, whilst the population of cyan cells grows over time.
Supplementary Video 7
In silico simulation of an off-lattice model using HAL. The movie shows that an off-lattice implementation of neighbor competition produces similar results to the on-lattice models. In the example shown, the mutant cells (green) have a larger bias towards dividing than the wild-type cells (black). Edge (red) or core (blue) subclones are induced in the mutant clones and show that, on average, the edge subclones grow larger. We ran simulations for three different mechanisms of mutant advantage and all three showed similar results (Extended Data Fig. 7g).
Rights and permissions
About this article
Cite this article
Colom, B., Alcolea, M.P., Piedrafita, G. et al. Spatial competition shapes the dynamic mutational landscape of normal esophageal epithelium. Nat Genet 52, 604–614 (2020). https://doi.org/10.1038/s41588-020-0624-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-0624-3
This article is cited by
-
Genetic variation across and within individuals
Nature Reviews Genetics (2024)
-
Spatial architectures of somatic mutations in normal prostate, benign prostatic hyperplasia and coexisting prostate cancer
Experimental & Molecular Medicine (2024)
-
Cell competition in development, homeostasis and cancer
Nature Reviews Molecular Cell Biology (2023)
-
Somatic mutations in facial skin from countries of contrasting skin cancer risk
Nature Genetics (2023)
-
Commonalities and differences in the mutational signature and somatic driver mutation landscape across solid and hollow viscus organs
Oncogene (2023)