Abstract
About half of all cancers have somatic integrations of retrotransposons. Here, to characterize their role in oncogenesis, we analyzed the patterns and mechanisms of somatic retrotransposition in 2,954 cancer genomes from 38 histological cancer subtypes within the framework of the Pan-Cancer Analysis of Whole Genomes (PCAWG) project. We identified 19,166 somatically acquired retrotransposition events, which affected 35% of samples and spanned a range of event types. Long interspersed nuclear element (LINE-1; L1 hereafter) insertions emerged as the first most frequent type of somatic structural variation in esophageal adenocarcinoma, and the second most frequent in head-and-neck and colorectal cancers. Aberrant L1 integrations can delete megabase-scale regions of a chromosome, which sometimes leads to the removal of tumor-suppressor genes, and can induce complex translocations and large-scale duplications. Somatic retrotranspositions can also initiate breakage–fusion–bridge cycles, leading to high-level amplification of oncogenes. These observations illuminate a relevant role of 22 L1 retrotransposition in remodeling the cancer genome, with potential implications for the development of human tumors.
Similar content being viewed by others
Main
L1 retrotransposons are widespread repetitive elements in the human genome, representing 17% of the entire DNA content1,2. Using a combination of cellular enzymes and self-encoded proteins with endonuclease and reverse transcriptase activity, L1 elements copy and insert themselves at new genomic sites, in a process called retrotransposition. Most of the approximately 500,000 L1 copies in the human reference genome are truncated, inactive elements that are unable to retrotranspose. A small subset of them, around 100–150 L1 loci, remain active in the average human genome, acting as source elements, a small number of which consists of highly active copies termed hot-L1s3,4,5. These L1 source elements are usually transcriptionally repressed, but epigenetic changes that occur in tumors may promote their expression and allow them to retrotranspose6,7. Somatic L1 retrotransposition usually introduces a new copy of the 3′ end of the L1 sequence, and can also mobilize unique DNA sequences located immediately downstream of the source element, in a process called 3′ transduction7,8,9. L1 retrotransposons can also promote the somatic trans-mobilization of Alu elements, SINE-VNTR-Alu (SVA) elements and processed pseudogenes, which are copies of mRNAs that have been reverse transcribed into DNA and inserted into the genome with the machinery of active L1 elements10,11,12.
Approximately 50% of human tumors contain somatic retrotranspositions of L1 elements7,13,14,15. Previous analyses indicate that although a fraction of somatically acquired L1 insertions in cancer may influence gene function, the majority of retrotransposon integrations in a single tumor represent passenger mutations with little or no effect on cancer development7,13. Nonetheless, L1 elements are capable of promoting other types of genomic structural alterations in the germline and somatically, in addition to canonical L1 insertion events16,17,18; the effect of these alterations remains largely unexplored in the context of human cancer19,20.
To further understand the roles of retrotransposons in cancer, we developed strategies to analyze the patterns and mechanisms of somatic retrotransposition in 2,954 cancer genomes from 38 histological cancer subtypes within the framework of the PCAWG project21, many of which had not been evaluated for retrotransposition. On the basis of the robustness of the retrotransposition calls, we retained 296 tumors that were preliminarily excluded by the PCAWG Consortium21 (see Methods). Our analyses identify patterns and mutational mechanisms of structural variation in human cancers that are mediated by L1 retrotransposition. We found that the aberrant integration of L1 retrotransposons has a relevant role in remodeling the architecture of the cancer genome in some human tumors, mainly by promoting megabase-scale deletions that, occasionally, generate genomic consequences that may promote cancer development through the removal of tumor-suppressor genes, such as CDKN2A, or trigger the amplification of oncogenes, such as CCND1.
Results
The landscape of somatic retrotransposition in a large cancer whole-genome dataset
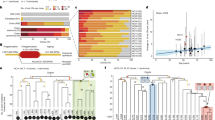
We ran our bioinformatic pipelines (Methods and Supplementary Note) to explore somatic retrotransposition on whole-genome sequencing data from 2,954 tumors and their matched normal pairs, across 38 cancer types (Supplementary Fig. 1 and Supplementary Table 1). The analysis retrieved a total of 19,166 somatically acquired retrotranspositions that were classified into six categories (Fig. 1a and Supplementary Table 2). Comprising 98% (18,739 out of 19,166) of the events, L1 integrations (14,967 solo-L1, 3,669 L1-transductions, and 103 L1-mediated rearrangements, which mainly comprised deletions) overwhelmingly dominate the landscape of somatic retrotransposition in the PCAWG dataset (Fig. 1a,b). By contrast, elements of the lineages Alu (Supplementary Fig. 2) and SVA (comprising 130 and 23 somatic copies, respectively) and processed pseudogenes, with 274 events, represent minor categories.
a) Number of somatic retrotransposition events identified in 2,954 cancer genomes across six categories: solo-L1, L1-mediated transductions (TD), L1-mediated rearrangements (RG), Alu, SVA and pseudogenes (PSD). b, Left, circos plot showing a head-and-neck tumor (Head-SCC) with high retrotransposition rate (638 somatic events). Right, a single pancreatic adenocarcinoma sample harboring around 26% (70 out of 274) of all processed pseudogenes identified in the PCAWG cohort. Chromosome ideograms are shown around the outer ring with individual rearrangements represented as arcs; colors match the type of rearrangement. c, For 31 PCAWG cancer types with sample size of n ≥ 15, data show the proportion of tumor samples with >100 (red), 10–100 (orange), 1–10 (yellow) and 0 (gray) somatic retrotranspositions. The number of samples analyzed for each tumor type is shown in parentheses. Retrotransposition enrichment or depletion for each tumor type together with the level of significance (zero-inflated negative binomial regression) is shown. *P < 0.05, **P < 0.01. NA, not applicable. d, Distribution of retrotransposition events per sample across the four tumor types significantly enriched in somatic retrotranspositions; the remaining tumors are grouped into ‘Other’. The number of samples from each group is shown in parentheses; point, median; box, 25th to 75th percentiles (interquartile range); whiskers, data within 1.5× the interquartile range. P values indicate significance from a two-tailed Mann–Whitney U-test. The y axis is shown on a logarithmic scale. e, For the same four tumor types in d, the fraction of structural variants (SV) belonging to six classes is shown: mobile element insertions (MEI), deletions (DEL), duplications (DUP), translocations (TRANS), head-to-head inversions (H2HINV) and tail-to-tail inversions (T2TINV). The total number of structural variants per cancer type is indicated on the right side of the panel.
The core pipeline, TraFiC-mem (Supplementary Fig. 3)—which was used to explore somatic retrotransposition in PCAWG—was validated by single-molecule whole-genome sequencing data analysis of one cancer cell line with high retrotransposition rate and its matched normal sample, confirming the somatic acquisition of 295 out of 308 retrotransposition events (false discovery rate <5%, Supplementary Fig. 4a,b). To further evaluate TraFiC-mem, we reanalyzed a mock cancer genome into which we had previously7 seeded somatic retrotransposition events at different levels of tumor clonality, and then simulated sequencing reads to the average level of coverage of the PCAWG dataset. The results confirmed a high precision (>99%) of TraFiC-mem, and a recall ranging from 90 to 94% for tumor clonalities from 25 to 100%, respectively (Supplementary Fig. 4c–e).
We observed marked variation in the retrotransposition rate across PCAWG tumor types (Fig. 1c and Supplementary Table 3). Overall, 35% (1,046 out of 2,954) of all cancer genomes have at least one retrotransposition event. However, esophageal adenocarcinoma, head-and-neck squamous carcinoma, lung squamous carcinoma and colorectal adenocarcinoma are significantly enriched in somatic retrotranspositions (Mann–Whitney U-test, P < 0.05; Fig. 1c,d and Supplementary Fig. 5). These four tumor types alone account for 70% (13,373 out of 19,166) of all somatic events in the PCAWG dataset, although they represent just 9% (266 out of 2,954) of the samples. This is particularly noticeable in esophageal adenocarcinoma, in which 27% (27 out of 99) of the samples show more than 100 separate somatic retrotranspositions (Fig. 1c), making L1 insertions the most frequent type of structural variation in esophageal adenocarcinoma (Fig. 1e). Furthermore, retrotranspositions are the second-most frequent type of structural variants in head-and-neck squamous and colorectal adenocarcinomas (Fig. 1e). To gain insights into the genetic causes that make some cancers more prone to retrotransposition than others, we looked for associations between retrotransposition and driver mutations in cancer-related genes. This analysis revealed an increased L1 retrotransposition rate in tumors with TP53 mutations (Mann–Whitney U-test, P < 0.05; Supplementary Fig. 6), and supports previous analyses that have suggested that TP53 functions to restrain mobile elements22,23. We also observe a widespread correlation between L1 retrotransposition and other types of structural variation (Spearman’s ρ = 0.44, P < 0.01; Supplementary Fig. 7), a finding that is most likely a consequence of a confounding effect of TP53-mutated genotypes (Supplementary Fig. 6).
We identified 43% (7,979 out of 18,636) somatic retrotranspositions of L1 inserted within gene regions including promoters, of which 66 events hit cancer-associated genes. The analysis of expression levels in samples with available transcriptome data, revealed four genes—including the ABL oncogene—with L1 retrotranspositions in the proximity of promoter regions that showed significant overexpression compared with the expression in the remaining samples of the same tumor type (Student’s t-test, q < 0.10; Supplementary Fig. 8a–c). The structural analysis of RNA-sequencing data identified instances in which portions of a somatic retrotransposition within a gene exonize, a process that sometimes involves cancer-associated genes (Supplementary Fig. 8d). In addition, we found evidence of aberrant fusion transcripts arising from the inclusion of processed pseudogenes in the target host gene and expression of processed pseudogenes landing in intergenic regions (Supplementary Fig. 8e).
Dissecting the genomic features that influence the landscape of L1 retrotranspositions in cancer
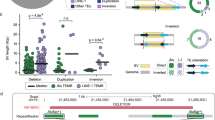
The genome-wide analysis of the distribution of somatic L1 insertions across the cancer genome revealed considerable variation in the rate of L1 retrotransposition (Fig. 2a and Supplementary Table 4). To understand the reasons behind such variation, we studied the association of L1 event rates with various genomic features. We first investigated whether the distribution of somatic L1s across the cancer genome could be determined by the occurrence of L1-endonuclease target-site motifs. We used a statistical approach based on negative binomial regression to deconvolute the influence of multiple overlapping genomic variables24; this analysis showed that close matches to the motif have a 244-fold increased L1 rate, compared with non-matched motifs (Fig. 2b and Supplementary Fig. 9a). Adjusting for this effect, we found a strong association with DNA replication time; the latest-replicating quarter of the genome was 8.9-fold enriched in L1 events (95% confidence interval, 8.25–9.71) compared with the earliest-replicating quarter (Fig. 2b,c and Supplementary Fig. 9b). Recent work25 has shown that L1 retrotransposition has a strong cell-cycle bias, and preferentially occurs during S phase. Our results are in agreement with these findings and suggest that L1 retrotransposition peaks in the later stages of nuclear DNA synthesis.
Genome-wide analysis of the distribution of 15,906 somatic L1 insertions, which include solo-L1 and L1 transductions with a 3′-poly(A) breakpoint characterized to base-pair resolution. a, The L1 insertion rate (purple) is shown together with the L1 endonuclease (EN) motif density (blue) and replication timing (orange). The data are represented per 1-Mb window. For illustrative purposes, only chromosome 3 is shown. b, Association between L1 insertion rate and multiple predictor variables at single-nucleotide resolution. Enrichment scores (thick dots) are adjusted for multiple covariates and compare the L1 insertion rate in bins 1–3 for a particular genomic feature (L1 endonuclease motif, replication timing, open chromatin, histone marks and expression level) versus bin 0 of the same feature, which therefore always has log-transformed enrichment = 0 by definition and is not shown. The error bars represent 95% confidence intervals. The number of observations per bin is provided in parentheses. MMs, the number of mismatches with respect to the consensus L1 endonuclease motif (see Supplementary Note). Heterochromatic regions and transcription elongation are defined based on H3K9me3 and H3K36me3 histone marks. Accessible chromatin is measured through DNase hypersensitivity. c, L1 insertion density, using kernel density estimate (KDE), along the replication timing spectrum. DNA replication timing is expressed on a scale from 80 (early) to 0 (late).
Next, we examined L1 rates in open chromatin measured using DNase hypersensitivity and, conversely, in closed heterochromatic regions by analyzing K9-trimethylated histone H3 (H3K9me3)26. When adjusting for the confounding effects of L1 motif content and replication time24, we found that somatic L1 events are enriched in open chromatin (1.27-fold in the top DNase hypersensitivity bin; 95% confidence interval, 1.14–1.41; Fig. 2b) and depleted in heterochromatin (1.72-fold, 95% confidence interval, 1.57–1.99; Fig. 2b). This finding differs from previous analyses, which have suggested that L1 insertions favored heterochomatin7—a discrepancy that we believe to be due to the confounding effect between heterochromatin and late-replicating DNA regions, which was not addressed in previous analyses. We also found a negative association of L1 rate with features of active transcription of chromatin, characterized by fewer L1 events at active promoters (1.63-fold; Supplementary Fig. 9c), a slight but significant reduction in L1 rates in highly expressed genes (1.25-fold lower; 95% confidence interval, 1.16–1.34; Fig. 2b) and a further depletion at H3K36me3 (1.90-fold reduction in the highest tertile; 95% confidence interval, 1.59–2.29; Fig. 2b), a mark of actively transcribed regions deposited in the body and at the 3′ end of active genes26. Further details on these associations are shown in Supplementary Fig. 9c–e and described in the Supplementary Note.
The contribution of L1 source elements to the pan-cancer retrotransposition burden
We used somatically mobilized L1 3′ transduction events to trace L1 activity to specific source elements7. This strategy revealed 124 germline L1 loci in the human genome that are responsible for most of the genomic variation generated by retrotransposition in the PCAWG dataset7,21 (Supplementary Table 5). To our knowledge, 52 of these loci represent previously unreported source elements in human cancer21. We analyzed the relative contribution of individual source elements to retrotransposition burden across cancer types, and found that retrotransposition is generally dominated by five hot-L1 source elements that alone give rise to half of all somatic transductions (Fig. 3a). This analysis revealed a dichotomous pattern of hot-L1 activity, with source elements that we have termed Strombolian and Plinian, given their similarity to these two types of volcanoes (Fig. 3b). Strombolian source elements are relatively indolent and produce small numbers of retrotranspositions in individual tumor samples, although they are often active and contribute substantially to overall retrotransposition in the PCAWG dataset. By contrast, Plinian elements are rarely active across tumors, but in these isolated cases, their activity is fulminant, causing large numbers of retrotranspositions.
a, The total number of transductions identified for each cancer type is shown as a blue-colored scale. The sample size for each tumor type is shown in parentheses. Contribution of each source element is defined as the proportion of the total number of transductions from each cancer type that is explained by each source locus. Only the top ten contributing source elements are shown, while the remaining are grouped into the category ‘Other’. b, Two extreme patterns of hot-L1 activity, Strombolian (blue) and Plinian (red), were identified. Dots show the number of transductions promoted by each source element in a given tumor sample. Arrows highlight violent eruptions (that is, strong peaks of somatic activity) in particular samples. c, Number of active germline L1 source elements per sample, across cancer types with source element activity. A source element is considered to be active in a given sample if it promotes at least one transduction. The enrichment or depletion of the number of active source elements for each tumor type together with the level of significance (zero-inflated negative binomial regression) is shown. *P < 0.05, **P < 0.01. The number of samples analyzed for each tumor type is shown in parentheses. d, Correlation between the number of somatic L1 insertions and the number of active germline L1 source elements in PCAWG samples. Each dot represents a tumor sample and colors match cancer types. Sample sizes (n), together with Spearman’s ρ and P values are shown above the panel.
At the individual tumor level, although we observed that the number of active source elements in a single cancer genome varied from 1 to 22, typically only 1 to 3 loci were operative (Fig. 3c). There is a correlation between somatic retrotranspositions and the number of active germline L1 source elements among PCAWG samples (Fig. 3d); this is likely one of the factors that explains why esophageal adenocarcinoma, lung and head-and-neck squamous carcinoma account for higher retrotransposition rates—in these three tumor types we also observed higher numbers of active germline L1 loci (Fig. 3c). Occasionally, somatic L1 integrations that retain their full length may also act as a source for subsequent somatic retrotransposition events7,27, and may reach high activity rates, leading them to dominate retrotransposition in a given tumor. For example, in a remarkable head-and-neck tumor sample, SA197656, we identified one somatic L1 integration at 4p16.1 that then triggered 18 transductions from its new site, with the next most active element being a germline L1 locus at 22q12.1, which accounted for 15 transductions (Supplementary Table 5).
Genomic deletions mediated by somatic L1 retrotransposition
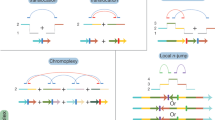
In cancer genomes with high somatic L1 activity rates, we observed that some L1 retrotransposition events followed a distinctive pattern that consisted of a single cluster of reads, associated with copy-number loss, for which the mates unequivocally identified one extreme of a somatic L1 integration with, apparently, no local, reciprocal cluster that supported the other extreme of the L1 insertion (Fig. 4a). Analysis of the associated copy-number changes identified the missing L1 reciprocal cluster at the far end of the copy-number loss, indicating that this pattern represents a deletion that occurred in conjunction with the integration of an L1 retrotransposon (Fig. 4b; see the Supplementary Note for additional information on how to interpret the paired-end mapping data from this and other figures). These rearrangements—called L1-mediated deletions—have been observed to occur somatically with engineered L1s in cultured human cells16,17 and naturally in the brain18, and are most likely the consequence of an aberrant mechanism of L1 integration.
a, In esophageal adenocarcinoma sample SA528802, we found a single cluster of reads on chromosome X, which is associated with one breakpoint of a copy-number loss, and for which the mates unequivocally identified one extreme of a somatic L1 integration. Paired-end reads are colored by the chromosome on which their mates can be found. Different colors for different reads from the same cluster indicate that mates are mapping a repetitive element. b, Analysis of the associated copy-number change on chromosome X identifies the missing L1 reciprocal cluster at the second breakpoint of the copy-number loss, and reveals a 3.9-kb deletion that occurs in conjunction with the integration of a 2.1-kb L1 somatic insertion. (A)n and (T)n represent poly(A) and poly(T) tails, respectively. c, Model of L1-mediated deletion. The integration of an L1 mRNA starts with L1-endonuclease cleavage promoting a 3′ overhang for reverse transcription. The cDNA (−) strand invades a second 3′ overhang from a pre-existing double-strand break upstream of the initial integration site. d, Distribution of the sizes of 90 L1-mediated deletions identified in the PCAWG dataset. e, In lung squamous carcinoma sample SA313800, a 34-bp truncated L1 insertion promotes a 1.1-kb deletion on chromosome 19. Because the L1 insertion was so short, we also identified discordant read pairs that span the L1 event and support the deletion. f, In esophageal adenocarcinoma sample SA528932, the integration on chromosome 3 of a 413-bp orphan L1 transduction from chromosome 7 causes a 2.5-kb deletion, which is supported by two clusters of discordant read pairs for which the mates map onto the transduced region of chromosome 7.
We developed specific algorithms to systematically identify L1-mediated deletions, and applied these methods across all PCAWG tumors. We identified 90 somatic events that matched the patterns described above, causing deletions of different size, which ranged in size from around 0.5 kb to 53.4 Mb (Fig. 4c and Supplementary Table 6). The reconstruction of the sequence at the breakpoint junctions in each case supports the presence of an L1-element—or L1-transduction—sequence and its companion polyadenylate tract, indicative of passage through an RNA intermediate. No target site duplication was found, which is also the typical pattern for L1-mediated deletions17. One potential mechanism for these events is that a molecule of L1 cDNA pairs with a distant 3′ overhang from a pre-existing double-strand DNA break generated upstream of the initial integration site, and the DNA region between the break and the original target site is subsequently removed by aberrant repair17 (Fig. 4d). Indeed, in 75% (47 out of 63) of L1-mediated deletions with a 5′-end breakpoint characterized to base-pair resolution, the analysis of the sequences at the junction revealed short (1–5 bp long, with median at 3 bp) microhomologies between the pre-integration site and the 5′ L1 sequence integrated right there (Supplementary Table 6). Furthermore, we found 14% (9 out of 63) instances in which short insertions (1–33 bp long, with median at 9 bp) are found at the 5′-breakpoint junction of the insertion. Both signatures are consistent with a non-homologous end-joining mechanism28, or other type of microhomology-mediated repair, for the 5′-end attachment of the L1 cDNA to a 3′ overhang from a pre-existing double-strand DNA break located upstream. L1-mediated deletions in which microhomologies or insertions are not found may follow alternative models17,29,30,31.
To confirm that these rearrangements are mediated by the integration of a single intervening retrotransposition event, we explored the PCAWG dataset for somatic L1-mediated deletions in which the L1 sequences at both breakpoints of the deletion could unequivocally be assigned to the same L1 insertion. These include small deletions and associated L1 insertions that were shorter than the library size, allowing sequencing read pairs to overlay the entire structure. For example, in a lung tumor sample, SA313800, we identified a deletion involving a 1-kb region of 19q12 with hallmarks of being generated by an L1 element (Fig. 4e). In this rearrangement, we found two different types of discordant read pairs at the deletion breakpoints: one cluster that supported the insertion of an L1 element and a second that spanned the L1 event and supported the deletion. Another type of L1-mediated deletion that could unequivocally be assigned to a single L1 insertion event is represented by those deletions generated by the integration of orphan L1 transductions. These transductions represent fragments of unique DNA sequence located downstream of an active L1 locus, which are mobilized without the companion L1 (refs. 7,15). For example, in one esophageal tumor sample, SA528932, we found a deletion of 2.5 kb on chromosome 3 mediated by the orphan transduction of a sequence downstream of an L1 locus on chromosome 7 (Fig. 4f).
Owing to the unavailability of PCAWG DNA specimens, we performed a validation of 16 additional somatic L1-mediated deletions that were identified by TraFiC-mem in two head-and-neck cancer cell lines with high retrotransposition rates, NCI-H2009 and NCI-H20877. We carried out two independent validation approaches, including PCR followed by single-molecule sequencing of amplicons, and Illumina whole-genome sequencing using mate-pair libraries with long insert size (3 kb and 10 kb). The results confirmed the somatic status of the rearrangements and a single L1-derived retrotransposition as the cause of the associated copy-number loss (Supplementary Figs. 10–12 and Supplementary Table 7).
Analysis of L1 3′-extreme insertion breakpoint sequences from L1-mediated deletions found in the PCAWG dataset revealed that 82% (74 out of 90) of the L1 events that caused deletions preferentially inserted into sequences that resemble L1-endonuclease consensus cleavage sites (for example, 5′-TTTT/A-3′ and related sequences32) (Supplementary Table 6). This confirms that the L1 machinery, through a target-primed reverse-transcription mechanism, is responsible for the integration of most of the L1 events that cause neighboring DNA loss32. Notably, in 16% (14 out of 90) of the events endonucleotidic cleavage occurred at the phosphodiester bond between a T and G instead of between the standard T and A site. In addition, we observed 8% (7 out of 90) instances in which the endonuclease motif was not found and the integrated element was truncated at both the 5′ and 3′ ends, suggesting that a small fraction of L1-associated deletions are the consequence of an L1-endonuclease-independent insertion mechanism30,31,32. Whatever mechanism of L1 integration is effective in each case, taken together, these data indicate that the somatic integration of L1 elements induces the associated deletions.
Megabase-size L1-mediated deletions cause loss of tumor-suppressor genes
Most L1-mediated deletions ranged from a few hundred to thousands of base pairs, although occasionally megabase-long regions of a chromosome were deleted (Fig. 4c and Supplementary Table 6). For example, in esophageal tumor sample SA528901, we found a 45.5-Mb interstitial deletion that involved the p31.3–p13.3 regions of chromosome 1 (Fig. 5a), in which both breakpoints of the rearrangement showed the hallmarks of a deletion mediated by integration of an L1 element. Here, the L1 element is 5′ truncated, which generated a small L1 insertion, allowing a fraction of the sequencing read pairs to span both breakpoints of the rearrangement. This unequivocally supports the model that the observed copy-number change is indeed a deletion mediated by retrotransposition of an L1 element. Similarly, in a lung tumor sample, SA313800, we found an interstitial L1-mediated deletion that induced the loss of 51.1 Mb from chromosome X, which included the centromere (Fig. 5b).
a, In esophageal adenocarcinoma sample SA528901, a 45.5-Mb interstitial deletion on chromosome 1 is generated after integration of a short L1 event. We observed a pair of clusters of discordant read pairs for which the mates support both extremes of the L1 insertion. Because the L1 element event is smaller than the library insert size, we also identified read pairs that span the L1 event and support the deletion. The L1-endonuclease 5′-TTTT/A-3′ motif identifies a target-primed reverse transcription (TPRT) L1-integration mechanism. b, In esophageal tumor sample SA313800, a partnered transduction7 (that is, the transduced region and its companion L1 source element) from chromosome 22 is integrated on chromosome X, promoting a 51.1-Mb deletion that removes the centromere. One negative cluster (green reads) supports a small region transduced from chromosome 22. c, L1-mediated deletions promote the loss of tumor-suppressor genes. In esophageal tumor sample SA528932, the somatic integration on chromosome 9 of a partnered transduction from chromosome 7, promotes a 5.3-Mb deletion that involves the loss of one copy of the tumor-suppressor gene CDKN2A. We observed a positive cluster of reads for which the mates map onto the 5′ extreme of an L1, and a negative cluster that contains split reads that match a poly(A) region and for which the mates map onto a region that is transduced from chromosome 7 (light blue). d, In a second esophageal adenocarcinoma sample, SA528899, the integration of an L1 retrotransposon generates an 8.6-Mb deletion that involves the same tumor-suppressor gene, CDKN2A. The sequencing data reveal two clusters—positive and negative—for which the mates support the L1 event.
L1-mediated deletions were, on occasion, driver events and caused the loss of tumor-suppressor genes. In esophageal tumor sample SA528932, the integration of an L1 transduction from chromosome 7p12.3 to the short arm of chromosome 9 caused a 5.3-Mb clonal deletion that involved the 9p21.3–9p21.2 region. This led to the loss of one copy of a key tumor-suppressor gene, CDKN2A (Fig. 5c), which is deleted in many cancer types including esophageal tumors33,34,35,36. Notably, the sequencing data revealed a somatic transduction that arose from this L1 element at its new insertion site, demonstrating that L1 events that promote deletions can be competent for retrotransposition (Supplementary Fig. 13). In a second esophageal tumor sample, SA528899, an L1 element integrated into chromosome 9 promoted an 8.6-Mb clonal deletion that encompasses the 9p22.1–9p21.1 region that removes one copy of the same tumor-suppressor gene, CDKN2A (Fig. 5d). Thus, L1-mediated deletions have clear oncogenic potential.
L1 retrotransposition generates other types of structural variation in human tumors
Somatic retrotransposition can also be involved in mediating or repairing more complex structural variants. In one esophageal tumor sample, SA528896, two separate L1-mediated structural variants were present within a complex cluster of rearrangements (Fig. 6a). In the first, an L1 transduction from a source element on chromosome 14q23.1 bridged an unbalanced translocation from chromosome 1p to 5q. A second somatic retrotransposition event bridged from chromosome 5p to an unknown part of the genome, completing a large interstitial copy-number loss on chromosome 5 that involves the centromere. This case suggests that retrotransposon transcripts and their reverse-transcriptase machinery can mediate breakage and repair of complex dsDNA breaks, spanning two chromosomes.
a, In esophageal adenocarcinoma sample SA528896, two separate L1 events mediate interchromosomal rearrangements. In the first, an L1 transduction from a source element on chromosome 14q23.1 bridged an unbalanced translocation from chromosome 1p to 5q. A second somatic retrotransposition event bridged from chromosome 5p to an unknown part of the genome, completing a 47.9-Mb interstitial copy-number loss on chromosome 5 that removes the centromere. b, In a cancer cell line, NCI-H2087, we found an interchromosomal translocation, between chromosomes 8 and 1, mediated by a region transduced from chromosome 6, which acts as a bridge and joins both chromosomes. We observed two read clusters, positive and negative, that demarcate the boundaries of the rearrangement, for which the mates support the transduction event. In addition, two reciprocal clusters span the insertion breakpoints, supporting the translocation between chromosomes 8 and 1. c, A model for megabase-size L1-mediated interchromosomal rearrangements. L1-endonuclease cleavage promotes a 3′ overhang in the negative strand, retrotranscription starts and the cDNA (−) strand invades a second 3′ overhang from a pre-existing double-strand break on a different chromosome, leading to translocation.
To explore this further, we identified single-L1 clusters with no reciprocal cluster in the cancer cell lines that were sequenced by using mate pairs with 3 kb and 10 kb inserts. Such events may correspond to hidden genomic translocations leading to the linkage of two different chromosomes, in which L1 retrotransposition is involved. One of the samples, NCI-H2087, showed translocation breakpoints at 1q31.1 and 8q24.12, both of which had the hallmarks of L1-mediated deletions, for which the mate-pair sequencing data identified an orphan L1 transduction from chromosome 6p24 that bridged both chromosomes (Fig. 6b). The configuration has also been confirmed by using long-read single-molecule sequencing (Supplementary Fig. 11). This interchromosomal rearrangement is likely mediated by the aberrant operation of L1-integration mechanism, in which the L1-transduced cDNA is wrongly paired with a second 3′ overhang from a pre-existing double-strand break generated in a second chromosome32 (Fig. 6c).
We also found evidence that L1 integrations can cause duplications of large genomic regions in human cancer. In esophageal tumor sample SA528848 (Fig. 7a), we identified two independent read clusters that support the integration of a small L1 event, coupled with a coverage drop at both breakpoints. Copy-number analysis revealed that the two L1 clusters demarcate the boundaries of a 22.6-Mb duplication that involves the 6q14.3–q21 region, suggesting that the L1 insertion could be the cause of such rearrangement by bridging sister chromatids during or after DNA replication (Fig. 7b). The analysis of the rearrangement data at the breakpoints identified read pairs that traverse the length of the L1 insertion breakpoints, and the L1-endonuclease motif is the L1 3′ insertion breakpoint, both confirming a single L1 event as the cause of a tandem duplication (Fig. 7a). Notably, this duplication increases the copy number of the cyclin C gene, CCNC, which is dysregulated in some tumors37.
a, In esophageal adenocarcinoma sample SA528848, we found a 22.6-Mb tandem duplication on the long arm of chromosome 6. The analysis of the sequencing data at the boundaries of the rearrangement breakpoints reveals two clusters of discordant read pairs for which the mates support the involvement of an L1 event. Because the L1 element was shorter than the library size, we also found two reciprocal clusters that aligned 22.6 Mb apart on the genome and in opposite orientation, spanning the insertion breakpoints and confirming the tandem duplication. An L1-endonuclease 5′-TTT/A-3′ degenerate motif was found. b, Large direct tandem duplications can be generated if the cDNA (−) strand invades a second 3′ overhang from a pre-existing double-strand break that occurred on a sister chromatid, and downstream to the initial integration site locus. c, In lung tumor sample SA313800, a small L1 insertion causes a 79.6-Mb duplication of the 14q arm through the induction of a fold-back inversion rearrangement. The analysis of the sequencing data at the breakpoint revealed two clusters of discordant read pairs (multi-colored reads) with the same orientation, aligning close together (5.5 kb apart) and demarcating a copy-number change for which the sequencing density is much greater on the right half of the rearrangement than the left. Both clusters of multi-colored reads support the integration of an L1. d, L1-mediated fold-back inversion model.
L1-mediated rearrangements can induce breakage–fusion–bridge cycles that trigger oncogene amplification
L1 retrotranspositions can also induce genomic instability by triggering breakage–fusion–bridge cycles. This form of genetic instability starts with end-to-end fusion of broken sister chromatids, and lead to a dicentric chromosome that forms an anaphase bridge during mitosis. Classically, the end-to-end chromosome fusions are thought to arise from telomere attrition38,39,40. We found, however, that somatic retrotransposition can induce the first inverted rearrangement that generates end-to-end fusion of sister chromatids. In lung tumor sample SA313800 (Fig. 7c), we found a small L1 event inserted on chromosome 14q that demarcates a copy-number change that involves a 79.6-Mb amplification of the 14q arm. Analysis of the sequencing data at the breakpoint revealed two discordant read clusters with the same orientation, which are 5.5 kb apart and support the integration of an L1. Both discordant clusters demarcate an increment of the sequencing coverage, for which the density is much greater in the right cluster. The only genomic structure that can explain this pattern is a fold-back inversion in which the two sister chromatids are bridged by an L1 retrotransposition in head-to-head (inverted) orientation (Fig. 7d).
In the example described above (Fig. 7c,d), no further breaks occurred, and the L1 retrotransposition generated an isochromosome (14q). In addition, we found examples in which the fusion of two chromatids by an L1 bridge induced further cycles of breakage–fusion–bridge repair. In esophageal tumor sample SA528848, we identified a cluster of reads on the long arm of chromosome 11 that had the typical hallmarks of an L1-mediated rearrangement (Fig. 8a). Copy-number data analysis showed that this L1 insertion point demarcated a 53-Mb deletion, which involved the loss of the telomeric region, from a region of massive amplification on chromosome 11. The amplified region on chromosome 11 contains the CCND1 oncogene, which is amplified in many human cancers41. The other end of this amplification was bound by a conventional fold-back inversion rearrangement (Fig. 8a), which is indicative of breakage–fusion–bridge repair42,43.
a, In esophageal adenocarcinoma sample SA528848, a single cluster of discordant reads (multi-colored reads) together with an L1-endonuclease cleavage site motif 5′-TTT/A-3′ supports the integration of an L1 event that demarcates a 53-Mb telomeric (that is, including the telomere) deletion, from a region of massive amplification that involves CCND1. Around 14 Mb upstream of the breakpoint of the deletion, we observed the presence of two clusters of read pairs (brown reads) that align close together and in the same orientation, which demarcate a change in copy number; this is a distinctive pattern of a fold-back inversion42,43, a rearrangement typically found to be associated with breakage–fusion–bridge (BFB) repair. In this fold-back inversion, the coverage shows much greater density on the right half of the rearrangement than the left, indicating that the abnormal chromosome is folded back on itself leading to duplicated genomic sequences in a head-to-head (inverted) orientation. The patterns described here suggest two independent breakage–fusion–bridge cycles, marked with (1) and (2). The copy-number plot shows the consensus total copy numbers (gold band) and the minor allele copy numbers (gray band). b, Models for the patterns described in a. The fold-back inversion model involves two breakage–fusion–bridge cycles, one induced by L1-mediated fold-back inversion (see Fig. 7d), and a second induced by standard breakage–fusion–bridge repair. The interchromosomal rearrangement model involves an interchromosomal rearrangement mediated by an L1, followed by one extra cycle of breakage–fusion–bridge repair. c, In lung cancer sample SA503541, the integration of an L1 retrotransposon is associated with a 50-Mb loss on 11q that includes the telomere, and activates breakage–fusion–bridge repair, which leads to the amplification of CCND1.
These patterns suggest the following sequence of events. During or soon after S phase, a somatic L1 retrotransposition bridges across sister chromatids in inverted orientation, breaking off the telomeric ends of 11q, which are then lost to the clone during the subsequent cell division (fold-back inversion model, Fig. 8b). The chromatids bridged by the L1 insertion now produce a dicentric chromosome. During mitosis, the two centromeres are pulled to opposite poles of the dividing cell, creating an anaphase bridge, which is resolved by further dsDNA breakage. This induces a second cycle of breakage–fusion–bridge repair, albeit not one mediated by L1 retrotransposition. These cycles lead to rapid-fire amplification of the CCND1 oncogene. Alternatively, an interchromosomal rearrangement mediated by L1 retrotransposition (interchromosomal rearrangement model, Fig. 8b) followed by two cycles of breakage–fusion–bridge repair could generate similar copy-number patterns with telomere loss and amplification of CCND1.
Our data show that L1-mediated retrotransposition is an alternative mechanism of creating the first dicentric chromosome that induces subsequent rounds of chromosomal breakage and repair. If this occurs near an oncogene, the resulting amplification can provide a powerful selective advantage to the clone. We searched the PCAWG dataset for other rearrangements that included copy-number amplifications from telomeric deletions that were mediated by L1 integration. We found four more such events across three cancer samples (Supplementary Fig. 14). In a lung tumor sample, SA503541, we found almost identical rearrangements to the one described above (Fig. 8c). In this case, a somatic L1 event also generated telomere loss that induced a second cycle of breakage–fusion–bridge repair. The megabase-size amplification of chromosomal regions also targeted the CCND1 oncogene, in which the boundaries were demarcated by the L1 insertion breakpoint and a fold-back inversion, which indicates breakage–fusion–bridge repair. The independent occurrence of these patterns, which involve the amplification of CCND1, in two different tumor samples (SA528848 and SA503541) demonstrates a mutational mechanism mediated by L1 retrotransposition, which likely contributes to the development of human cancer.
Discussion
Here we characterize the patterns and mechanisms of cancer retrotransposition on a multidimensional scale, across 2,954 cancer genomes, integrated with rearrangement, transcriptomic and copy-number data. Our analyses provide a new perspective on the long-standing question of whether the activation of retrotransposons is relevant in human oncogenesis. Our findings demonstrate that major restructuring of cancer genomes can sometimes emerge from aberrant L1 retrotransposition events in tumors with high retrotransposition rates, particularly in esophageal, lung and head-and-neck cancers. L1-mediated deletions can promote the loss of megabase-scale regions of a chromosome that may involve centromeres and telomeres. It is likely that the majority of such genomic rearrangements would be harmful for a cancer clone. However, occasionally, L1-mediated deletions may promote cancer-driving rearrangements that involve the loss of tumor-suppressor genes and/or the amplification of oncogenes, representing another mechanism by which cancer clones acquire new mutations that help them to survive and grow. We expect that structural variants induced by somatic retrotransposition in human cancer are more frequent than we could unambiguously characterize here, given the constraints on the fragment sizes of paired-end sequencing libraries. Long-read sequencing technologies should be able to provide a more comprehensive picture of how frequent such events are. Relatively few germline L1 loci in a given tumor, typically one to three copies, are responsible for such marked structural remodeling. Given the role that these L1 copies may have in some cancer types, this work underscores the importance of characterizing cancer genomes for patterns of L1 retrotransposition.
Methods
Pan-cancer datasets
Whole-genome sequencing dataset
We analyzed Illumina whole-genome paired-end sequencing reads (100–150 bp) from 2,954 tumors and matched normal samples across 38 cancer types21. On the basis of the robustness of the retrotransposition calls (false discovery rate of <5%), we opted to retain all samples that were preliminarily excluded by the PCAWG Consortium21, as they were excluded from SNV and structural variation analyses on the basis of read direction biases from PCR artifacts or poor sequence quality, but were not found to be problematic for retrotransposition analysis. For the majority of donors, the tumor specimens consisted of a fresh frozen sample, whereas the normal specimens consisted of a blood sample. Most of the tumor samples came from treatment-free primary cancers, although there was also a small number of donors with multiple samples of primary, metastatic and/or recurrent tumors. The average coverage was 30 reads per genome for normal samples, whereas tumor samples had a bimodal coverage distribution with maxima at 38 and 60 reads per bp (Supplementary Fig. 1 and Supplementary Table 1). BWA-mem44 v.0.7.8-r455 was used to align each tumor and normal sample to human reference build GRCh37. Additional technical details of the sequencing metrics are provided in Supplementary Table 1 and in the PCAWG lead paper21. The Ethics oversight for the PCAWG protocol was undertaken by the TCGA Program Office and the Ethics and Governance Committee of the ICGC.
Transcriptome dataset
About half of the donors (1,188) with whole-genome data in PCAWG had at least one tumor specimen with whole transcriptome obtained by RNA sequencing (RNA-seq). Mapping onto the reference was carried out using two independent read aligners, STAR45 v.2.4.0i, two-pass and TopHat2 (ref. 46) v.2.0.12. Gene expression was quantified with HTSeq47 v.0.6.1p1 and consensus normalized expression values, in fragments per kilobase of transcript per million mapped reads (FPKM), were obtained by averaging the expression from STAR and TopHat2. A more detailed description of RNA-seq data processing is provided by the PCAWG Integration of Transcriptome and Genome Working Group48.
Copy-number dataset
We analyzed copy-number profiles obtained by the PCAWG Evolution and Heterogeneity Working Group, using a consensus approach that combines six different state-of-the-art copy-number calling methods49. GC content corrected logR values were extracted using the Battenberg algorithm50, smoothed using a running median and transformed into copy-number space according to n = [(2(1 − ρ) + ψρ)2logR − 2(1 − ρ)]/ρ where ρ and ψ are consensus tumor purity and ploidy, respectively.
Structural variant dataset
The structural variation call set was generated by the PCAWG Somatic Structural Variation Working Group by merging the structural variant calls from four independent calling pipelines51. The merged structural variant calls were further required to have a consistent change in copy number.
Analysis of somatic retrotransposition
Detection of mobile element insertions using TraFiC-mem
BAM files from tumor and matched normal pairs were processed with TraFiC-mem v.1.1.0 to identify somatic mobile element insertions (MEIs) including solo-L1, L1-mediated transductions, Alu, SVA and ERV-K using Illumina paired-end mapping data. TraFiC-mem starts by identifying candidate somatic MEIs by analyzing discordant read pairs. In contrast to a previous version of the algorithm7, the new pipeline uses BWA-mem v.0.7.17 instead of RepeatMasker as the search engine for the identification of retrotransposon-like sequences in the sequencing reads. Calls obtained at this step are preliminary, in which MEI features are outlined and insertion coordinates represent ranges that surround the breakpoints. Then, a new module of TraFiC-mem, called MEIBA (from Mobile Element Insertion Breakpoint Analyzer), is used to identify the integration breakpoints to base-pair resolution and to perform a detailed characterization of MEI features, including structure, subfamily assignment and insertion site annotation. TraFiC-mem is illustrated in Supplementary Fig. 3. Detailed information about the pipeline is provided in the Supplementary Note.
Identification of germline and somatic L1 source elements
Because L1-mediated transductions are defined by the retrotransposition of unique, non-repetitive genomic sequences, we can unambiguously identify the L1 source element from which they derive7. The method relies on the detection of unique DNA regions retrotransposed somatically elsewhere in the cancer genome from a single locus that matches the 10-kb downstream region of a reference full-length L1 element or a putative non-reference polymorphic L1 element detected by TraFiC-mem across the matched normal samples in the PCAWG cohort21. When transduced regions were derived from the downstream region of a putative L1 event present in the tumor genome but not in the matched normal genome, we catalogued these elements as somatic L1 source loci.
Identification of processed pseudogene insertions
An additional separate module of TraFiC-mem was implemented for the identification of somatic insertions of processed pseudogenes. The method relies on the same principle as for the identification of somatic MEI events, through the detection of two reciprocal clusters of discordant read pairs, namely positive and negative, that supports an insertion event in the reference genome. However, the method differs from standard MEI calling to which the read mates map, as in this case mates are required to map to exons that belong to the same source protein-coding gene in GENCODE v.19. To avoid misclassification with standard genomic rearrangements that involve coding regions, we use MEIBA—described above—to reconstruct the insertion breakpoint junctions looking for hallmarks of retrotransposition, including the poly(A) tract and duplication of the target site. Candidate insertions without a poly(A) tail were discarded.
Identification of L1-mediated deletions
Independent read clusters, identified with TraFic-mem, supporting an L1 event (that is, clusters of discordant read pairs with no apparent reciprocal cluster within the proximal 500 bp, and for which the mates support a somatic L1 retrotransposition event) were interrogated for the presence of an associated change in copy number in its proximity. In brief, we looked for copy-number loss calls from the PCAWG Evolution and Heterogeneity Working Group for which the following conditions were fulfilled: (1) the upstream breakpoint matches an independent L1 cluster in positive orientation, (2) the corresponding downstream breakpoint, if any, from the same change in copy number matches an independent L1 cluster in negative orientation, and (3) the reconstruction of the structure of the putative insertion causing the deletion is compatible with one-single retrotransposition event. We used MEIBA (Supplementary Note) to reconstruct the insertion breakpoint junctions to confirm the ends of the events and identify hallmarks of retrotransposition, including the poly(A) tract and duplication of the target site.
An additional strategy was used for L1-mediated deletions that were shorter than 100 kb. Read-depth drops in the proximity of independent clusters were detected by, first, normalizing the read depth on each side of the cluster, using the matched normal sample as a reference. Then, the ratio between the normalized read depth on both sides of the cluster was computed for windows of 200–5,000 bp. Adjacent buffer regions of 300 bp on each side of the cluster were omitted from read-depth calculations to avoid false positives caused by sequence repeats. Pairs of independent reciprocal (positive–negative) clusters were selected such that: (1) both clusters were located less than 100 kb apart, (2) a potential drop in the read-depth ratio was identified, extending from the positive cluster to the negative cluster, and (3) the reconstruction of the structure of the putative insertion that caused the deletion was compatible with a single L1 event. For each cluster pair, the continuity and reliability of the copy-number drop was assessed by measuring the normalized read-depth ratio between non-overlapping 500-bp windows that spanned the region between the positive and negative clusters (that is the putative deletion) and windows upstream and downstream of the positive and negative clusters, respectively. The significance of each read-depth ratio drop was estimated nonparametrically using a null distribution of normalized read-depth ratios. This distribution was obtained for each tumor sample by randomly sampling 100,000 genomic locations (from copy-number segments showing the predominant copy number), and calculating read-depth ratios between both sides of each position. Nonparametric P values were calculated by comparing observed read-depth ratios with this null distribution, and adjusted using the Benjamini–Hochberg correction. Two cluster groups were produced: tier 1, pairs of reciprocal clusters with both clusters that had P < 0.1, and tier 2, pairs of reciprocal clusters with only one of both clusters having P < 0.1.
Retrotransposition rate enrichment and depletion across tumor types
For each tumor type with a minimum sample size of 15, we assessed whether it was enriched or depleted in retrotransposition compared to the overall retrotransposition burden using zero-inflated negative binomial regression, as implemented in the zeroinfl function of the pscl R package. This type of model takes into account the excess of zeros and the overdispersion that is present in this dataset. The MEI counts per sample were regressed on a binary factor that expressed whether they belonged to that particular type of cancer or to any other cancer type. On each regression, the magnitude and sign of the z-score indicates the effect size and directionality of the association. More specifically, positive z-scores indicate that a higher number of counts in the samples belongs to a particular cancer type compared with the rest (enrichment), whereas negative scores indicate a lower number of counts (depletion). Each z-score is accompanied by its P value to indicate the level of statistical significance.
Association between mutation in tumor-suppressor genes and retrotransposition and structural variantion rates
To assess whether the disruption of a particular tumor-suppressor gene was associated with a high level of retrotransposition, we used the whole-genome panorama of cancer driver events per sample produced by the PCAWG Drivers and Functional Interpretation Working Group21. This panorama includes coding and non-coding SNVs, insertions and deletions, copy-number alterations, structural variants and potentially predisposing germline variants. For each tumor-suppressor gene in the Cancer Gene Census database with mutational data, we stratified the samples into two groups—mutated tumor-suppressor genes and non-mutated tumor-suppressor genes. Then, we compared the distribution of MEI counts between both groups using a Mann–Whitney U-test to identify significant differences. P values were corrected for multiple testing using the Benjamini–Hochberg procedure. Adjusted P < 0.05 were considered significant. This analysis was done at both the level of the individual cancer type and the level of pan-cancer to identify tumor-type-specific associations. We further investigated whether there was a TP53 dosage effect as follows: every PCAWG sample was classified into three groups according to TP53 mutational status, namely wild-type, monoallelic and biallelic driver mutation. Then, the MEI counts distribution was compared for all possible group pair combinations using a Mann–Whitney U-test. The same analysis described above was applied to investigate the association between TP53 mutation and other types of structural variation.
Correlation between L1 insertion and structural variation rate
For each sample, we computed the number of MEIs, the total number of structural variants and the number of five different structural variant classes: deletions, duplications, translocations, head-to-head inversions and tail-to-tail inversions, when data were available. Then, the correlation between the number of MEIs and the structural variant burden was assessed at both the level of the individual tumor type and the level of the pan-cancer using a Spearman’s rank test.
Association between L1 insertion rate and genomic features
The L1 insertion rate was calculated as the total number of somatic L1 insertions, identified across the complete PCAWG cohort per 1-Mb window. The density of L1 endonuclease motifs was computed as the number of canonical endonuclease motifs, here defined as TTTT|R (where R is A or G) or Y|AAAA (where Y is C or T) per 1-Mb window. To study the association of L1 insertion rate with multiple variables at single-nucleotide resolution, we used a statistical framework based on negative binomial regression, as described in detail previously24, and adapted to adjust for the genome-wide distribution of the L1 endonuclease motif; we stratified the genome into four bins (0–3) by the closeness of match to the motif. Bin 0 contains dissimilar DNA motifs, with four or five (out of five) mismatches, encompassing 1,150 Mb. Bins 1, 2 and 3 contain loci with three, two and at most one mismatches, encompassing 749 Mb, 380 Mb and 114 Mb, respectively. The closer match of either of the two DNA strands at each locus was considered. Histone mark data and DNase hypersensitivity data were obtained from the Roadmap Epigenomics Consortium by averaging the fold-enrichment signal across eight cell types and processed by stratifying into four genomic bins as described previously24: bin 0 contains regions with below-baseline signal (fold enrichment versus input <1), while bins 1–3 are approximately equal-sized bins that cover the remainder of the genome. RNA-seq data from Roadmap were processed by averaging across eight cell types; bin 0 contains non-expressed genes (FPKM = 0) and intergenic DNA not listed as expressed, while bins 1–3 included genes with up to 0.59, 5.68 and above 5.68 FPKM, respectively. Replication time was averaged across the eight ENCODE cell types and divided into four equal-sized genomic bins, where bin 0 is latest and bin 3 is earliest replicating. Essential genes were estimated from CRISPR screens in cell lines52. All enrichment scores shown compare bins 1–3 for a particular feature (replication time, histone marks, gene expression, L1 motif) versus bin 0 of the same feature. Bin 0 therefore always has log enrichment = 0 by definition and is not shown on plots. The analyses were restricted to regions of the genome with perfect CRG75 alignability.
Impact of retrotransposition insertions on gene expression
To study the transcriptional impact of a somatic L1 insertion within COSMIC cancer genes and promoters, we used RNA-seq data to compare gene-expression levels in samples with and without somatic L1 insertion. For each somatic L1 insertion within a cancer gene or promoter, we compared the gene FPKM between the sample having the insertion (study sample) and the remaining samples of the same tumor type (control samples). Using the distribution of gene-expression levels in control samples, we calculated the normalized gene expression differences using a Student’s t-test. To overcome the problems due to multiple testing, false discovery rate–adjusted P values (q values) were calculated using the Benjamini–Hochberg procedure, and adjusted P < 0.1 was considered to be significant.
Analysis of processed pseudogene expression
We analyzed the PCAWG RNA-seq data to identify and characterize the transcriptional consequences of somatic integrations of processed pseudogenes (PSD). We interrogated RNA-seq split reads and discordant read pairs, looking for chimeric retrocopies that involved PSDs and target genomic regions. For each PSD insertion somatic call, we extracted all of the RNA-seq reads (when available), mapping the source gene and the insertion target region, together with the RNA-seq unmapped reads for the corresponding sample. Then, we used these reads as a query in BLASTn53 v.2.7.1 searches against a database that contained all isoforms of the source gene described in RefSeq54, together with the genomic sequence ranging from −5 kb to +5 kb around the PSD integration site. Finally, we looked for RNA-seq discordant read pairs and/or RNA-seq clipped reads that supported the joint expression of PSD and target site. Only read pairs with one of the mates aligned to the host gene mRNA with >98% identity were considered. All expression signals were confirmed by visual inspection with Integrative Genomics Viewer v.2.4.10.
Validation of somatic retrotransposition algorithms
In silico validation of TraFiC-mem
To evaluate the precision and recall of our algorithm TraFiC-mem, we reanalyzed a mock cancer genome into which we had previously seeded known somatic retrotransposition events at different levels of tumor clonality7. To create the artificial, tumoral genome, 10,000 L1 insertion breakpoints—including solo-L1, partnered and orphan transductions—were randomly distributed in the standard reference genome using BedTools v.2.25.0, of which 9,227 were inserted out of un-sequenced gaps. Then, ART55 (v.MountRainier-2016-06-05) was used to generate paired-end read sequencing data for both the standard and the artificial reference genomes to a 38× coverage. The simulation FASTQ files were aligned into the standard reference genome with BWA-mem56 v.0.7.17. Reads from the normal and tumor BAM files were randomly subsampled and merged with samtools v.1.7 at three distinct proportions to also produce tumor samples with 25%, 50% and 75% clonalities. After that, the four possible tumor and matched normal pairs were processed with TraFiC-mem to call MEIs. For each clonality, the identified MEIs were compared with the list of simulated MEIs to compute the number of true-positive (TP), false-positive (FP), true-negative (TN) and false-negative (FN) calls. Finally, precision and recall were computed as follows: Precision = TP/(TP + FP); Recall = TP/(TP + FN).
Validation of TraFiC-mem calls using single-molecule sequencing
We performed validation of 308 putative somatic retrotranspositions identified with TraFiC-mem in one cancer cell line (NCI-H2087) with high retrotransposition rate, and absent in the matched normal cell line (NCI-BL2087) derived from blood, by single-molecule sequencing using Oxford Nanopore technology. Genomic DNA was sheared to 10-kb fragments using Covaris g-TUBEs, and cleaned with 0.4× Ampure XP Beads. After end-repairing and dA-tailing using the NEBNext End Repair/dA-tailing module (NEB), whole-genome libraries were constructed with the Oxford Nanopore Sequencing 1D ligation library prep kit (SQK-LSK108, Oxford Nanopore Technologies). Genomic libraries were loaded on MinION R9.4 flowcells. We used the Oxford Nanopore basecaller Albacore v.2.0.1 to generate fastq files. After quality filtering of the fastq files and read trimming of the data with Porechop v.0.2.3, we used minimap2 (ref. 57) v.2.10-r764-dirty to map sequencing reads onto the hs37d5 reference genome. Sequencing coverages were 8.2× (NCI-BL2087) and 9.17× (NCI-H2087), and average read sizes of mapped reads were ~4.5 kb (NCI-BL2087) and ~11 kb (NCI-H2087). After obtaining the whole-genome BAM files for each of the 308 putative somatic retrotransposition calls identified with TraFic-mem, we interrogated the long-read tumor BAM file to find reads that validated the event. MEIs supported by at least one Nanopore read in the tumor and absent in the matched normal sample were considered true-positive somatic events, while MEIs not supported by long reads in the tumor and/or present in the matched normal were considered false-positive calls. Overall, we found 4.22% (13/308) false-positive events. False discovery rate (FDR) was estimated as follows: FDR = FP/(TP + FP).
Validation of L1-mediated rearrangements with PCR and single-molecule sequencing
We performed validation of 20 somatic L1-mediated rearrangements, mostly deletions, identified in two cancer cell lines with high retrotransposition rates (NCI-H2009 and NCI-H2087). We carried out PCR followed by single-molecule sequencing of amplicons from the two tumor cell lines and their matched normal samples (NCI-BL2009 and NCI-BL2087) using a MinION from Oxford Nanopore. PCR primers were designed to amplify three regions from each event (namely, 5′-extreme, 3′-extreme and target sites) as shown in Supplementary Fig. 10.
Validation of L1-mediated rearrangements using mate pairs
To further validate and characterize L1-mediated rearrangements, we performed 10× mate-pair whole-genome sequencing using libraries with two different insert sizes (4 kb and 10 kb), which can span the integrated L1 element that caused the deletion, enabling the validation of the involvement of L1 in the generation of such rearrangements. Mate-pair reads (100 nucleotides long) were aligned to the human reference with BWA-mem v.0.7.17. Then, for each candidate L1-mediated rearrangement, we searched for discordant mate-pair clusters that spanned the breakpoints and supported the L1-mediated event. Each event was confirmed by visual inspection of the BAM files using Integrative Genomics Viewer v.2.4.10.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Somatic and germline variant calls, mutational signatures, subclonal reconstructions, transcript abundance, splice calls and other core data generated by the ICGC/TCGA PCAWG Consortium are available for download at https://dcc.icgc.org/releases/PCAWG. Additional information on accessing the data, including raw read files, can be found at https://docs.icgc.org/pcawg/data/. In accordance with the data access policies of the ICGC and TCGA projects, most molecular, clinical and specimen data are in an open tier, which does not require access approval. To access potentially identifying information, such as germline alleles and underlying sequencing data, researchers will need to apply to the TCGA Data Access Committee (DAC) via dbGaP (https://dbgap.ncbi.nlm.nih.gov/aa/wga.cgi?page=login) for access to the TCGA portion of the dataset, and to the ICGC Data Access Compliance Office (DACO; http://icgc.org/daco) for the ICGC portion. In addition, to access somatic SNVs derived from TCGA donors, researchers will also need to obtain dbGaP authorization.
In addition, the analyses in this paper used a number of datasets that were derived from the raw sequencing data and variant calls (Supplementary Table 8). The individual datasets are available at Synapse (https://www.synapse.org/), which are also mirrored at DCC portal (https://dcc.icgc.org). Full links, filenames, accession numbers and descriptions are detailed in Supplementary Table 8. VCF files for somatic mobile element insertions described specifically in this manuscript can be found at Synapse, under accession number syn21052009, and in DCC portal at https://dcc.icgc.org/releases/PCAWG/retrotransposition.
Code availability
The core computational pipelines used by the PCAWG Consortium for alignment, quality control and variant calling are available to the public at https://dockstore.org/search?search=pcawg under a GNU General Public License v.3.0, which allows for reuse and distribution. The algorithm for the identification of somatic retrotransposition events (TraFiC-mem) is available at https://gitlab.com/mobilegenomesgroup/TraFiC (v.1.2.0).
Change history
21 March 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41588-023-01319-9
References
International Human Genome Sequencing Consortium Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Kazazian, H. H. Jr. Mobile elements: drivers of genome evolution. Science 303, 1626–1632 (2004).
Sassaman, D. M. et al. Many human L1 elements are capable of retrotransposition. Nat. Genet. 16, 37–43 (1997).
Brouha, B. et al. Hot L1s account for the bulk of retrotransposition in the human population. Proc. Natl Acad. Sci. USA 100, 5280–5285 (2003).
Beck, C. R. et al. LINE-1 retrotransposition activity in human genomes. Cell 141, 1159–1170 (2010).
Menendez, L., Benigno, B. B. & McDonald, J. F. L1 and HERV-W retrotransposons are hypomethylated in human ovarian carcinomas. Mol. Cancer 3, 12 (2004).
Tubio, J. M. et al. Extensive transduction of nonrepetitive DNA mediated by L1 retrotransposition in cancer genomes. Science 345, 1251343 (2014).
Holmes, S. E., Dombroski, B. A., Krebs, C. M., Boehm, C. D. & Kazazian, H. H. Jr. A new retrotransposable human L1 element from the LRE2 locus on chromosome 1q produces a chimaeric insertion. Nat. Genet. 7, 143–148 (1994).
Moran, J. V., DeBerardinis, R. J. & Kazazian, H. H. Jr. Exon shuffling by L1 retrotransposition. Science 283, 1530–1534 (1999).
Kazazian, H. H. Jr. Processed pseudogene insertions in somatic cells. Mob. DNA 5, 20 (2014).
Cooke, S. L. et al. Processed pseudogenes acquired somatically during cancer development. Nat. Commun. 5, 3644 (2014).
Ewing, A. D. et al. Retrotransposition of gene transcripts leads to structural variation in mammalian genomes. Genome Biol. 14, R22 (2013).
Lee, E. et al. Landscape of somatic retrotransposition in human cancers. Science 337, 967–971 (2012).
Helman, E. et al. Somatic retrotransposition in human cancer revealed by whole-genome and exome sequencing. Genome Res. 24, 1053–1063 (2014).
Solyom, S. et al. Extensive somatic L1 retrotransposition in colorectal tumors. Genome Res. 22, 2328–2338 (2012).
Symer, D. E. et al. Human L1 retrotransposition is associated with genetic instability in vivo. Cell 110, 327–338 (2002).
Gilbert, N., Lutz-Prigge, S. & Moran, J. V. Genomic deletions created upon LINE-1 retrotransposition. Cell 110, 315–325 (2002).
Erwin, J. A. et al. L1-associated genomic regions are deleted in somatic cells of the healthy human brain. Nat. Neurosci. 19, 1583–1591 (2016).
Burns, K. H. Transposable elements in cancer. Nat. Rev. Cancer 17, 415–424 (2017).
Kazazian, H. H. Jr. & Moran, J. V. Mobile DNA in health and disease. N. Engl. J. Med 377, 361–370 (2017).
The ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. Pan-cancer analysis of whole genomes. Nature https://doi.org/10.1038/s41586-020-1969-6 (2020).
Wylie, A. et al. p53 genes function to restrain mobile elements. Genes Dev. 30, 64–77 (2016).
Jung, H., Choi, J. K. & Lee, E. A. Immune signatures correlate with L1 retrotransposition in gastrointestinal cancers. Genome Res. 28, 1136–1146 (2018).
Supek, F. & Lehner, B. Clustered mutation signatures reveal that error-prone DNA repair targets mutations to active genes. Cell 170, 534–547 (2017).
Mita, P. et al. LINE-1 protein localization and functional dynamics during the cell cycle. eLife 7, e30058 (2018).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Kimberland, M. L. et al. Full-length human L1 insertions retain the capacity for high frequency retrotransposition in cultured cells. Hum. Mol. Genet 8, 1557–1560 (1999).
Chang, H. H. Y., Pannunzio, N. R., Adachi, N. & Lieber, M. R. Non-homologous DNA end joining and alternative pathways to double-strand break repair. Nat. Rev. Mol. Cell Biol. 18, 495–506 (2017).
Han, K. et al. Genomic rearrangements by LINE-1 insertion-mediated deletion in the human and chimpanzee lineages. Nucleic Acids Res. 33, 4040–4052 (2005).
Sen, S. K., Huang, C. T., Han, K. & Batzer, M. A. Endonuclease-independent insertion provides an alternative pathway for L1 retrotransposition in the human genome. Nucleic Acids Res. 35, 3741–3751 (2007).
Farkash, E. A., Kao, G. D., Horman, S. R. & Prak, E. T. Gamma radiation increases endonuclease-dependent L1 retrotransposition in a cultured cell assay. Nucleic Acids Res. 34, 1196–1204 (2006).
Gilbert, N., Lutz, S., Morrish, T. A. & Moran, J. V. Multiple fates of L1 retrotransposition intermediates in cultured human cells. Mol. Cell Biol. 25, 7780–7795 (2005).
Zhou, C., Li, J. & Li, Q. CDKN2A methylation in esophageal cancer: a meta-analysis. Oncotarget 8, 50071–50083 (2017).
The Cancer Genome Atlas Research Network Comprehensive genomic characterization of squamous cell lung cancers. Nature 489, 519–525 (2012).
The Cancer Genome Atlas Network Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 517, 576–582 (2015).
The Cancer Genome Atlas Research Network Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Xu, W. & Ji, J. Y. Dysregulation of CDK8 and cyclin C in tumorigenesis. J. Genet. Genomics 38, 439–452 (2011).
Artandi, S. E. et al. Telomere dysfunction promotes non-reciprocal translocations and epithelial cancers in mice. Nature 406, 641–645 (2000).
O’Hagan, R. C. et al. Telomere dysfunction provokes regional amplification and deletion in cancer genomes. Cancer Cell 2, 149–155 (2002).
Maciejowski, J. & de Lange, T. Telomeres in cancer: tumour suppression and genome instability. Nat. Rev. Mol. Cell Biol. 18, 175–186 (2017).
Beroukhim, R. et al. The landscape of somatic copy-number alteration across human cancers. Nature 463, 899–905 (2010).
Campbell, P. J. et al. The patterns and dynamics of genomic instability in metastatic pancreatic cancer. Nature 467, 1109–1113 (2010).
Li, Y. et al. Constitutional and somatic rearrangement of chromosome 21 in acute lymphoblastic leukaemia. Nature 508, 98–102 (2014).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner Bioinformatics 29, 15–21 (2013).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
PCAWG Transcriptome Core Group et al. Genomic basis for RNA alterations in cancer. Nature https://doi.org/10.1038/s41586-020-1970-0 (2020).
Gerstung, M. et al. The evolutionary history of 2,658 cancers. Nature https://doi.org/10.1038/s41586-019-1907-7 (2020).
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012).
Rheinbay, E. et al. Analyses of non-coding somatic drivers in 2,693 cancer whole genomes. Nature https://doi.org/10.1038/s41586-020-1965-x (2020).
Meyers, R. M. et al. Computational correction of copy number effect improves specificity of CRISPR–Cas9 essentiality screens in cancer cells. Nat. Genet. 49, 1779–1784 (2017).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
O'Leary, N.A. et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 44, D733–D745 (2016).
Huang, W., Li, L., Myers, J. R. & Marth, G. T. ART: a next-generation sequencing read simulator. Bioinformatics 28, 593–594 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Acknowledgements
J.M.C.T. is supported by European Research Council (ERC) Starting Grant 716290 ‘SCUBA CANCERS’, Ramon y Cajal grant RYC-2014-14999 and Spanish Ministry of Economy, Industry and Competitiveness (MINECO) grant SAF2015-66368-P. B.R.-M., E.G.A., M.S.G. and S.Z. are supported by PhD fellowships from Xunta de Galicia (Spain) ED481A-2016/151, ED481A-2017/299, ED481A-2017/306 and ED481A-2018/199, respectively. F.S. was supported by ERC Starting Grant 757700 ‘HYPER-INSIGHT’, MINECO grant BFU2017-89833-P ‘RegioMut’, and further acknowledges institutional funding from the MINECO Severo Ochoa award and from the CERCA Programme of the Catalan Government. Y.S.J. was supported by Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (grant number HI16C2387). A.L.B. is supported by MINECO PhD fellowship BES-2016-078166. M.T. was supported by MINECO grant SAF2015-73916-JIN. R.B. received funding through the National Institutes of Health (U24CA210978 and R01CA188228). M.G.B. received funding through MINECO, AEI, Xunta de Galicia and FEDER (BFU2013-41554-P, BFU2016-78121-P, ED431F 2016/019). N.B. is supported by a My First AIRC grant from the Associazione Italiana Ricerca sul Cancro (number 17658). J.D. is a postdoctoral fellow of the Research Foundation Flanders (FWO) and the European Union’s Horizon 2020 research and innovation program (Marie Skłodowska-Curie grant agreement number 703594-DECODE). K.C. and Z.C. are supported by NIH R01 CA172652 and U41 HG007497. Z.C. is supported by an American Heart Association Institutional Data Fellowship Award (17IF33890015). P.A.E. is supported by Cancer Research UK. E.A.L. is supported by K01AG051791. I.M. is supported by Cancer Research UK (C57387/A21777). F.M. is supported by A.I.L. (Associazione Italiana Contro le Leucemie-Linfomi e Mieloma ONLUS) and by S.I.E.S. (Società Italiana di Ematologia Sperimentale). S.M.W. received funding through a SNSF Early Postdoc Mobility fellowship (P2ELP3_155365) and an EMBO Long-Term Fellowship (ALTF 755-2014). J.W. received funding from the Danish Medical Research Council (DFF-4183-00233). D.C.W. is funded by the Li Ka Shing foundation and the National Institute for Health Research (NIHR) Oxford Biomedical Research Centre. J.O.K. is supported by an ERC Starting Grant. P.V.L. is a Winton Group Leader in recognition of the Winton Charitable Foundation’s support towards the establishment of The Francis Crick Institute. This work is supported by The Francis Crick Institute, which receives its core funding from Cancer Research UK (FC001202), the UK Medical Research Council (FC001202) and the Wellcome Trust (FC001202). H.H.K. is supported by grants from the National Institute of General Medical Sciences (P50GM107632 and 1R01GM099875). K.H.B. is supported by P50GM107632, R01CA163705 and R01GM124531. This work was supported by the TransTumVar project PN013600. R.C.F. thanks Cancer Research UK Programme Grant for esophageal ICGC, Cambridge BRC and ECMC infrastructure support. This work was supported by the Wellcome Trust grant 09805. We acknowledge the contributions of the many clinical networks across the ICGC and TCGA who provided samples and data to the PCAWG Consortium, and the contributions of the Technical Working Group and the Germline Working Group of the PCAWG Consortium for collation, realignment and harmonized variant calling of the cancer genomes used in this study. We thank the patients and their families for their participation in the individual ICGC and TCGA projects.
Author information
Authors and Affiliations
Consortia
Contributions
J.M.C.T. and P.J.C. designed the study and wrote the manuscript with assistance from B.R.-M., H.H.K. and K.H.B. B.R.-M., A.B.-O., J.Z. and J.M.C.T. designed the somatic retrotransposition algorithms and performed retrotransposition analyses with support from A.L.B., E.G.A., S.Z., Z.C., K.C., E.A.L., D.A., J.M., G.B., M.G.B., H.D., F.C.P.N., S.M.W., M.G., P.A.E., P.J.P., H.H.K. and K.H.B. F.S. carried out genomic features analyses and wrote the associated manuscript section. Y.S.J. performed transcriptomic analyses. A.D.-B., M.P., B.R.-M., D.T. and J.M.C.T. performed the analysis of processed pseudogenes. J.D., A.B.-O., S.C.D. and P.V.L. carried out copy-number analyses. Y.L., N.D.R., S.E.S., J.W., J.A.W., R.B., J.O.K. and P.J.C. performed structural variation analyses. B.R.-M., J.Z., I.M., D.C.W. and P.V.L. carried out statistical analyses. J.T., J.R.-C., M.T., M.S. and D.G.-S. performed sample preparation, experimental validation and evaluation of the sequencing data. K.M.R., J.Z. and A.B. supported variant calling algorithms and sequencing analysis platforms. F.M., N.B., G.C. and R.C.F. provided additional genomic data, samples and experimental materials. All authors reviewed the manuscript during its preparation. R.B. and P.J.C. were working group or project co-leaders.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–14 and Notes
Supplementary Tables
Supplementary Tables 1–8
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rodriguez-Martin, B., Alvarez, E.G., Baez-Ortega, A. et al. Pan-cancer analysis of whole genomes identifies driver rearrangements promoted by LINE-1 retrotransposition. Nat Genet 52, 306–319 (2020). https://doi.org/10.1038/s41588-019-0562-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0562-0
This article is cited by
-
Characterization of active transposable elements and their new insertions in tuber propagated greater yam (Dioscorea alata)
BMC Genomics (2024)
-
ZNF689 deficiency promotes intratumor heterogeneity and immunotherapy resistance in triple-negative breast cancer
Cell Research (2024)
-
Locus-specific LINE-1 expression in clinical ovarian cancer specimens at the single-cell level
Scientific Reports (2024)
-
Towards targeting transposable elements for cancer therapy
Nature Reviews Cancer (2024)
-
The genomic landscape of Vk*MYC myeloma highlights shared pathways of transformation between mice and humans
Nature Communications (2024)