Abstract
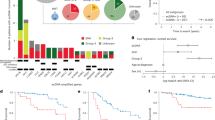
Extrachromosomal circularization of DNA is an important genomic feature in cancer. However, the structure, composition and genome-wide frequency of extrachromosomal circular DNA have not yet been profiled extensively. Here, we combine genomic and transcriptomic approaches to describe the landscape of extrachromosomal circular DNA in neuroblastoma, a tumor arising in childhood from primitive cells of the sympathetic nervous system. Our analysis identifies and characterizes a wide catalog of somatically acquired and undescribed extrachromosomal circular DNAs. Moreover, we find that extrachromosomal circular DNAs are an unanticipated major source of somatic rearrangements, contributing to oncogenic remodeling through chimeric circularization and reintegration of circular DNA into the linear genome. Cancer-causing lesions can emerge out of circle-derived rearrangements and are associated with adverse clinical outcome. It is highly probable that circle-derived rearrangements represent an ongoing mutagenic process. Thus, extrachromosomal circular DNAs represent a multihit mutagenic process, with important functional and clinical implications for the origins of genomic remodeling in cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The WGS and RNA-seq data that support the findings of this study have been deposited with the European Genome-phenome Archive (https://www.ebi.ac.uk/ega/) under accession nos. EGAS00001001308 and EGAS00001004022. The Circle-seq data that support the findings of this study are available from the corresponding author upon request. Source data for Fig. 1 are available online.
Code availability
The scripts used to analyze the sequencing data have been uploaded to www.github.com/henssenlab. Data on tree-shaped rearrangements can be accessed and visualized online (https://kons.shinyapps.io/trees/).
Change history
27 February 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41588-020-0598-1
References
Turner, K. M. et al. Extrachromosomal oncogene amplification drives tumour evolution and genetic heterogeneity. Nature 543, 122–125 (2017).
Møller, H. D. et al. Circular DNA elements of chromosomal origin are common in healthy human somatic tissue. Nat. Commun. 9, 1069 (2018).
Shibata, Y. et al. Extrachromosomal microDNAs and chromosomal microdeletions in normal tissues. Science 336, 82–86 (2012).
Pennisi, E. Circular DNA throws biologists for a loop. Science 356, 996 (2017).
Verhaak, R. G. W., Bafna, V. & Mischel, P. S. Extrachromosomal oncogene amplification in tumour pathogenesis and evolution.Nat. Rev. Cancer 19, 283–288 (2019).
Møller, H. D., Parsons, L., Jørgensen, T. S., Botstein, D. & Regenberg, B. Extrachromosomal circular DNA is common in yeast. Proc. Natl Acad. Sci. USA 112, E3114–E3122 (2015).
Tjio, J. H. & Levan, A. The chromosome number of man. Hereditas 42, 1–6 (1956).
Garsed, D. W. et al. The architecture and evolution of cancer neochromosomes. Cancer Cell 26, 653–667 (2014).
Rausch, T. et al. Genome sequencing of pediatric medulloblastoma links catastrophic DNA rearrangements with TP53 mutations. Cell 148, 59–71 (2012).
Kohl, N. E. et al. Transposition and amplification of oncogene-related sequences in human neuroblastomas. Cell 35, 359–367 (1983).
deCarvalho, A. C. et al. Discordant inheritance of chromosomal and extrachromosomal DNA elements contributes to dynamic disease evolution in glioblastoma. Nat. Genet. 50, 708–717 (2018).
Nikolaev, S. et al. Extrachromosomal driver mutations in glioblastoma and low-grade glioma. Nat. Commun. 5, 5690 (2014).
Schwab, M. et al. Amplified DNA with limited homology to myc cellular oncogene is shared by human neuroblastoma cell lines and a neuroblastoma tumour. Nature 305, 245–248 (1983).
Balaban-Malenbaum, G. & Gilbert, F. Double minute chromosomes and the homogeneously staining regions in chromosomes of a human neuroblastoma cell line. Science 198, 739–741 (1977).
Cox, D., Yuncken, C. & Spriggs, A. I. Minute chromatin bodies in malignant tumours of childhood. Lancet 1, 55–58 (1965).
Sanborn, J. Z. et al. Double minute chromosomes in glioblastoma multiforme are revealed by precise reconstruction of oncogenic amplicons. Cancer Res. 73, 6036–6045 (2013).
Deshpande, V. et al. Exploring the landscape of focal amplifications in cancer using AmpliconArchitect. Nat. Commun. 10, 392 (2019).
Avet-Loiseau, H. et al. Morphologic and molecular cytogenetics in neuroblastoma. Cancer 75, 1694–1699 (1995).
Dillon, L. W. et al. Production of extrachromosomal microDNAs is linked to mismatch repair pathways and transcriptional activity. Cell Rep. 11, 1749–1759 (2015).
Forbes, S. A. et al. COSMIC: somatic cancer genetics at high-resolution. Nucleic Acids Res. 45, D777–D783 (2017).
Bouzas-Rodriguez, J. et al. Neurotrophin-3 production promotes human neuroblastoma cell survival by inhibiting TrkC-induced apoptosis. J. Clin. Invest. 120, 850–858 (2010).
Xu, K. Structure and evolution of double minutes in diagnosis and relapse brain tumors. Acta Neuropathol. 137, 123–137 (2019).
Storlazzi, C. T. et al. Gene amplification as double minutes or homogeneously staining regions in solid tumors: origin and structure. Genome Res. 20, 1198–1206 (2010).
Villamón, E. et al. Genetic instability and intratumoral heterogeneity in neuroblastoma with MYCN amplification plus 11q deletion. PLoS ONE 8, e53740 (2013).
Marrano, P., Irwin, M. S. & Thorner, P. S. Heterogeneity of MYCN amplification in neuroblastoma at diagnosis, treatment, relapse, and metastasis. Genes Chromosomes Cancer 56, 28–41 (2017).
Gröbner, S. N. et al. The landscape of genomic alterations across childhood cancers. Nature 555, 321–327 (2018).
Northcott, P. A. et al. Enhancer hijacking activates GFI1 family oncogenes in medulloblastoma. Nature 511, 428–434 (2014).
Henssen, A. G. et al. PGBD5 promotes site-specific oncogenic mutations in human tumors. Nat. Genet. 49, 1005–1014 (2017).
Henssen, A. G. et al. Genomic DNA transposition induced by human PGBD5. eLife 4, e10565 (2015).
Henssen, A. G. et al. Forward genetic screen of human transposase genomic rearrangements. BMC Genomics 17, 548 (2016).
Guzmán, C., Bagga, M., Kaur, A., Westermarck, J. & Abankwa, D. ColonyArea: an ImageJ plugin to automatically quantify colony formation in clonogenic assays. PLoS ONE 9, e92444 (2014).
Henssen, A. G., MacArthur, I., Koche, R. & Dorado-García, H. Purification and sequencing of large circular DNA from human cells. Protoc Exch (2019); https://doi.org/10.1038/protex.2019.006
Boeva, V. et al. Control-FREEC: a tool for assessing copy number and allelic content using next-generation sequencing data. Bioinformatics 28, 423–425 (2012).
Van Loo, P. et al. Allele-specific copy number analysis of tumors. Proc. Natl Acad. Sci. USA 107, 16910–16915 (2010).
Chong, Z. et al. novoBreak: local assembly for breakpoint detection in cancer genomes. Nat. Methods 14, 65–67 (2017).
Wala, J. A. et al. SvABA: genome-wide detection of structural variants and indels by local assembly. Genome Res. 28, 581–591 (2018).
Rausch, T. et al. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 28, i333–i339 (2012).
Moncunill, V. et al. Comprehensive characterization of complex structural variations in cancer by directly comparing genome sequence reads. Nat. Biotechnol. 32, 1106–1112 (2014).
Helmsauer, K. Tree-shaped Rearrangement Patterns in Pediatric Cancer Genomes (2019); https://kons.shinyapps.io/trees
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
We thank A. Kentsis, S. Armstrong, B. Regenberg, F. Speleman, S. Perner, U. Ohler and N. Hübener for critical discussions, and K. Astrahantseff for editorial advice. We thank D. Sunaga-Franze, C. Quedenau, M. Sohn, K. Richter and C. Langnick for technical support. A.G.H. is supported by the Deutsche Forschungsgemeinschaft (German Research Foundation; grant no. 398299703) and the Wilhelm Sander Stiftung (2018.011.1). A.G.H., A.K. and S.F. are participants in the Berlin Institute of Health-Charité Clinical Scientist Program funded by the Charité-Universitätsmedizin Berlin and the Berlin Institute of Health. A.G.H., S.F., K.H. and V.B. are supported by Berliner Krebsgesellschaft e.V. K.H. is supported by Boehringer Ingelheim Fonds. This work was also supported by the TransTumVar project (project no. PN013600). This project was supported by the Berlin Institute of Health within the collaborative research project TERMINATE-NB (CRG04). We thank the patients and their parents for granting access to the tumor specimens and clinical information that were analyzed in this study. We thank B. Hero, H. Düren and N. Hemstedt of the Neuroblastoma Biobank and Neuroblastoma Trial Registry (University Children’s Hospital Cologne) of the GPOH for providing samples and clinical data.
Author information
Authors and Affiliations
Contributions
R.P.K., E.R.F., K.H., J.M., F.H., I.C.M., R.C., A.W., M.B., M.P., C. Röefzaad, T.T., H.D., R.F.S., C. Rosswog, J. Theissen, V.B., N.M.P., H.D.G., Y.B., A.S., N. Timme, K.K., S.F., N. Thiessen, E.B., K.S, A.K., P.H., J. Toedling, M.F., D.B., S.S., A.E., D.T., J.H.S. and A.G.H. contributed to the study design and the collection and interpretation of the data. R.P.K. performed the analysis of Circle-seq and WGS. E.R.F. performed the data analysis of the WGS data. I.C.M., R.C., N.M.P. and H.D.G. performed the circular DNA extraction and PCR-based validation. V.B., C. Röefzaad, A.W., P.H., K.S., M.F., A.S. and F.H. collected and prepared the patient samples. C. Rosswog. and J. Theissen performed and analyzed the FISH. M.B. and R.F.S. performed the haplotype phasing analyses. M.P. and J. Toedling. analyzed the tumor genome sequencing data. M.B. performed the allele-specific analysis of Circle-seq, WGS and RNA-seq. S.F., F.K., R.P.K, K.H. and J.M. performed the RNA-seq data analysis. P.H., H.D.G., N.M.P., A.S., D.B. and K.S. performed the experiments and analyzed the data. R.P.K., A.K., A.E. and J.H.S. contributed to the study design. R.P.K. and A.G.H. led the study design, performed the data analysis and wrote the manuscript. All authors contributed to manuscript drafting.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–12, Tables 1–4 and Note
Source data
Source Data Fig. 1
Unprocessed Western blots for Supplementary Fig. 10f–h.
Rights and permissions
About this article
Cite this article
Koche, R.P., Rodriguez-Fos, E., Helmsauer, K. et al. Extrachromosomal circular DNA drives oncogenic genome remodeling in neuroblastoma. Nat Genet 52, 29–34 (2020). https://doi.org/10.1038/s41588-019-0547-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0547-z
This article is cited by
-
Differential expression and analysis of extrachromosomal circular DNAs as serum biomarkers in pulmonary arterial hypertension
Respiratory Research (2024)
-
High-fidelity (repeat) consensus sequences from short reads using combined read clustering and assembly
BMC Genomics (2024)
-
Dynamics of extrachromosomal circular DNA in rice
Nature Communications (2024)
-
Machine learning-based extrachromosomal DNA identification in large-scale cohorts reveals its clinical implications in cancer
Nature Communications (2024)
-
Lineage specific transcription factor waves reprogram neuroblastoma from self-renewal to differentiation
Nature Communications (2024)