Abstract
A large number of putative cis-regulatory sequences have been annotated in the human genome, but the genes they control remain poorly defined. To bridge this gap, we generate maps of long-range chromatin interactions centered on 18,943 well-annotated promoters for protein-coding genes in 27 human cell/tissue types. We use this information to infer the target genes of 70,329 candidate regulatory elements and suggest potential regulatory function for 27,325 noncoding sequence variants associated with 2,117 physiological traits and diseases. Integrative analysis of these promoter-centered interactome maps reveals widespread enhancer-like promoters involved in gene regulation and common molecular pathways underlying distinct groups of human traits and diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All raw and processed data have been deposited in the GEO database under accession number GSE86189. Visualization of processed pcHi-C data is available at http://www.3div.kr/capture_hic.
Code availability
Code for pcHi-C interaction detection can be made available on request. For other data analysis, we used publicly available software.
References
Welter, D. et al. The NHGRI GWAS Catalog, a curated resource of SNP-trait associations. Nucleic Acids Res. 42, D1001–D1006 (2014).
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Hindorff, L. A. et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl Acad. Sci. USA 106, 9362–9367 (2009).
Lettice, L. A. et al. A long-range Shh enhancer regulates expression in the developing limb and fin and is associated with preaxial polydactyly. Hum. Mol. Genet 12, 1725–1735 (2003).
Uslu, V. V. et al. Long-range enhancers regulating Myc expression are required for normal facial morphogenesis. Nat. Genet. 46, 753–758 (2014).
Claussnitzer, M. et al. FTO obesity variant circuitry and adipocyte browning in humans. N. Engl. J. Med. 373, 895–907 (2015).
Smemo, S. et al. Obesity-associated variants within FTO form long-range functional connections with IRX3. Nature 507, 371–375 (2014).
Yu, M. & Ren, B. The three-dimensional organizationof mammalian genomes. Annu. Rev. Cell Dev. Biol. 33, 265–289 (2017)..
de Wit, E. et al. The pluripotent genome in three dimensions is shaped around pluripotency factors. Nature 501, 227–231 (2013).
Sanyal, A., Lajoie, B. R., Jain, G. & Dekker, J. The long-range interaction landscape of gene promoters. Nature 489, 109–113 (2012).
Dixon, J. R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015).
Jin, F. et al. A high-resolution map of the three-dimensional chromatin interactome in human cells. Nature 503, 290–294 (2013).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Tang, Z. et al. CTCF-mediated human 3D genome architecture reveals chromatin topology for transcription. Cell 163, 1611–1627 (2015).
Sahlen, P. et al. Genome-wide mapping of promoter-anchored interactions with close to single-enhancer resolution. Genome Biol. 16, 156 (2015).
Jager, R. et al. Capture Hi-C identifies the chromatin interactome of colorectal cancer risk loci. Nat. Commun. 6, 6178 (2015).
Mifsud, B. et al. Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C. Nat. Genet. 47, 598–606 (2015).
Dryden, N. H. et al. Unbiased analysis of potential targets of breast cancer susceptibility loci by Capture Hi-C. Genome Res. 24, 1854–1868 (2014).
Martin, P. et al. Capture Hi-C reveals novel candidate genes and complex long-range interactions with related autoimmune risk loci. Nat. Commun. 6, 10069 (2015).
Javierre, B. M. et al. Lineage-specific genome architecture links enhancers and non-coding disease variants to target gene promoters. Cell 167, 1369–1384 e19 (2016).
Freire-Pritchett, P. et al. Global reorganisation of cis-regulatory units upon lineage commitment of human embryonic stem cells. eLife 6, e21926 (2017).
Siersbaek, R. et al. Dynamic rewiring of promoter-anchored chromatin loops during adipocyte differentiation. Mol. Cell 66, 420–435 e5 (2017).
Rubin, A. J. et al. Lineage-specific dynamic and pre-established enhancer-promoter contacts cooperate in terminal differentiation. Nat. Genet. 49, 1522–1528 (2017).
Orlando, G. et al. Promoter capture Hi-C-based identification of recurrent noncoding mutations in colorectal cancer. Nat. Genet. 50, 1375–1380 (2018).
Leung, D. et al. Integrative analysis of haplotype-resolved epigenomes across human tissues. Nature 518, 350–354 (2015).
Consortium, G. T. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Schmitt, A. D. et al. A compendium of chromatin contact maps reveals spatially active regions in the human genome. Cell Rep. 17, 2042–2059 (2016).
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Zhang, Y. et al. Chromatin connectivity maps reveal dynamic promoter-enhancer long-range associations. Nature 504, 306–310 (2013).
Rajagopal, N. et al. High-throughput mapping of regulatory DNA. Nat. Biotechnol. 34, 167–174 (2016).
Diao, Y. et al. A tiling-deletion-based genetic screen for cis-regulatory element identification in mammalian cells. Nat. Methods 14, 629–635 (2017).
Engreitz, J. M. et al. Local regulation of gene expression by lncRNA promoters, transcription and splicing. Nature 539, 452–455 (2016).
Dao, L. T. M. et al. Genome-wide characterization of mammalian promoters with distal enhancer functions. Nat. Genet. 49, 1073–1081 (2017).
Roadmap Epigenomics, C. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Richard, M., Drouin, R. & Beaulieu, A. D. ABC50, a novel human ATP-binding cassette protein found in tumor necrosis factor-alpha-stimulated synoviocytes. Genomics 53, 137–145 (1998).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Yang, D. et al. 3DIV: A 3D-genome interaction viewer and database. Nucleic Acids Res. 46, D52–D57 (2018).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Acknowledgments
We thank members of the Ren laboratory for critical suggestions in the course of this work. We thank N. Nariai (UCSD) for sharing LD information. This work was funded in part by the Ludwig Institute for Cancer Research (to B.R.), NIH (grant no.1R01ES024984 to B.R.), the Ministry of Science, ICT and Future Planning through the National Research Foundation in Republic of Korea (grant no. 2017R1C1B2008838 to I.J.), Korean Ministry of Health and Welfare (grant no. HI17C0328 to I.J.) and SUHF Fellowship (to I.J.).
Author information
Authors and Affiliations
Contributions
I.J., A.S., Y.D. and B.R. conceived the study. I.J., A.S. and Y.D. performed experiments with assistance from T.L., C.T. and S.C., I.J., A.J.L. and D.Y. performed data analysis with assistance from J.E., M.C., Z.C. and C.L.B., D.K. supervised data analysis by D.Y., C.K., E.M. and C.L.B. contributed to provide human brain tissue samples. B.L. and S.K. contributed to sequencing and initial data processing. I.J. prepared the manuscript with assistance from A.S., Y.D., A.J.L., J.E. and B.R. All authors read and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
B.R. is a co-founder of Arima Genomics, Inc. A.S. is an employee of Arima Genomics.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Capture Hi-C design, probe synthesis, and workflow of target enrichment.
a, Schematic of probe design for Promoter Capture Hi-C experiments. For each promoter (black rectangle), two flanking HindIII cut sites were identified. Three 120-mer capture probes were designed for each side of the HindIII cut site, with a 30bp shift between the adjacent probes. In total, 12 capture probes were designed for each promoter and all probes were targeted to the Watson Strand. b, Schematic of custom RNA probe synthesis. Single stranded DNA (ssDNA) probe synthesis by CustomArray, Inc., is shown from top to bottom; PCR amplification with SP6 recognition sequence completion and purification, BsrDI digestion and purification, in vitro transcription in the presence of biotinylated UTP and purification, and pooling of probe batches using equal mass ratios. c, Schematic workflow of target enrichment of Hi-C libraries (Promoter Capture Hi-C). From top to bottom, preparation of library mix, hybridization buffer, and probe mix, followed by combining the mixes and overnight incubation to bind probes to Hi-C template. Then, preparation of streptavidin beads and wash buffers, followed by binding of RNA:DNA duplexes to streptavidin beads and rigorous washing to remove off-target binding. And lastly, PCR amplification of the resulting Promoter Capture Hi-C library.
Supplementary Figure 2 Overview of samples and capture probe quality control.
a, Overview of cell and tissue types analyzed by Promoter Capture-Hi-C and note of other datasets available for these samples. Embryonic or embryonic-derived cell types are on the left and tissues are tabled on the right according to their developmental origin. b, Histogram of DNA fragments containing different number of TSS. c, A scatter plot showing the reproducibility of probe density from RNA-seq data between two probe synthesis experiments. Each dot on the scatter plot represents a single promoter and the value is the aggregated probe density from all probes assigned to that given promoter (n = 19,462). Pearson correlation coefficient value is shown together. d, Venn diagram showing the number of targeted regions that contain detectable probe density based on RNA-seq of the capture probes from each replicate of probe synthesis. e, Snapshot of Promoter Capture-Hi-C probe density from RNA-seq analysis of the capture probes. Two replicates of probe synthesis and subsequent RNA-seq are shown, followed by GENCODE gene annotations. f, A genome-browser view of the distribution of the capture probes, as determined by RNA-seq, in a ~200kb window. Below the replicate RNA-seq datasets are the HindIII cut sites and GENCODE gene annotations, illustrating that the vast majority of probe density is only found around HindIII restriction sites flanking promoters. g, h, Histograms of the probe densities measured by RNA-seq (x-axis) in each promoter from replicate 1 (g) and replicate 2 (h) of probe synthesis.
Supplementary Figure 3 General characterization of promoter-centered long-range chromatin interactions.
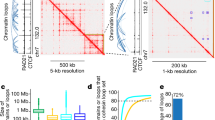
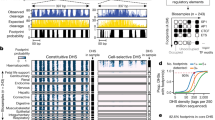
a, Genome Browser views of the identified pcHi-C chromatin contacts in different cell/tissue types. The darkness of blue denotes the strength of interactions. RefSeq genes are presented below the snapshots. b, A histogram of the number of chromatin interactions that span a given genomic distance. Orange line indicates the accumulated fraction of chromatin interactions from all 27 tissues/cell types. c, Fraction of pcHi-C interactions uniquely detected in one cell/tissue type (green) or also detected in other cell/tissue types (orange). The average fraction of cell/tissue-specific interactions is not over-estimated due to the number of tested samples (at 22 samples the fraction of cell/tissue-specific interactions reach plateau) and tissue-heterogeneity (similar trend was observed when we only considered pcHi-C interactions obtained from cell lines). d, Snapshot of a locus showing promoter-centered long-range interactions revealed by pcHi-C in H1-hESC (bottom, purple loops) in the context of TAD annotations (blue rectangles) (top, red) in H1-hESC. RefSeq genes are shown at the bottom. e, Fraction of P-O pcHi-C chromatin interactions in the context of TAD annotations with the respective cell/tissue types.
Supplementary Figure 4 Validation of the identified promoter capture Hi-C interactions.
a, A genome-browser snapshot of the CCL gene cluster, highlighting the similarity of promoter-centered interactions from Promoter Capture Hi-C and the high resolution Hi-C data in IMR90. The top two tracks show histone modification signals for H3K4me3 and H3K27ac, followed by a track showing the RefSeq genes. Below these tracks are pcHi-C chromatin interactions in IMR90 (blue colored arcs) and promoter-centered chromatin interactions from high-resolution Hi-C data in IMR90 (reddish brown arcs). b-e, ROC plots illustrating the performance of predicting Promoter Capture Hi-C result for in situ Hi-C loops anchored at promoters in lymphoblastoid (b), IMR90 (c), hippocampus (d), and dorsolateral prefrontal cortex (e). Promoter centered interactions for in situ Hi-C loops were considered as true interactions, and ROC plots are drawn for the corresponding pcHi-C result. ROC scores are shown in the ROC plot. f, ROC plots showing the reproducibility of pcHi-C chromatin interactions between two biological replicates. pcHi-C interactions from one replicate are used as true interactions, and ROC plots are drawn for the other replicate. g-k. Venn diagrams presenting the number of commonly identified pcHi-C interactions between biological replicates (n=2) for lymphoblastoid (g), dorsolateral prefrontal cortex (h), mesenchymal stem cell (i), lymphoblastoid processed by CHICAGO (j), and GM12878 with previously published pcHi-C data18 (k). Hypergeometric p-values are shown together. l-m, Illustration of interaction intensity in the n=2 biological replicates of lymphoblastoid (l) and mesenchymal stem cells (m), depending on the replicate consistency. For the boxplots, the box represents the interquartile range (IQR), and the whiskers correspond to the highest and lowest points within 1.5×IQR.
Supplementary Figure 5 Integrative analysis of long-range chromatin interactions with chromatin modification and accessibility.
a, b, Histograms of the number of cREs per promoter (a) and number of putative target promoters per cRE (b). Y-axis indicates frequency and the x-axis indicates the number of cREs per promoter (a) or number of promoters per cRE (b). c, Depiction of identified long-range promoter-centered interactions across a 0.84Mb locus in lymphoblastoid (top). Shown below are histone modification signals obtained from ChIP-seq analyses35, accessible chromatin measured from DNaseI hypersensitivity assay. d, Depiction of extensively interacting DNA fragments (EIF) from P-P and P-O interactions, and transcription factor (TF) binding clusters identified in GM12878 cells for the same region shown in Supplementary Fig. 5c. Below the tracks are TF binding profiles (n=67) obtained from ChIP-seq results performed in GM12878 cells listed in Supplementary Table 9. Highlighted in translucent blue are overlapping EIF and TF binding clusters. EIF was defined in each cell/tissue type by selecting frequently interacting DNA fragments with multiple promoters in terms of 0.01 Poisson p value cutoff. e, f, Boxplots showing the number of P-O EIF overlapping with TF clusters (blue dot) compared to random expectation in lymphoblastoid (e) and H1-hESC (f). Random expectation values (n = 1,000) were calculated by using typical TF peaked regions (blue) and generating random genomic regions (green). One-sided empirical p-value shows statistical significance (*** p value < 0.001). For the boxplots, the box represents the interquartile range (IQR), and the whiskers correspond to the highest and lowest points within 1.5×IQR. g, h, Boxplots showing the number of P-P EIF overlapping with TF clusters compared to random expectation in lymphoblastoid (g) and H1-hESC (h). Random expectation values (n=1,000) were calculated by randomly selecting typical TF peaked regions (blue) and randomly generating genomic regions (green). One-sided empirical p-value shows statistical significance (*** p value < 0.001). For the boxplots, the box represents the interquartile range (IQR), and the whiskers correspond to the highest and lowest points within 1.5×IQR. i, An array of boxplots showing the number of P-O EIF overlapping with super-enhancers (orange dot), compared to typical enhancers (blue) and random genomic regions (purple). Random expectation values obtained by random permutations (n=10,000). One-sided empirical p-value showed statistical significance for all tested cell/tissue types compared to random genomic regions (p value < 0.0001). For the boxplots, the box represents the interquartile range (IQR), and the whiskers correspond to the highest and lowest points within 1.5×IQR.
Supplementary Figure 6 Enrichment of long-range chromatin interactions at various chromatin states generated by a 50-state ChromHMM model.
a, Boxplots showing the fold change of chromatin interaction frequencies between active/bivalent promoters and each chromatin state over the expected values. The 50 chromatin states (E01-E50) were obtained from the 50-state ChromHMM model. Two-sided KS-tests were performed between active promoters and bivalent promoters across multiple cell lines (n=5) (two adjacent boxplots) (** p value < 0.01 and *** p value < 0.001). The chromatin states of a DNA fragment that interacts more frequently with active promoters than bivalent promoters were highlighted in pink asterisk. The chromatin states of a DNA fragment that interacts more frequently with bivalent promoters than active promoters were highlighted in blue asterisk. For the boxplots, the box represents the interquartile range (IQR), and the whiskers correspond to the highest and lowest points within 1.5×IQR. b, A heatmap showing an emission parameter matrix of each chromatin state in which each row corresponds to a different chromatin state and each column corresponds to an emission probability of a chromatin mark shown at the top. The pre-calculated emission parameter heatmap was downloaded from the 50-state ChromHMM model established by Roadmap Epigenomics Project.
Supplementary Figure 7 Comparison between the P-O pcHi-C interactions and eQTL-associations found in different tissues/cell-types.
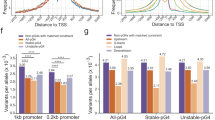
a-c, Illustrative LocusZoom plots of eQTLs for the HS3ST1 (a), the METTL25 (b), and the DAAM1 (c) gene expression in left ventricle, dorsolateral prefrontal cortex, and aorta, respectively. RefSeq genes are plotted below the LocusZoom plot. Identified pcHi-C interactions are shown as loops (purple) in the bottom. d, Boxplots showing fold enrichment between the number of eQTL-associations matched to P-O pcHi-C interactions and that of distance matched random P-O pcHi-C interactions (n=1,000) for 12 corresponding tissue types. P-O interactions in all 12 tissues were significantly enriched for eQTL associations (one-sided empirical p value < 0.01). For the boxplots, the box represents the interquartile range (IQR), and the whiskers correspond to the highest and lowest points within 1.5×IQR. e, Array of boxplots showing the number of matched eQTL associations between P-O pcHi-C chromatin interactions after exclusion of DNA fragment shared promoters and random expectation (n=1,000) across 14 matched tissue types from GTEx database. All P-O pcHi-C interactions are significantly enriched by eQTL associations compared to random P-O pcHi-C interactions with or without distance match (* one-sided empirical p-value <0.05, ** one-sided empirical p-value <0.01, *** one-sided empirical p-value <0.001). For the boxplots, the box represents the interquartile range (IQR), and the whiskers correspond to the highest and lowest points within 1.5×IQR.
Supplementary Figure 8 Analysis of correlation between P-O pcHi-C interactions and cRE-promoter activities.
a, Shown are boxplots of the distribution of PCC between H3K27ac signals in cRE-promoter pairs connected by pcHi-C interactions after exclusion of multiple fragment spanning cREs (Orange, n=154,055), compared to the distribution of random expectation with matched distance (dark gray, n=154,055) and without matched distance (gray, n=154,055). We only considered P-O pairs where other DNA fragments are marked by H3K27ac peaks in at least one cell/tissue type analyzed. Statistical test was performed between P-O pairs and random control, demonstrating that P-O pairs showed significant positive correlation (*** Two-sided Welch’s t-test p value < 2.2e-16). For the boxplots, the box represents the interquartile range (IQR), and the whiskers correspond to the highest and lowest points within 1.5×IQR. b, Similar to Supplementary Fig. 8a, but the distribution of PCC between H3K27ac signals at a cRE and target gene expressions of the cRE connected by pcHi-C interactions. Statistical test was performed between P-O pairs (orange, n=154,055), distance matched random control (dark gray, n=154,055), and random control (gray, n=154,055), revealing that P-O pairs showed significant positive correlation (*** Two-sided Welch’s t-test p value < 2.2e-16). For the boxplots, the box represents the interquartile range (IQR), and the whiskers correspond to the highest and lowest points within 1.5×IQR. c, Boxplots showing transcript levels of the putative target genes (n = 2,339) connected by hippocampus specific pcHi-C interactions. Genes are significantly expressed in hippocampus compared to other cell/tissues types (one-sided KS-test p value < 0.005) except dorsolateral prefrontal cortex (one-sided KS-test p value 0.27) and mesenchymal stem cell (one-sided KS-test p value 0.02). For the boxplots, the box represents the interquartile range (IQR), and the whiskers correspond to the highest and lowest points within 1.5×IQR.
Supplementary Figure 9 Characterization of promoter-promoter interactions.
a, Pie chart showing the fraction of promoter-promoter interactions (P-P) among all pcHi-C interactions. The fraction of P-P pcHi-C interactions modestly decreases to 6.5% after excluding fragments that harbor multiple promoters. b, An array of boxplots showing the number of eQTL associations matched to P-P pcHi-C interactions (purple dot), compared to random expectation (n=1,000) with matched distance (middle, blue) and without matched distance (right, light blue). One-sided empirical p values are shown at the top (* < 0.05, ** < 0.01, *** < 0.001). For the boxplots, the box represents the interquartile range (IQR), and the whiskers correspond to the highest and lowest points within 1.5×IQR. c, d, Illustrative LocusZoom plots of FHOD1 eQTLs (c) and POFUT2 eQTLs (d) in left ventricle and aorta, respectively. Promoters that contain significant eQTLs and target promoters are highlighted in translucent orange. Dots along the LocusZoom plot represent SNPs, and their significance of association with FHOD1 and POFUT2 gene expression is plotted along the left y-axis, respectively. The blue line traveling across the scatterplot indicates the estimated recombination rate, as plotted along the right y-axis. RefSeq genes and RNA-seq are plotted below the LocusZoom plot. pcHi-C interactions are shown as purple in the bottom.
Supplementary Figure 10 Experimental validation of enhancer-like promoter functions.
a, An illustrative LocusZoom plot of eQTLs for DACT3 gene expression in dorsolateral prefrontal cortex. Both the DACT3 gene promoter region and the AP2S1 gene promoter that contains significant eQTLs are highlighted in translucent orange, dots along the LocusZoom plot represent SNPs, and their significance of association with the DACT3 gene expression is plotted along the left y-axis. Dots are also color-coded based on their LD score with a tag SNP (rs78730097). The blue line indicates the estimated recombination rate, as plotted along the right y-axis. Gene expression levels detected by RNA-seq and RefSeq genes are plotted below the LocusZoom plot. b, Barplots showing the eQTL associations between the SNP rs78730097 and surrounding genes, showing the most significant association with the distal gene DACT3. Y-axis indicates –log10(eQTL association p value) obtained from GTEx database. c, d, Barplots showing FPKM values of distal target gene expressions upon deletion of core promoter regions of the ARIH2OS (f) and the ZNF891 (g). Two biological replicates of one mutant clone for the NCKIPSD (n=2) and two biological replicates of three mutant clones for the ZNF84 (n=6) were measured using RNA-seq, respectively. FDR-adjusted p value obtained from cuffdiff is shown together. N.S indicates statistically non-significant. e, Barplots showing FPKM values of two nearby genes of the ARIH2OS and one nearest gene of the NCKIPSD (y-axis) upon deletion of core promoter regions of the ARIH2OS. The ARIH2, a DNA fragment sharing gene with the ARIH2OS, is excluded. FDR-adjusted p value obtained from cuffdiff between WT and mutants for biological replicates (n=2) is shown together. Corresponding gene name is shown on the top of barplots. f, The relative mRNA expression levels of distal target genes (orange) and nearby genes (gray) of promoter-proximal eQTLs quantified by RT-qPCR are shown. Error bars indicate standard deviation from total six mutant clones for two separate sgRNAs with three mutant clones for each. Y-axis indicates mean values. One-sided KS-test p value is 6.14e-06 for the ABCF3 gene, 0.716 for the EIF4G1 gene, 6.14e-06 for the TOP3A gene, 0.862 for the LLG1 gene, 6.14e-06 for the SEC16A gene, 0.963 for the PMPCA gene, 0.552 for the SDCCAG3 gene, 1.00 for the ZFYVE21 gene (*** p value < 0.001).
Supplementary Figure 11 Identification of target genes of disease-associated genetic variants.
a, Illustration of the strategy to identify target genes of each GWAS trait. An example result is shown for Alzheimer disease. Both known and novel target genes were identified according to literature search. b, Venn diagram showing number of target genes by pcHi-C interactions and by nearby gene information for the GWAS-SNPs associated with Parkinson disease. c, Number of matched disease-associated genes in each group of target genes identified in Parkinson disease. d, Fraction of distal genes (blue) and nearby genes (gray) among the identified target genes of GWAS-SNPs based on pcHi-C interactions (left). Expected fraction is shown by calculating the fraction of nearby genes when we consider a nearest gene over 15kb as a GWAS-SNP target gene (right). e, Barplot showing the relative mRNA expression levels of GNL3 upon induced mutation of GWAS-SNPs with sgRNA as quantified by RT-qPCR as a control of MEP_L_fig4Fig. 4a. Y-axes indicates mean values and error bars indicate standard deviation of two mutant clones with technical triplicates. f, Barplot showing RT-qPCR results of relative target gene expression (y-axis) between mutant and control. Y-axes indicates mean values and error bars indicate standard deviation of two mutant clones with technical triplicates. The mutants showing more than 30% reduced expression of target genes are shown in orange. sgRNA target GWAS-SNP genomic coordinate, rsID, associated disease, distal target gene information, high LD SNP on coding region, and related publication PMID information are shown together.
Supplementary Figure 12 Analysis of the similarities in molecular genetics between human traits and diseases.
a, Illustration of the strategy to calculate the similarity between GWAS mapped traits using target gene similarity information. b, c, Shown are similarities, as measured by Pearson correlation coefficient (PCC), between traits (n=687) in the same order as Fig. 4b based on similarities of the putative GWAS-SNP target genes without shared promoters (b) and without genes located in HLA and HIST locus (c). The color intensity of each dot indicates Pearson correlation coefficient (PCC) of the putative target genes between two diseases or traits. d, Shown are similarities, as measured by Pearson correlation coefficient (PCC), between traits (n=687) based on the 5 nearest genes of the GWAS SNPs. The color intensity of each dot indicates PCC of target gene similarities between GWAS mapped traits. e, Barplots showing the fraction of number of TSS in a DNA fragment between all TSS and TSS corresponding genes in cluster 38 of Fig. 4b.
Supplementary information
Supplementary Information
Supplementary Figs. 1–12, Tables 1, 2, 5–11, 13, 15–18 and 21, and Note
Supplementary Table 3
List of all the significant P–O pcHi-C interactions
Supplementary Table 4
List of all the significant P–P pcHi-C interactions
Supplementary Table 12
List of P–O pcHi-C interactions and matched eQTL relationships
Supplementary Table 14
List of P–P pcHi-C interactions and matched eQTL relationships
Supplementary Table 19
List of putative target genes of GWAS SNPs
Supplementary Table 20
List of GWAS mapped traits and enriched GO biological processes in Fig. 4b
Rights and permissions
About this article
Cite this article
Jung, I., Schmitt, A., Diao, Y. et al. A compendium of promoter-centered long-range chromatin interactions in the human genome. Nat Genet 51, 1442–1449 (2019). https://doi.org/10.1038/s41588-019-0494-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0494-8
This article is cited by
-
Priority index for critical Covid-19 identifies clinically actionable targets and drugs
Communications Biology (2024)
-
Epigenetic profiling reveals key genes and cis-regulatory networks specific to human parathyroids
Nature Communications (2024)
-
Epigenomic insights into common human disease pathology
Cellular and Molecular Life Sciences (2024)
-
Dynamic enhancer landscapes in human craniofacial development
Nature Communications (2024)
-
Convergence of YAP/TAZ, TEAD and TP63 activity is associated with bronchial premalignant severity and progression
Journal of Experimental & Clinical Cancer Research (2023)