Abstract
Lung cancer is the world’s leading cause of cancer death and shows strong ancestry disparities. By sequencing and assembling a large genomic and transcriptomic dataset of lung adenocarcinoma (LUAD) in individuals of East Asian ancestry (EAS; n = 305), we found that East Asian LUADs had more stable genomes characterized by fewer mutations and fewer copy number alterations than LUADs from individuals of European ancestry. This difference is much stronger in smokers as compared to nonsmokers. Transcriptomic clustering identified a new EAS-specific LUAD subgroup with a less complex genomic profile and upregulated immune-related genes, allowing the possibility of immunotherapy-based approaches. Integrative analysis across clinical and molecular features showed the importance of molecular phenotypes in patient prognostic stratification. EAS LUADs had better prediction accuracy than those of European ancestry, potentially due to their less complex genomic architecture. This study elucidated a comprehensive genomic landscape of EAS LUADs and highlighted important ancestry differences between the two cohorts.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Raw sequencing data have been deposited in the European Genome-phenome Archive (EGA, http://www.ebi.ac.uk/ega/) under accession codes EGAD00001004421 and EGAD00001004422. All clinical records, somatic mutations, copy number variations and histological images from our study are hosted in OncoSG (https://src.gisapps.org/OncoSG/) under dataset ‘Lung Adenocarcinoma (GIS, 2019)’ which is publicly available (Supplementary Note).
References
Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68, 394–424 (2018).
Cheng, T.-Y. D. et al. The international epidemiology of lung cancer: latest trends, disparities, and tumor characteristics. J. Thorac. Oncol. 11, 1653–1671 (2016).
Imielinski, M. et al. Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell 150, 1107–1120 (2012).
The Cancer Genome Atlas Research Network. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Campbell, J. D. et al. Distinct patterns of somatic genome alterations in lung adenocarcinomas and squamous cell carcinomas. Nat. Genet. 48, 607–616 (2016).
Kadara, H. et al. Whole-exome sequencing and immune profiling of early-stage lung adenocarcinoma with fully annotated clinical follow-up. Ann. Oncol. 28, 75–82 (2017).
Garon, E. B. et al. Pembrolizumab for the treatment of non-small-cell lung cancer. N. Engl. J. Med. 372, 2018–2028 (2015).
Borghaei, H. et al. Nivolumab versus docetaxel in advanced nonsquamous non–small-cell lung cancer. N. Engl. J. Med. 373, 1627–1639 (2015).
Fehrenbacher, L. et al. Updated efficacy analysis including secondary population results for OAK: a randomized phase III study of Atezolizumab versus Docetaxel in patients with previously treated advanced non-small cell lung cancer. J. Thorac. Oncol. 13, 1156–1170 (2018).
Ayers, M. et al. IFN-γ-related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest. 127, 2930–2940 (2017).
Cristescu, R. et al. Pan-tumor genomic biomarkers for PD-1 checkpoint blockade-based immunotherapy. Science 362, eaar3593 (2018).
Tan, W.-L. et al. Novel therapeutic targets on the horizon for lung cancer. Lancet Oncol. 17, e347–e362 (2016).
Shigematsu, H. et al. Clinical and biological features associated with epidermal growth factor receptor gene mutations in lung cancers. J. Natl. Cancer Inst. 97, 339–346 (2005).
Jain, A. et al. Impact of smoking and brain metastasis on outcomes of advanced EGFR mutation lung adenocarcinoma patients treated with first line epidermal growth factor receptor tyrosine kinase inhibitors. PloS ONE 10, e0123587 (2015).
Kris, M. G. et al. Using multiplexed assays of oncogenic drivers in lung cancers to select targeted drugs. JAMA 311, 1998–2006 (2014).
Clinical Lung Cancer Genome Project (CLCGP) & Network Genomic Medicine (NGM). A genomics-based classification of human lung tumors. Sci. Transl. Med. 5, 209ra153 (2013).
Wu, K. et al. Frequent alterations in cytoskeleton remodelling genes in primary and metastatic lung adenocarcinomas. Nat. Commun. 6, 10131 (2015).
Wang, C. et al. Whole-genome sequencing reveals genomic signatures associated with the inflammatory microenvironments in Chinese NSCLC patients. Nat. Commun. 9, 2054 (2018).
Luo, W. et al. Characteristics of genomic alterations of lung adenocarcinoma in young never-smokers. Int. J. Cancer 143, 1696–1705 (2018).
Nahar, R. et al. Elucidating the genomic architecture of Asian EGFR-mutant lung adenocarcinoma through multi-region exome sequencing. Nat. Commun. 9, 216 (2018).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Ramos, A. H. et al. Oncotator: cancer variant annotation tool. Hum. Mutat. 36, E2423–E2429 (2015).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Saunders, C. T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor–normal sample pairs. Bioinformatics 28, 1811–1817 (2012).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Tokheim, C. J., Papadopoulos, N., Kinzler, K. W., Vogelstein, B. & Karchin, R. Evaluating the evaluation of cancer driver genes. Proc. Natl Acad. Sci. USA 113, 14330–14335 (2016).
Nicorici, D. et al. FusionCatcher—a tool for finding somatic fusion genes in paired-end RNA-sequencing data. Preprint at bioRxiv https://doi.org/10.1101/011650 (2014).
Liu, L. et al. Comprehensive genomic profiling of lung cancer using a validated panel to explore therapeutic targets in East Asian patients. Cancer Sci. 108, 2487–2494 (2017).
Hu, X. et al. TumorFusions: an integrative resource for cancer-associated transcript fusions. Nucleic Acids Res. 46, D1144–D1149 (2018).
Weischenfeldt, J. et al. Integrative genomic analyses reveal an androgen-driven somatic alteration landscape in early-onset prostate cancer. Cancer Cell 23, 159–170 (2013).
Liao, S. et al. A genetic interaction analysis identifies cancer drivers that modify EGFR dependency. Genes Dev. 31, 184–196 (2017).
Gibson, B. A. & Kraus, W. L. New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat. Rev. Mol. Cell Biol. 13, 411–424 (2012).
Ikeda, Y. et al. Germline PARP4 mutations in patients with primary thyroid and breast cancers. Endocr. Relat. Cancer 23, 171–179 (2016).
Long, N. P. et al. Novel biomarker candidates for colorectal cancer metastasis: a meta-analysis of in vitro studies. Cancer Inform. 15, 11–17 (2016).
Katsyv, I. et al. EPRS is a critical regulator of cell proliferation and estrogen signaling in ER+ breast cancer. Oncotarget 7, 69592–69605 (2016).
Wang, J.-W. et al. Deregulated expression of LRBA facilitates cancer cell growth. Oncogene 23, 4089–4097 (2004).
Sung, H. et al. Inactivation of RASA1 promotes melanoma tumorigenesis via R-Ras activation. Oncotarget 7, 23885–23896 (2016).
Zhu, Y.-J., Xu, B. & Xia, W. Hsa-mir-182 downregulates RASA1 and suppresses lung squamous cell carcinoma cell proliferation. Clin. Lab. 60, 155–159 (2014).
Wang, W.-H., Studach, L. L. & Andrisani, O. M. Proteins ZNF198 and SUZ12 are down-regulated in hepatitis B virus (HBV) X protein-mediated hepatocyte transformation and in HBV replication. Hepatology 53, 1137–1147 (2011).
Tamborero, D. et al. Cancer genome interpreter annotates the biological and clinical relevance of tumor alterations. Genome Med. 10, 25 (2018).
Lee, Y. J. et al. Activating mutations within the EGFR kinase domain: a molecular predictor of disease-free survival in resected pulmonary adenocarcinoma. J. Cancer Res. Clin. Oncol. 135, 1647–1654 (2009).
D’Angelo, S. P. et al. Distinct clinical course of EGFR-mutant resected lung cancers: results of testing of 1118 surgical specimens and effects of adjuvant Gefitinib and Erlotinib. J. Thorac. Oncol. 7, 1815–1822 (2012).
Jeon, J. H. et al. Prognostic and predictive role of epidermal growth factor receptor mutation in recurrent pulmonary adenocarcinoma after curative resection. Eur. J. Cardiothorac. Surg. 47, 556–562 (2015).
Favero, F. et al. Sequenza: allele-specific copy number and mutation profiles from tumor sequencing data. Ann. Oncol. 26, 64–70 (2015).
Mermel, C. H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41 (2011).
Gaujoux, R. & Seoighe, C. A flexible R package for nonnegative matrix factorization. BMC Bioinformatics 11, 367 (2010).
Blokzijl, F., Janssen, R., van Boxtel, R. & Cuppen, E. MutationalPatterns: comprehensive genome-wide analysis of mutational processes. Genome Med. 10, 33 (2018).
McGranahan, N. et al. Clonal status of actionable driver events and the timing of mutational processes in cancer evolution. Sci. Transl. Med. 7, 283ra54 (2015).
Bruin, E. Cde et al. Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science 346, 251–256 (2014).
Haradhvala, N. J. et al. Mutational strand asymmetries in cancer genomes reveal mechanisms of DNA damage and repair. Cell 164, 538–549 (2016).
Hayes, D. N. et al. Gene expression profiling reveals reproducible human lung adenocarcinoma subtypes in multiple independent patient cohorts. J. Clin. Oncol. 24, 5079–5090 (2006).
Wilkerson, M. D. et al. Differential pathogenesis of lung adenocarcinoma subtypes involving sequence mutations, copy number, chromosomal instability, and methylation. PLoS ONE 7, e36530 (2012).
Hoshida, Y., Brunet, J.-P., Tamayo, P., Golub, T. R. & Mesirov, J. P. Subclass mapping: identifying common subtypes in independent disease data sets. PLoS ONE 2, e1195 (2007).
Nirmal, A. J. et al. Immune cell gene signatures for profiling the microenvironment of solid tumors. Cancer Immunol. Res. 6, 1388–1400 (2018).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. et al. The molecular signatures database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Sanchez-Vega, F. et al. Oncogenic signaling pathways in the cancer genome atlas. Cell 173, 321–337.e10 (2018).
Chakravarty, D. et al. OncoKB: A precision oncology knowledge base. JCO Precis. Oncol. https://doi.org/10.1200/PO.17.00011 (2017).
Wu, Y.-L. et al. A consensus on immunotherapy from the 2017 Chinese Lung Cancer Summit expert panel. Transl. Lung Cancer Res. 7, 436–436 (2018). 428-.
Morris, L. G. T. et al. Pan-cancer analysis of intratumor heterogeneity as a prognostic determinant of survival. Oncotarget 7, 10051–10063 (2016).
Yuan, Y. et al. Assessing the clinical utility of cancer genomic and proteomic data across tumor types. Nat. Biotechnol. 32, 644 (2014).
Jamal-Hanjani, M. et al. Tracking the evolution of non-small-cell lung cancer. N. Engl. J. Med. 376, 2109–2121 (2017).
Tan, D. S. W., Mok, T. S. K. & Rebbeck, T. R. Cancer genomics: diversity and disparity across ethnicity and geography. J. Clin. Oncol. 34, 91–101 (2016).
Haiman, C. A. et al. Genome-wide association study of prostate cancer in men of African ancestry identifies a susceptibility locus at 17q21. Nat. Genet. 43, 570–573 (2011).
Haiman, C. A. et al. A common variant at the TERT-CLPTM1L locus is associated with estrogen receptor-negative breast cancer. Nat. Genet. 43, 1210–1214 (2011).
Ziegler, R. G. et al. Migration patterns and breast cancer risk in Asian-American women. J. Natl Cancer Inst. 85, 1819–1827 (1993).
Kuniholm, M. H. et al. Prevalence of hepatitis C virus infection in US Hispanic/Latino adults: results from the NHANES 2007-2010 and HCHS/SOL studies. J. Infect. Dis. 209, 1585–1590 (2014).
Teng, A. M., Blakely, T., Baker, M. G. & Sarfati, D. The contribution of Helicobacter pylori to excess gastric cancer in Indigenous and Pacific men: a birth cohort estimate. Gastric Cancer 20, 752–755 (2017).
Haiman, C. A. et al. Ethnic and racial differences in the smoking-related risk of lung cancer. N. Engl. J. Med. 354, 333–342 (2006).
Wu, C. et al. Genetic variants on chromosome 15q25 associated with lung cancer risk in Chinese populations. Cancer Res. 69, 5065–5072 (2009).
Wang, J. et al. Genetic predisposition to lung cancer: comprehensive literature integration, meta-analysis, and multiple evidence assessment of candidate-gene association studies. Sci. Rep. 7, 8371 (2017).
Seow, A. et al. Fumes from meat cooking and lung cancer risk in Chinese women. Cancer Epidemiol. Biomarkers Prev. 9, 1215–1221 (2000).
Lee, T. & Gany, F. Cooking oil fumes and lung cancer: a review of the literature in the context of the U.S. population. J. Immigr. Minor. Health 15, 646–652 (2013).
Lortet-Tieulent, J. et al. Convergence of decreasing male and increasing female incidence rates in major tobacco-related cancers in Europe in 1988–2010. Eur. J. Cancer 51, 1144–1163 (2015).
Jemal, A. et al. Higher lung cancer incidence in young women than young men in the United States. N. Engl. J. Med. 378, 1999–2009 (2018).
Takeuchi, T. et al. Expression profile-defined classification of lung adenocarcinoma shows close relationship with underlying major genetic changes and clinicopathologic behaviors. J. Clin. Oncol. 24, 1679–1688 (2006).
Davoli, T., Uno, H., Wooten, E. C. & Elledge, S. J. Tumor aneuploidy correlates with markers of immune evasion and with reduced response to immunotherapy. Science 355, eaaf8399 (2017).
Taylor, A. M. et al. Genomic and functional approaches to understanding cancer aneuploidy. Cancer Cell 33, 676–689.e3 (2018).
Coudray, N. et al. Classification and mutation prediction from non-small cell lung cancer histopathology images using deep learning. Nat. Med. 24, 1559–1567 (2018).
Cann, H. M. et al. A human genome diversity cell line panel. Science 296, 261–262 (2002).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997v2 (2013).
Ding, L. et al. Somatic mutations affect key pathways in lung adenocarcinoma. Nature 455, 1069–1075 (2008).
Govindan, R. et al. Genomic landscape of non-small cell lung cancer in smokers and never-smokers. Cell 150, 1121–1134 (2012).
Lawrence, M. S. et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505, 495–501 (2014).
Kandoth, C. et al. Mutational landscape and significance across 12 major cancer types. Nature 502, 333–339 (2013).
Bailey, M. H. et al. Comprehensive characterization of cancer driver genes and mutations. Cell 173, 371–385.e18 (2018).
Tan, V. Y. F. & Févotte, C. Automatic relevance determination in nonnegative matrix factorization with the /spl beta/-divergence. IEEE Trans. Pattern Anal. Mach. Intell. 35, 1592–1605 (2013).
Kim, J. et al. Somatic ERCC2 mutations are associated with a distinct genomic signature in urothelial tumors. Nat. Genet. 48, 600–606 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Korotkevich, G., Sukhov, V. & Sergushichev, A. Fast gene set enrichment analysis. Preprint at bioRxiv https://doi.org/10.1101/060012 (2019).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Andor, N., Harness, J. V., Müller, S., Mewes, H. W. & Petritsch, C. EXPANDS: expanding ploidy and allele frequency on nested subpopulations. Bioinformatics 30, 50–60 (2014).
Merlo, L. M. F. et al. A comprehensive survey of clonal diversity measures in Barrett’s esophagus as biomarkers of progression to esophageal adenocarcinoma. Cancer Prev. Res. (Phila) 3, 1388–1397 (2010).
Mroz, E. A. & Rocco, J. W. MATH, a novel measure of intratumor genetic heterogeneity, is high in poor-outcome classes of head and neck squamous cell carcinoma. Oral Oncol. 49, 211–215 (2013).
Harrell, F. Regression Modeling Strategies: With Applications to Linear Models, Logistic Regression, and Survival Analysis (Springer, 2001).
Harrell, F. E., Lee, K. L. & Mark, D. B. Multivariable prognostic models: issues in developing models, evaluating assumptions and adequacy, and measuring and reducing errors. Stat. Med. 15, 361–387 (1996).
Schröder, M. S., Culhane, A. C., Quackenbush, J. & Haibe-Kains, B. survcomp: an R/Bioconductor package for performance assessment and comparison of survival models. Bioinformatics 27, 3206–3208 (2011).
Acknowledgements
This work was funded by Glaxo Wellcome Manufacturing Pte Ltd, the Agency for Science Technology and Research (A*STAR) (grant no. GIS/15-IAF100), the National Medical Research Council, Singapore (grant nos. NMRC/OFLCG/002c/2018, NMRC/OFIRG/0064/2017 and NMRC/TCR/007-NCC/2013), the National Research Foundation, Singapore (grant no. NRF-NRFF2015–04), German Cancer Aid (grant no. 70113510) and Lung Cancer Consortium Singapore (LCCS). LCCS is jointly supported by philanthropy (including Singapore Millennium Foundation), and institutional and industrial grants. W.Z. is supported in part by the National Key R&D program of China (grant nos. 2018YFC0910400 and 2018YFC1406902) and the National Science Foundation of China (grant no. 31970566). We thank Beijing Genomics Institute for providing the published sequencing data. We thank T. Zhang for contribution to genomic data analysis, C. T. J. Ong, Y. L. Lee, I. M. L. Chua and W. W. J. Soon for the next-generation sequencing work, the GIS Research Pipeline Development team for support with analysis pipelines and Y. Matsuoka for administrative support. We thank Y. Cun for helpful discussions.
Author information
Authors and Affiliations
Contributions
A.M.H., W.Z. and D.S.W.T. conceived the study, and B. Lim, W.L.T. and E.-H.T. contributed. A.M.H. coordinated the genomics work. D.S.W.T. coordinated the clinical work. J.C. and W.Z. coordinated work on data analysis. J.C. performed genomic data analysis with contributions from H.Y., C.Q.T., B. Lu, J.J.S.A., J.Q.L., F.G.S., R.N., Y.Y.L., C.Z.J.P., K.P.C., Y.F.L. and J.L. A.W. contributed to analysis pipeline development. A.S.M.T. and L.B.A. performed nucleic acid extraction, exome library preparations and fusion gene validation, with contributions from F.G.S. who also performed SNV validation. A.T., with assistance from Z.W.A. and T.K.H.L., performed sectoring and histology studies and led the pathological work. P.S.C. and P.Y.N. contributed to RNA-seq library preparations and sequencing. T.P.T.K., B.-H.O., D.A., A.A.L.H., A.G. and C.W.T. performed surgery and biopsy procedures. D.S.W.T., A.T., W.-T.L., C.K.T., L.W. and E.-H.T. coordinated patient tissue banking, specimen transfer and clinical data curation. P.J.C., M.M.C., J.J.S.A. and A.J.S. implemented the OncoSG data portal. L.S., Z.W.A. and J.P.S.Y. performed multiplex immunohistochemistry. J.C., W.Z., A.M.H., D.S.W.T., C.L.C. and E.-H.T. interpreted the data and conceptualized the manuscript. J.C. created figures with contributions from F.G.S., C.Q.T., J.J.S.A., S.M., K.P.C. and W.Z. J.C. and W.Z. wrote the manuscript, with contributions from D.S.W.T., A.M.H., C.L.C., S.M., A.S.M.T., J.J.S.A., H.Y., B. Lu, K.P.C. and E.-H.T.
Corresponding authors
Ethics declarations
Competing interests
A.M.H, D.S.W.T., and W.Z. received research funding from Glaxo Wellcome Manufacturing Pte Ltd.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
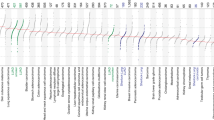
Extended Data Fig. 1 Kaplan-Meier plots of driver genes that can stratify patient survival outcome.
Survival outcomes of patients harbouring the driver mutation were compared against those did not in the (a) East Asian- ancestry (EAS, n=293) and (b) European-ancestry (EUR, n=225) cohort. Genes tested are from a curated list of LUAD drivers (Methods), and the plotted genes are those show significant coefficients (FDR<0.2 in EAS and p-values<0.05 in EUR, two-sided t-test) in a multivariate Cox model including stage, age, gender and smoker in any cohort. (c) Comparision of the survival outcomes of mutant or wildtype EGFR carriers among the EAS non-smokers (n=185) and smokers (n=110). EGFR mutant carriers showed better outcome, especially among non-smokers. Mut, mutant; WT, wildtype.
Extended Data Fig. 2 PCA of RNA profiles using both tumor and normal samples.
PCA of (a) East Asian-ancestry (EAS) and (b) European-ancestry (EUR) tumor (EAS, n=172; EUR, n=249) and normal (EAS, n=88; EUR, n=42) samples to illustrate the relationship of LUAD transcriptomic subtypes and the normal samples. In the two-group partitions, the TRU clusters were closer to the normal samples in both cohorts. In the three-group partitions, the TRU and TRU-I sub-clusters in the Asian were closer to the normal samples, while the TRU sub-cluster in the EUR was closer to the normal samples.
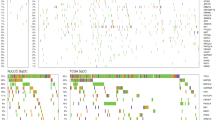
Extended Data Fig. 3 Phenotypes of the RNA sub-clusters in the EUR cohort.
The top two rows indicate the cluster assignment of the patients. The following rows show the normalized mean expression of GSEA enriched gene sets from the differential expressed genes between the TRU and non-TRU clusters and between the PI and PP sub-clusters, and the values of immune-related signatures. High values were shown in red and low in blue. The oncoprint plot shows major driver mutations across sub-clusters. The clinical and other genomic phenotypes are shown at the bottom.
Extended Data Fig. 4 Kaplan-Meier plots of the survival groups derived from genomic features only.
Using only genomic features (driver genes, molecular and ITH features), patients were divided evenly into three survival groups based on the predicted hazard from the multivariate Cox model. For both East Asian-ancestry (a) and European-ancestry (b) cohort, these survival models can clearly stratify patient survival outcome. They could stratify survival outcome even within early or late stage patients, indicating the prediction power of genomic features independent of clinical features. Statistical test used can be found under Methods section “Statistics and Reproducibility”.
Extended Data Fig. 5 Comparison of the prediction accuracies between the balanced EAS and EUR cohorts.
Related to Fig. 5c, box plots showing prediction accuracy calculated as Harrell’s concordance index (c-index) from the multivariate Cox models with different set of predictors. For fair comparisons across cohorts, the proportion of smokers were balanced by randomly down-sampling non-smokers in the East Asian-ancestry cohort and smokers in the European-ancestry cohort (a). To rule out the effect of EGFR mutation and possible TKI treatment on patient survival, the comparison was narrowed down to only patients with wildtyp EGFR (b). Statistical test used and the definition of boxplot elements can be found under Methods section “Statistics and Reproducibility”.
Extended Data Fig. 6 Summary of the ancestry differences.
A summary of major ancestry differences across the two cohorts in this study (top), and the differences seen when comparing among smokers and non-smokers (bottom). Red, higher/more; blue, lower/less; ≈, similar; ♂, male; ♀, female; IO, immuno-oncology; NA, not available.
Supplementary information
Supplementary Information
Supplementary Notes 1–7 and Figs. 1–41
Supplementary Tables
Supplementary Tables 1–12
Rights and permissions
About this article
Cite this article
Chen, J., Yang, H., Teo, A.S.M. et al. Genomic landscape of lung adenocarcinoma in East Asians. Nat Genet 52, 177–186 (2020). https://doi.org/10.1038/s41588-019-0569-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0569-6
This article is cited by
-
Prognostic biomarker discovery based on proteome landscape of Chinese lung adenocarcinoma
Clinical Proteomics (2024)
-
CellCharter reveals spatial cell niches associated with tissue remodeling and cell plasticity
Nature Genetics (2024)
-
Lung cancer in patients who have never smoked — an emerging disease
Nature Reviews Clinical Oncology (2024)
-
PET/CT-based deep learning grading signature to optimize surgical decisions for clinical stage I invasive lung adenocarcinoma and biologic basis under its prediction: a multicenter study
European Journal of Nuclear Medicine and Molecular Imaging (2024)
-
EGFR and ERBB2 Exon 20 Insertion Mutations in Chinese Non-small Cell Lung Cancer Patients: Pathological and Molecular Characterization, and First-Line Systemic Treatment Evaluation
Targeted Oncology (2024)