Abstract
Genome-wide association studies have discovered numerous genomic loci associated with Alzheimer’s disease (AD); yet the causal genes and variants are incompletely identified. We performed an updated genome-wide AD meta-analysis, which identified 37 risk loci, including new associations near CCDC6, TSPAN14, NCK2 and SPRED2. Using three SNP-level fine-mapping methods, we identified 21 SNPs with >50% probability each of being causally involved in AD risk and others strongly suggested by functional annotation. We followed this with colocalization analyses across 109 gene expression quantitative trait loci datasets and prioritization of genes by using protein interaction networks and tissue-specific expression. Combining this information into a quantitative score, we found that evidence converged on likely causal genes, including the above four genes, and those at previously discovered AD loci, including BIN1, APH1B, PTK2B, PILRA and CASS4.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Summary statistics from the meta-analysis are available through the National Human Genome Research Institute-European Bioinformatics Institute GWAS catalog under accession nos. GCST90012877 and GCST90012878 (https://www.ebi.ac.uk/gwas/downloads/summary-statistics). The eQTL Catalogue is available at http://www.ebi.ac.uk/eqtl/. The GTEx Portal is available at https://www.gtexportal.org/home/. The Roadmap Epigenomics is available at http://www.roadmapepigenomics.org/. DeepSEA is available at http://deepsea.princeton.edu/job/analysis/create/. SpliceAI is available at https://github.com/Illumina/SpliceAI. FANTOM5 enhancers are available at https://fantom.gsc.riken.jp/5/data/. GERP is available at hgdownload.cse.ucsc.edu/gbdb/hg19/bbi/All_hg19_RS.bw. PhyloP is available at hgdownload.cse.ucsc.edu/goldenpath/hg19/phyloP100way. PhastCons is available at hgdownload.cse.ucsc.edu/goldenpath/hg19/phastCons100way. The brain eQTL meta-analysis summary statistics are avilable at https://www.synapse.org/#!Synapse:syn16984815. Primary microglia eQTL summary statistics are available under EGA accession no. EGAD00001005736. Primary microglia ATAC-seq data are available under dbGaP accession no. phs001373.v1.p1. The Allen Brain Institute is available at http://portal.brain-map.org/atlases-and-data/rnaseq. The IntAct molecular interaction database is available at https://www.ebi.ac.uk/intact/. The BioGRID database is available at https://thebiogrid.org/. The STRING database is available at https://string-db.org/.
Code availability
The code for the analyses described in this article can be found at https://github.com/jeremy37/AD_finemap.
Change history
26 February 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41588-021-00822-1
References
Liu, J. Z., Erlich, Y. & Pickrell, J. K. Case-control association mapping by proxy using family history of disease. Nat. Genet. 49, 325–331 (2017).
Marioni, R. E. et al. GWAS on family history of Alzheimer’s disease. Transl. Psychiatry 8, 99 (2018).
Jansen, I. E. et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 51, 404–413 (2019).
Claussnitzer, M. et al. FTO obesity variant circuitry and adipocyte browning in humans. N. Engl. J. Med. 373, 895–907 (2015).
Sekar, A. et al. Schizophrenia risk from complex variation of complement component 4. Nature 530, 177–183 (2016).
Huang, H. et al. Fine-mapping inflammatory bowel disease loci to single-variant resolution. Nature 547, 173–178 (2017).
Malik, M. et al. CD33 Alzheimer’s risk-altering polymorphism, CD33 expression, and exon 2 splicing. J. Neurosci. 33, 13320–13325 (2013).
Guerreiro, R. et al. TREM2 variants in Alzheimer’s disease. N. Engl. J. Med. 368, 117–127 (2013).
Sims, R. et al. Rare coding variants in PLCG2, ABI3, and TREM2 implicate microglial-mediated innate immunity in Alzheimer’s disease. Nat. Genet. 49, 1373–1384 (2017).
Kerimov, N. et al. eQTL Catalogue: a compendium of uniformly processed human gene expression and splicing QTLs. Preprint at bioRxiv https://doi.org/10.1101/2020.01.29.924266 (2020).
GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Kunkle, B. W. et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat. Genet. 51, 414–430 (2019).
Lambert, J. C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
Young, A. M. H. et al. A map of transcriptional heterogeneity and regulatory variation in human microglia. Preprint at bioRxiv https://doi.org/10.1101/2019.12.20.874099 (2019).
Leung, Y. Y. et al. Identifying amyloid pathology-related cerebrospinal fluid biomarkers for Alzheimer’s disease in a multicohort study. Alzheimers Dement. (Amst) 1, 339–348 (2015).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Moreno-Grau, S. et al. Genome-wide association analysis of dementia and its clinical endophenotypes reveal novel loci associated with Alzheimer’s disease and three causality networks: the GR@ACE project. Alzheimers Dement. 15, 1333–1347 (2019).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Sieberts, S. K. et al. Large eQTL meta-analysis reveals differing patterns between cerebral cortical and cerebellar brain regions. Sci. Data 7, 340 (2020).
Ng, B. et al. An xQTL map integrates the genetic architecture of the human brain’s transcriptome and epigenome. Nat. Neurosci. 20, 1418–1426 (2017).
Schmiedel, B. J. et al. Impact of genetic polymorphisms on human immune cell gene expression. Cell 175, 1701–1715.e16 (2018).
Jaffe, A. E. et al. Developmental and genetic regulation of the human cortex transcriptome illuminate schizophrenia pathogenesis. Nat. Neurosci. 21, 1117–1125 (2018).
Buil, A. et al. Gene-gene and gene-environment interactions detected by transcriptome sequence analysis in twins. Nat. Genet. 47, 88–91 (2015).
Fairfax, B. P. et al. Genetics of gene expression in primary immune cells identifies cell type-specific master regulators and roles of HLA alleles. Nat. Genet. 44, 502–510 (2012).
Fairfax, B. P. et al. Innate immune activity conditions the effect of regulatory variants upon monocyte gene expression. Science 343, 1246949 (2014).
Naranbhai, V. et al. Genomic modulators of gene expression in human neutrophils. Nat. Commun. 6, 7545 (2015).
Chen, L. et al. Genetic drivers of epigenetic and transcriptional variation in human immune cells. Cell 167, 1398–1414.e24 (2016).
Alasoo, K. et al. Shared genetic effects on chromatin and gene expression indicate a role for enhancer priming in immune response. Nat. Genet. 50, 424–431 (2018).
Gutierrez-Arcelus, M. et al. Passive and active DNA methylation and the interplay with genetic variation in gene regulation. eLife 2, e00523 (2013).
Lappalainen, T. et al. Transcriptome and genome sequencing uncovers functional variation in humans. Nature 501, 506–511 (2013).
Kilpinen, H. et al. Common genetic variation drives molecular heterogeneity in human iPSCs. Nature 546, 370–375 (2017).
Nédélec, Y. et al. Genetic ancestry and natural selection drive population differences in immune responses to pathogens. Cell 167, 657–669.e21 (2016).
Quach, H. et al. Genetic adaptation and Neandertal admixture shaped the immune system of human populations. Cell 167, 643–656.e17 (2016).
Schwartzentruber, J. et al. Molecular and functional variation in iPSC-derived sensory neurons. Nat. Genet. 50, 54–61 (2018).
van de Bunt, M. et al. Transcript expression data from human islets links regulatory signals from genome-wide association studies for type 2 diabetes and glycemic traits to their downstream effectors. PLoS Genet. 11, e1005694 (2015).
Momozawa, Y. et al. IBD risk loci are enriched in multigenic regulatory modules encompassing putative causative genes. Nat. Commun. 9, 2427 (2018).
Chun, S. et al. Limited statistical evidence for shared genetic effects of eQTLs and autoimmune-disease-associated loci in three major immune-cell types. Nat. Genet. 49, 600–605 (2017).
Salazar, S. V. et al. Alzheimer’s disease risk factor Pyk2 mediates amyloid-β-induced synaptic dysfunction and loss. J. Neurosci. 39, 758–772 (2019).
Raj, T. et al. Integrative transcriptome analyses of the aging brain implicate altered splicing in Alzheimer’s disease susceptibility. Nat. Genet. 50, 1584–1592 (2018).
Calafate, S., Flavin, W., Verstreken, P. & Moechars, D. Loss of Bin1 promotes the propagation of Tau pathology. Cell Rep. 17, 931–940 (2016).
Nott, A. et al. Brain cell type-specific enhancer–promoter interactome maps and disease-risk association. Science 366, 1134–1139 (2019).
Rathore, N. et al. Paired immunoglobulin-like type 2 receptor alpha G78R variant alters ligand binding and confers protection to Alzheimer’s disease. PLoS Genet. 14, e1007427 (2018).
Chan, G. et al. CD33 modulates TREM2: convergence of Alzheimer loci. Nat. Neurosci. 18, 1556–1558 (2015).
Raj, T. et al. CD33: increased inclusion of exon 2 implicates the Ig V-set domain in Alzheimer’s disease susceptibility. Hum. Mol. Genet. 23, 2729–2736 (2014).
Jonsson, T. et al. Variant of TREM2 associated with the risk of Alzheimer’s disease. N. Engl. J. Med. 368, 107–116 (2013).
Claes, C. et al. Human stem cell-derived monocytes and microglia-like cells reveal impaired amyloid plaque clearance upon heterozygous or homozygous loss of TREM2. Alzheimers Dement. 15, 453–464 (2019).
Aerts, J. et al. Bayesian refinement of association signals for 14 loci in 3 common diseases. Nat. Genet. 44, 1294–1301 (2012).
Benner, C. et al. FINEMAP: efficient variable selection using summary data from genome-wide association studies. Bioinformatics 32, 1493–1501 (2016).
Kichaev, G. et al. Integrating functional data to prioritize causal variants in statistical fine-mapping studies. PLoS Genet. 10, e1004722 (2014).
Benner, C. et al. Prospects of fine-mapping trait-associated genomic regions by using summary statistics from genome-wide association studies. Am. J. Hum. Genet. 101, 539–551 (2017).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science 356, eaal3222 (2017).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
McLaren, W. et al. The Ensembl Variant Effect Predictor. Genome Biol. 17, 122 (2016).
Davydov, E. V. et al. Identifying a high fraction of the human genome to be under selective constraint using GERP++. PLoS Comput. Biol. 6, e1001025 (2010).
Zhou, J. & Troyanskaya, O. G. Predicting effects of noncoding variants with deep learning-based sequence model. Nat. Methods 12, 931–934 (2015).
Jaganathan, K. et al. Predicting splicing from primary sequence with deep learning. Cell 176, 535–548.e24 (2019).
Steinberg, S. et al. Loss-of-function variants in ABCA7 confer risk of Alzheimer’s disease. Nat. Genet. 47, 445–447 (2015).
De Roeck, A. et al. An intronic VNTR affects splicing of ABCA7 and increases risk of Alzheimer’s disease. Acta Neuropathol. 135, 827–837 (2018).
Canela-Xandri, O., Rawlik, K. & Tenesa, A. An atlas of genetic associations in UK Biobank. Nat. Genet. 50, 1593–1599 (2018).
Liu, J. Z. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 47, 979–986 (2015).
Lanoiselée, H.-M. et al. APP, PSEN1, and PSEN2 mutations in early-onset Alzheimer disease: a genetic screening study of familial and sporadic cases. PLoS Med. 14, e1002270 (2017).
Fang, H. et al. A genetics-led approach defines the drug target landscape of 30 immune-related traits. Nat. Genet. 51, 1082–1091 (2019).
Amin, L. & Harris, D. A. Aβ receptors specifically recognize molecular features displayed by fibril ends and neurotoxic oligomers. Preprint at bioRxiv https://doi.org/10.1101/822361 (2019).
Nordestgaard, L. T., Tybjærg-Hansen, A., Nordestgaard, B. G. & Frikke-Schmidt, R. Loss-of-function mutation in ABCA1 and risk of Alzheimer’s disease and cerebrovascular disease. Alzheimers Dement. 11, 1430–1438 (2015).
Pickrell, J. K. Joint analysis of functional genomic data and genome-wide association studies of 18 human traits. Am. J. Hum. Genet. 94, 559–573 (2014).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Bakken, T. E. et al. Evolution of cellular diversity in primary motor cortex of human, marmoset monkey, and mouse. Preprint at bioRxiv https://doi.org/10.1101/2020.03.31.016972 (2020).
Mukherjee, S., Klaus, C., Pricop-Jeckstadt, M., Miller, J. A. & Struebing, F. L. A microglial signature directing human aging and neurodegeneration-related gene networks. Front. Neurosci. 13, 2 (2019).
Zhang, B. et al. Integrated systems approach identifies genetic nodes and networks in late-onset Alzheimer’s disease. Cell 153, 707–720 (2013).
Patel, K. R. et al. Single cell-type integrative network modeling identified novel microglial-specific targets for the phagosome in Alzheimer’s disease. Preprint at bioRxiv https://doi.org/10.1101/2020.06.09.143529 (2020).
Novikova, G. et al. Integration of Alzheimer’s disease genetics and myeloid genomics reveals novel disease risk mechanisms. Preprint at bioRxiv https://doi.org/10.1101/694281 (2019).
Biundo, F., Ishiwari, K., Del Prete, D. & D’Adamio, L. Deletion of the γ-secretase subunits Aph1B/C impairs memory and worsens the deficits of knock-in mice modeling the Alzheimer-like familial Danish dementia. Oncotarget 7, 11923–11944 (2016).
Nicolas, G. et al. Somatic variants in autosomal dominant genes are a rare cause of sporadic Alzheimer’s disease. Alzheimers Dement. 14, 1632–1639 (2018).
Zhang, X. et al. Negative evidence for a role of APH1B T27I variant in Alzheimer’s disease. Hum. Mol. Genet. 29, 955–966 (2020).
Matthews, A. L. et al. Regulation of leukocytes by TspanC8 tetraspanins and the ‘molecular scissor’ ADAM10. Front. Immunol. 9, 1451 (2018).
Schlepckow, K. et al. An Alzheimer‐associated TREM2 variant occurs at the ADAM cleavage site and affects shedding and phagocytic function. EMBO Mol. Med. 9, 1356–1365 (2017).
Juul Rasmussen, I., Tybjærg-Hansen, A., Rasmussen, K. L., Nordestgaard, B. G. & Frikke-Schmidt, R. Blood–brain barrier transcytosis genes, risk of dementia and stroke: a prospective cohort study of 74,754 individuals. Eur. J. Epidemiol. 34, 579–590 (2019).
Zhao, J. et al. Rare 3-O-sulfation of heparan sulfate enhances Tau interaction and cellular uptake. Angew. Chem. Int. Ed. Engl. 59, 1818–1827 (2020).
Jun, G. R. et al. Transethnic genome-wide scan identifies novel Alzheimer’s disease loci. Alzheimers Dement. 13, 727–738 (2017).
Andersen, O. M., Rudolph, I.-M. & Willnow, T. E. Risk factor SORL1: from genetic association to functional validation in Alzheimer’s disease. Acta Neuropathol. 132, 653–665 (2016).
Sassi, C. et al. Influence of coding variability in APP-Aβ metabolism genes in sporadic Alzheimer’s disease. PLoS ONE 11, e0150079 (2016).
Lu, Q. et al. Systematic tissue-specific functional annotation of the human genome highlights immune-related DNA elements for late-onset Alzheimer’s disease. PLoS Genet. 13, e1006933 (2017).
Ghanbari, M. et al. A functional variant in the miR‐142 promoter modulating its expression and conferring risk of Alzheimer disease. Hum. Mutat. 40, 2131–2145 (2019).
Chung, C.-M. et al. Fine-mapping angiotensin-converting enzyme gene: separate QTLs identified for hypertension and for ACE activity. PLoS ONE 8, e56119 (2013).
Nylocks, K. M. et al. An angiotensin-converting enzyme (ACE) polymorphism may mitigate the effects of angiotensin-pathway medications on posttraumatic stress symptoms. Am. J. Med. Genet. B Neuropsychiatr. Genet. 168B, 307–315 (2015).
Kamboh, M. I. et al. Genome-wide association study of Alzheimer’s disease. Transl. Psychiatry 2, e117 (2012).
Bernstein, A. I. et al. 5-Hydroxymethylation-associated epigenetic modifiers of Alzheimer’s disease modulate Tau-induced neurotoxicity. Hum. Mol. Genet. 25, 2437–2450 (2016).
Witoelar, A. et al. Meta-analysis of Alzheimer’s disease on 9,751 samples from Norway and IGAP study identifies four risk loci. Sci. Rep. 8, 18088 (2018).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Bellenguez, C. et al. A robust clustering algorithm for identifying problematic samples in genome-wide association studies. Bioinformatics 28, 134–135 (2012).
Manichaikul, A. et al. Robust relationship inference in genome-wide association studies. Bioinformatics 26, 2867–2873 (2010).
Loh, P.-R. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Pirinen, M., Donnelly, P. & Spencer, C. C. A. Efficient computation with a linear mixed model on large-scale data sets with applications to genetic studies. Ann. Appl. Stat. 7, 369–390 (2013).
Zhao, H. et al. CrossMap: a versatile tool for coordinate conversion between genome assemblies. Bioinformatics 30, 1006–1007 (2014).
Wakefield, J. Bayes factors for genome-wide association studies: comparison with P-values. Genet. Epidemiol. 33, 79–86 (2009).
Kumasaka, N., Knights, A. J. & Gaffney, D. J. Fine-mapping cellular QTLs with RASQUAL and ATAC-seq. Nat. Genet. 48, 206–213 (2016).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
Zhang, Y. et al. Model-based analysis of ChIP–Seq (MACS). Genome Biol. 9, R137 (2008).
Alasoo, K. wiggleplotr: Make read coverage plots from BigWig files. R package version 1.10.1 https://bioconductor.org/packages/release/bioc/html/wiggleplotr.html (2019).
Orchard, S. et al. The MIntAct project: IntAct as a common curation platform for 11 molecular interaction databases. Nucleic Acids Res. 42, D358–D363 (2014).
Chatr-Aryamontri, A. et al. The BioGRID interaction database: 2017 update. Nucleic Acids Res. 45, D369–D379 (2017).
Szklarczyk, D. et al. The STRING database in 2017: quality-controlled protein–protein association networks, made broadly accessible. Nucleic Acids Res. 45, D362–D368 (2017).
Fogaras, D., Rácz, B., Csalogány, K. & Sarlós, T. Towards scaling fully personalized PageRank: algorithms, lower bounds, and experiments. Internet Math. 2, 333–358 (2005).
Csardi, G., Nepusz, T. The igraph software package for complex network research. InterJ. Complex Syst. 1695, 1–9 (2006).
Raudvere, U. et al. g:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 47, W191–W198 (2019).
Battle, A. et al. Characterizing the genetic basis of transcriptome diversity through RNA-sequencing of 922 individuals. Genome Res. 24, 14–24 (2014).
Barbeira, A. N. et al. Exploiting the GTEx resources to decipher the mechanisms at GWAS loci. Preprint at bioRxiv https://doi.org/10.1101/814350 (2019).
Stacey, D. et al. ProGeM: a framework for the prioritization of candidate causal genes at molecular quantitative trait loci. Nucleic Acids Res. 47, e3 (2019).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Acknowledgements
This work was funded by Open Targets (OTAR037). We thank J. Barrett for guidance during the initiation of the project and K. Alasoo for early access to the eQTL Catalogue. We thank A. Ruiz for support in using the summary results from the GR@ACE study. We thank the participants and investigators of the FinnGen study and UK Biobank. R.J.M.F. is supported by grants from the UK Multiple Sclerosis Society (MS 50), the Adelson Medical Research Foundation and a core support grant from the Wellcome Trust and Medical Research Council (MRC) to the Wellcome-MRC Cambridge Stem Cell Institute (no. 203151/Z/16/Z). A.M.H.Y. is supported by a Wellcome Trust PhD for Clinicians fellowship.
Author information
Authors and Affiliations
Contributions
J.S. planned and conducted the analyses and wrote the paper. J.Z.L. performed the GWAX and meta-analysis. S.C. and E.B. assisted with fine-mapping, variant and gene prioritization. I.B.-H. and P.B. performed and supervised the gene network analysis. R.J.M.F. designed and A.M.H.Y. performed the isolation of human microglia from brain biopsies. N.K. performed the microglia eQTL mapping. A.B., T.J., D.J.G. and K.E. conceived and supervised the study.
Corresponding authors
Ethics declarations
Competing interests
J.Z.L. was an employee of Biogen at the time of the study and is now an employee of GSK. D.J.G. is an employee of Genomics PLC. T.J. is an employee of GSK. K.E. is an employee of BioMarin Pharmaceutical.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
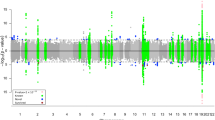
Extended Data Fig. 1 Association of AD loci in discovery + replication (‘global’) meta-analysis.
Association of AD loci in discovery + replication dataset (‘global’) meta-analysis. For most loci, association significance is increased in the global meta-analysis (blue bars) relative to the discovery analysis (grey bars). The dashed vertical line shows P = 5 × 10−8. P-values were computed by inverse variance weighted meta-analysis, and bars show the -log10(P) for the SNP with minimum P value at the locus in either the discovery or global meta-analysis.
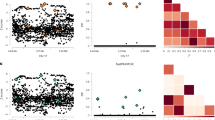
Extended Data Fig. 2 Comparison of fine-mapping in the meta-analysis vs. Kunkle et al.
Comparison of fine-mapping in the meta-analysis vs. Kunkle et al. Scatterplots showing, for each locus, SNP probabilities from FINEMAP applied to either the Kunkle et al. + UK Biobank meta-analysis (x-axis), or to only Kunkle et al. The number of causal variants at each locus was set to the number detected by GCTA in the meta-analysis. For most of the 36 loci, SNP probabilities are well correlated. For a few loci that are well powered in Kunkle et al., this is not the case, namely ABCA7, EPHA1, ECHDC3, and HLA. For these loci, fine-mapping results should be interpreted with caution. Six other loci are not well correlated (ADAMTS4, APH1B, IKZF1, PLCG2, TMEM163, and VKORC1), but these loci are poorly powered in Kunkle et al. (lead P values 2.1 × 10−6 to 2.1 × 10−3).
Extended Data Fig. 3 Network enrichment.
a, The Pagerank percentile of all genes (within 500 kb) at each AD GWAS locus containing a seed gene is shown, with seed genes highlighted in blue. b, A violin/boxplot shows that seed genes have a markedly higher network Pagerank percentile than remaining genes (P = 2.4 × 10−9, one-tailed Wilcoxon rank sum test). c, Log odds ratio enrichment of AD risk among SNPs nearest to genes with network Pagerank percentile in different bins, determined using fgwas (whiskers represent 95% confidence intervals).
Extended Data Fig. 4 Gene expression enrichments.
Expression enrichments for GTEx + microglia. Shown are the log odds ratio enrichments of AD risk among SNPs with relative gene expression in each tissue above the 80th (or 90th) percentile across tissues. Whiskers represent 95% confidence intervals determined by fgwas.
Extended Data Fig. 5 Colocalization scores.
a, Genes with maximum colocalization H4 probability >0.9 have higher Pagerank percentile (left boxplot) and higher total score (sum of the four non-coloc predictors, right boxplot) than do genes without colocalisation (<0.5). Genes with intermediate colocalisation evidence (bins 0.5 - 0.8 and 0.8 - 0.9) show little evidence of having higher scores by the other metrics. Based on this, we chose a maxColoc probability of 0.9 as the lower bound for our colocalization score. b, Boxplot of the total score (excluding coloc) for genes that have a colocalisation probability > 0.9 in at least one QTL dataset within each tissue group. The most significant difference is between totalScore for genes with microglial colocalizations vs. the genes with colocalization in ‘other’ tissues (non-immune GTEx tissues), but the difference is weak (P = 0.041, Wilcoxon rank sum test). In all cases, boxplots show the 25th, median, and 75th percentile of the distribution, with whiskers extending to the largest (and smallest) value no further than 1.5 times the interquartile range from the boxplot hinge.
Extended Data Fig. 6 Gene distance score.
The distance score assigned to genes near an AD GWAS peak, which decreases approximately linearly (past a distance of 1 kb) with increasing log-scaled distance up to 500 kb.
Supplementary information
Supplementary Information
Supplementary Note
Supplementary Tables
Supplementary Tables 1–14
Rights and permissions
About this article
Cite this article
Schwartzentruber, J., Cooper, S., Liu, J.Z. et al. Genome-wide meta-analysis, fine-mapping and integrative prioritization implicate new Alzheimer’s disease risk genes. Nat Genet 53, 392–402 (2021). https://doi.org/10.1038/s41588-020-00776-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-00776-w
This article is cited by
-
Unraveling the intercellular communication disruption and key pathways in Alzheimer’s disease: an integrative study of single-nucleus transcriptomes and genetic association
Alzheimer's Research & Therapy (2024)
-
Association of long-term exposure to various ambient air pollutants, lifestyle, and genetic predisposition with incident cognitive impairment and dementia
BMC Public Health (2024)
-
Deciphering the effect of phytosterols on Alzheimer’s disease and Parkinson’s disease: the mediating role of lipid profiles
Alzheimer's Research & Therapy (2024)
-
Priority index for critical Covid-19 identifies clinically actionable targets and drugs
Communications Biology (2024)
-
Leveraging electronic health records and knowledge networks for Alzheimer’s disease prediction and sex-specific biological insights
Nature Aging (2024)