Abstract
Like many other crops, the cultivated peanut (Arachis hypogaea L.) is of hybrid origin and has a polyploid genome that contains essentially complete sets of chromosomes from two ancestral species. Here we report the genome sequence of peanut and show that after its polyploid origin, the genome has evolved through mobile-element activity, deletions and by the flow of genetic information between corresponding ancestral chromosomes (that is, homeologous recombination). Uniformity of patterns of homeologous recombination at the ends of chromosomes favors a single origin for cultivated peanut and its wild counterpart A. monticola. However, through much of the genome, homeologous recombination has created diversity. Using new polyploid hybrids made from the ancestral species, we show how this can generate phenotypic changes such as spontaneous changes in the color of the flowers. We suggest that diversity generated by these genetic mechanisms helped to favor the domestication of the polyploid A. hypogaea over other diploid Arachis species cultivated by humans.
Similar content being viewed by others
Main
The domestication of plants, thousands of years ago, increased food supply and allowed the formation of large, complex human societies. Out of many thousands of wild species, only a few became domesticated crops and they now provide most of the food consumed by humans. It has long been noted that many of these crops are polyploid: their nuclei have more than two sets of chromosomes that are often derived from different species. Although it has been surprisingly difficult to rigorously demonstrate, it has long been thought that domestication may favor polyploids1,2.
Peanut (also called groundnut; Arachis hypogaea L.) is an important food crop (annual production of ~44 million tons based on FAOSTAT data for 2016 (http://www.fao.org/faostat/en/#home)). Whereas almost all related species in the genus Arachis are diploid (two sets of ten chromosomes; mostly 2n = 2× = 20 chromosomes), A. hypogaea is polyploid3,4. The seeds of all of these species are an attractive food, and several have been cultivated for thousands of years5 (Supplementary Note 1). Indeed, the action of humans was key to the formation of A. hypogaea itself. About 9,400 years ago (estimated by nucleotide divergence6), the human transport of the ‘B’ genome species, A. ipaensis Krapov. & W.C. Greg., into the range of the ‘A’ genome species A. duranensis Krapov. & W.C. Greg. enabled their hybridization and the formation of A. hypogaea6. It has two sets of chromosome pairs, one from each of the ancestral species: a type of polyploid termed allotetraploid (AABB-type genome; 2n = 4× = 40 chromosomes; genome size of ~2.7 Gb).
The origin of A. hypogaea was associated with a particularly severe population bottleneck7,8,9. This could, in principle, have reduced the variability on which, over generations, human selection could act. However, A. hypogaea evolved, becoming completely dependent on cultivation and morphologically very diverse5. Two subspecies (hypogaea and fastigiata) and six botanical varieties (hypogaea, hirsuta, fastigiata, vulgaris, aequatoriana and peruviana) are recognized5,10,11. Different grain colors and sizes, pod shapes and growth habits distinguish thousands of landraces and cultivars5,11 (see also United States Department of Agriculture (USDA) Germplasm Resources Information Network (https://www.ars-grin.gov)). It seems notable that, in spite of the higher genetic diversity of the diploid species7,9, and their cultivation starting earlier (Supplementary Note 1), it was the derived allotetraploid, A. hypogaea, that underwent the transformation to become the crop of worldwide importance.
Some time ago, while planning to sequence and assemble the peanut genome, we realized that it would not be possible using the short-read data (~100–200 bp DNA) that were generated by the only technology that was economically feasible at the time; such sequences were too short to reliably resolve the very similar A and B genomes, which frequently have more than 98% DNA identity between corresponding genes6,12,13. This level of similarity is due to the progenitor species that gave rise to the two subgenomes having diverged only around 2.2 million years ago (refs. 6,9,14). Therefore, as a foundation for understanding the genome of cultivated peanut, we first sequenced the genomes of both the diploid ancestral species6. These diploid genomes afforded new insights into peanut genetics. Notably, it was possible to infer that some chromosome ends of A. hypogaea had changed from the expected AABB structure to AAAA or BBBB, implying a particular complexity in peanut genetics6,15,16,17,18.
Here, using the much longer-read data obtained with PacBio technology19, and scaffolding using Hi-C20,21, a method used for determining the conformation of DNA in the nucleus, we report the complete chromosome-scale genome sequence of A. hypogaea cv. Tifrunner, a runner-type peanut. We also characterize the genomes of a diverse selection of cultivated peanuts, together with its wild counterpart, A. monticola Krapov. & Rigoni, and induced allotetraploid hybrids derived from the ancestral species. We are able to visualize, in considerable detail, the products of variable deletions from, and genetic recombination between, the A and B subgenomes. It seems likely that these variations in genome structure generated phenotypic variation on which selection could act, and helped to favor A. hypogaea over its diploid relatives during the process of domestication.
Results
Sequencing and assembly of the peanut genome
Arachis hypogaea cv. Tifrunner22, a runner-type peanut (registration number CV-93, PI 644011) was sequenced using whole-genome shotgun sequencing. Twenty chromosome sequences were produced (for assembly metrics see Supplementary Tables 1 and 2). They were numbered Arahy.01–Arahy.20, where the A subgenome is represented as Arahy.01–Arahy.10 and the B subgenome as Arahy.11–Arahy.20. The chromosome sequences contain 99.3% of the assembled sequence and are 2.54 Gb, 93% of the size estimated by flow cytometry23.
Chromosome architecture
The chromosomes of A. hypogaea cv. Tifrunner largely reflect their ancestral structures; the homeologous chromosomes mostly have a one-to-one correspondence: Arahy.02/12, 03/13, 04/14 and 10/20 are almost completely collinear; 06/16 and 09/19 are differentiated by a large inversion in one arm; 05/15 are differentiated by two large inversions; and 01/11 are differentiated by three large inversions. Chromosomes 17/18 have undergone reciprocal translocations relative to 07/08 (Supplementary Figs. 1–12). Gene densities are highest in distal chromosome regions (Supplementary Fig. 13). Gene counts are 11% higher in the B subgenome, with 35,110 predicted genes, compared to 31,359 genes in the A subgenome. Long terminal repeat (LTR) retrotransposons are highly abundant in pericentromeric regions, whereas DNA transposons are more frequent in euchromatic arms (Supplementary Fig. 14). Other transposable elements, together with approximately 3,300 pararetrovirus sequences account for 74% of the assembled genome sequence (Supplementary Tables 3 and 4). Notably, this compares to 64% repetitive content estimated by reassociation kinetics24, indicating the high quality and relative lack of collapse of repeats in this long read-based assembly. The chloroplast genome of A. hypogaea and a chloroplastic plasmid were inherited from A. duranensis (Supplementary Fig. 15).
DNA methylation and small RNAs
Genic methylation patterns were typical for plants, with lower methylation in transcribed regions and characteristic dips in methylation at transcription start and end sites (Supplementary Fig. 16). Genome-wide methylation per cytosine content was higher in pericentromeric regions than chromosome arms (Supplementary Fig. 17). Methylation was lower in the A subgenome than the B subgenome; with 76.0% and 80.5% methylation at CG sites, 61.7% and 65.1% methylation at CHG sites (where H is an A, T or C) and 5.14% and 5.51% methylation at CHH sites, respectively (Supplementary Table 5 and Supplementary Fig. 18a). Greater densities of DNA sequences corresponding to small RNAs were found in proximal, repetitive-rich regions of chromosomes (Supplementary Fig. 19). However, greater densities of DNA sequences that corresponded to uniquely mapping small RNAs were found in gene-rich chromosomal regions (Supplementary Fig. 20). Within genes, the B subgenome was enriched relative to the A subgenome for DNA sequences that corresponded to small RNAs (Supplementary Fig. 18b).
Comparison of gene expression in subgenomes
The expression of homeologous gene pairs (dataset 1a in ref. 25) from the A and B subgenomes of Tifrunner was investigated in diverse tissues and developmental stages (dataset 1b,c in ref. 25). As has been reported in other recent polyploids26,27, overall, the number of homeologous gene pairs with expression biased towards the A subgenome was not significantly different from the number biased towards the B subgenome (P = 0.2, two-sided binomial test; n = 3,648 and 3,759 for A and B, respectively). However, when tissues were considered separately, all but one had slightly more B than A subgenome-biased genes from homeologous pairs. In three reproductive tissues and in roots this difference was significant (P < 0.05, one-sided binomial test; Supplementary Fig. 21; dataset 1 in ref. 25).
Broadly, homeologous pairs with the highest asymmetry in expression (log2(expression ratios) > 3, Benjamini–Hochberg-adjusted P < 0.05, Wald test; Supplementary Fig. 22) were more commonly involved in oxidation–reduction processes, pollen recognition, lipid and chitin metabolic processes and response to biotic stimulus (Supplementary Fig. 23a; dataset 1c in ref. 25).
Taking the example of the subterranean peg tip (a unique reproductive structure in peanut), the A subgenome-biased homeologous pairs were enriched for genes involved in mannose metabolic processes, nitrate assimilation and cell wall assembly, whereas the B subgenome-biased homeologous pairs were enriched for genes involved in the response to biotic stimulus, sucrose transport and glucan metabolic processes. In the maturing pericarp (Pattee stage 6), the A subgenome-biased homeologous pairs were enriched for genes involved in phosphorylation signal transduction, carbohydrate metabolism and cell wall biogenesis, whereas B subgenome-biased homeologous pairs were enriched for genes involved in inorganic ion transport and response to biotic stimulus (dataset 1d,e in ref. 25). Additionally, we identified homeologous gene pairs with the highest asymmetry in expression (n = 4,062; log2(expression ratios) > 3, Benjamini–Hochberg-adjusted P < 0.05, Wald test; Supplementary Fig. 22) and a set of 394 pairs that displayed consistent asymmetrical expression patterns in at least half of the evaluated tissues (Supplementary Fig. 23b). Highly asymmetrically expressed homeologous pairs were more commonly involved in oxidation–reduction processes, pollen recognition, lipid and chitin metabolic processes and response to biotic stimulus (Supplementary Fig. 23a, dataset 1c in ref. 25) and, as might be expected, the consistently asymmetrically expressed homeologous pairs were mainly enriched for functions associated with fundamental biological processes such as organelle organization, molecular transport and protein complex biogenesis (dataset 1c in ref. 25).
Changes following polyploidy
Genetic exchange between subgenomes and deletions
For allotetraploids, chromosome associations during meiosis and genetic exchange are mostly limited to corresponding chromosomes within the same subgenome (that is, homologous chromosomes); however, as has been characterized in other plants such as Brassica26,28,29, these may also occur at lower frequency between corresponding chromosomes from the other subgenome (that is, homeologous chromosomes)3,6,16. We investigated genetic exchange between the subgenomes and deletions in more than 200 diverse genotypes comprising the wild tetraploid peanut (A. monticola), landraces and cultivars of A. hypogaea, and new allotetraploid hybrids made from the ancestral species (dataset 2 in ref. 25). Two different approaches were used: observation of mapping densities of short-read whole-genome sequences onto the combined sequenced diploid ancestral species genomes, and analysis of the short-read whole-genome sequences for single-nucleotide polymorphisms (SNPs) that consistently differentiate representatives of A and B genome diploid species5,9,30,31,32 (Supplementary Fig. 24). (It should be noted that, except for the assembled reference genotype of Tifrunner, these methods are not capable of detecting genome changes that result from balanced homeologous exchanges or chromosome rearrangements.)
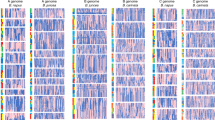
Genetic exchange between ancestral genomes could be inferred towards the ends of colinear pairs of homeologous chromosomes. In these regions, the genome structure was not the expected AABB, but may be better described as AAAA or BBBB, that is, ‘tetrasomic’ conformations. The abrupt junctions of these segments indicate that they may have occurred by crossover (Figs. 1 and 2 and Supplementary Figs. 1–12 and 25; datasets 3–5 in ref. 25). In Tifrunner, 14.8 Mb of the A genome has been transferred, in blocks, into B chromosomes, and 3.1 Mb of the B genome has been transferred, in blocks, into A chromosomes (Supplementary Tables 6 and 7). Most of these tetrasomic regions are at the very distal ends of chromosomes—for example, the lower regions of Arahy.02/Arahy.12, Arahy.04/Arahy.14, Arahy.06/Arahy.16 and the upper regions of Arahy.05/Arahy.15—and these were present in all of the A. hypogaea and A. monticola genotypes surveyed (but not in induced allotetraploids derived from the same diploid ancestral species; Fig. 1; dataset 4a,b in ref. 25). However, in slightly more proximal regions, the tetrasomic regions were variable. Notably, in different accessions, in some genome regions, genetic exchange had occurred in opposite directions, creating AAAA structures in some accessions, and BBBB structures in others (Fig. 2 and Supplementary Fig. 25; dataset 4a,b in ref. 25). Although clearly identifiable as A or B, these tetrasomic regions contain a significant number of SNPs that are characteristic of the corresponding subgenome (Supplementary Table 7). This may be the result of genetic exchange by gene conversion prior to the large-scale transfer of genetic material between subgenomes. The fine-scale patterns of these SNPs represent substantially fixed, or fossilized, genetic signals (‘fingerprints’) from past events. Their uniformity in all six botanical varieties and the wild counterpart of peanut A. monticola favors a single polyploid origin for the two species (Fig. 1f,g; dataset 5 in ref. 25). In Tifrunner, chromosome segments transferred between subgenomes mostly form tetrasomic regions, although one region at the lower end of Arahy.16 contains a chromosome segment with predominantly ancestral A genome characteristics that is absent from Arahy.06 itself (Supplementary Fig. 6). This region on Arahy.16 forms a peculiar structure in which B and A homeologous chromosome segments are retained in tandem.
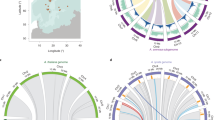
a–e, Overviews of genetic exchange between ancestral A and B genomes; f,g, visualization of fine-scale exchange at the ends of chromosomes. In a–e, data are of log2-transformed values of ratios of mapping densities of whole-genome sequences onto 17,373 orthologous A/B gene pairs from A. ipaensis and A. duranensis, ordered according to chromosome number and position in A. ipaensis. Where values cluster around zero, as is the case in the diploid hybrid in a, A and B genes are present in equal number and are unaltered by genetic flux between them; in tetraploid genotypes this indicates a genome structure of AABB. Deviations from zero indicate genetic flux between the orthologous gene pairs, or complete replacement of A genes by B, or vice versa. b, The ninth generation tetraploid hybrid shows such deviations, with a change in genome structure from AABB to AAAA for chromosomes A04/B04 and the upper regions of B07/A08. c–e, These patterns are very different from those of A. monticola and A. hypogaea, which are similar to each other (and throughout diverse genotypes). Note deviations are mostly at chromosome ends. f,g, Fine-scale recombination (fingerprints) between A and B subgenomes are shown in two distal chromosome regions in which the genome structure approximates AAAA; the presence of SNPs characteristic of the ancestral B that form barcode-like patterns that are uniform in all A. monticola and A. hypogaea are observed. These patterns emphasize the similarities between A. monticola and A. hypogaea and favor a single polyploid origin (Supplementary Fig. 25; dataset 4a in ref. 25).
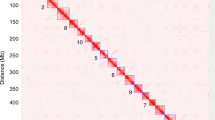
a–c, Genetic exchange between Arahy.04 and Arahy.14 in three A. hypogaea, visualized by mapping the densities of short-read whole-genome sequences onto A. duranensis and A. ipaensis (normalized values, in blue and red respectively; distances scaled to Arahy.14). Where mapping densities cluster around the expected value of one, the genome composition is AABB. Where mapping densities on one genome increase to approximately two and on the other genome decrease to near zero, the genome composition is better described as AAAA or BBBB. Mapping densities decrease to around zero on one genome and remain around one on the other indicate a deletion (common in b). d, A panel of 68 representative diverse genotypes in a region of Arahy.04 and Arahy.14 in which hypervariability has been created by differential recombination between subgenomes. The panel represents a heat map of log2-transformed ratios of mapping densities on the B and A genomes; blue represents AAAA and red represents BBBB, Tif., PI and Chib. are the genotypes repesented in a–c, respectively (for full visualizations, see dataset 4b in ref. 25).
The signals of disperse genetic exchange were also detectable through the bodies of chromosomes. Overall, this dispersed genetic exchange has had a greater total effect than the transfer of chromosome segments. In Tifrunner, almost twice as many B alleles have been transferred to A chromosomes than vice versa (Supplementary Table 6; dataset 3 in ref. 25). In addition, variable deletions were frequent in proximal chromosome regions (Fig. 2; dataset 4c in ref. 25). Notably, a large deletion (around 10 Mb) was common on Arahy.14 of botanical varieties fastigiata and vulgaris (e.g., Fig. 2b).
In Tifrunner, genome deletions have disproportionately affected some gene families. The genes most frequently lost were members of the serine/threonine-protein phosphatase (around 89 genes) and FAR1-related families (around 83 genes). Genes in these families tend to occur in large genomic clusters or arrays, which can expand or contract through slipped-strand mispairing33. There have also been apparent increases in gene families; these include an increase of around 118 SAUR-like auxin-responsive protein family genes and around 50 NBS-LRR–encoding genes (the latter family of genes encode plant nucleotide-binding-site leucine-rich repeats and are associated with pest and disease resistance). For an overview of loss of ancestral SNP alleles through homeologous recombination and deletions in 39 diverse genotypes, see Supplementary Fig. 26.
Mobile-element activity
Transposable elements generate extrachromosomal circular DNAs when active34. Circular DNAs were detected from a MUTATOR (MU4) and TY3-GYPSY (ZUHE) element in A. duranensis, A. ipaensis, their hybrids and A. hypogaea, and from a TY1-COPIA element (YARA) in A. duranensis and A. hypogaea. However, no abundant circular DNAs were detected in induced allotetraploids or A. hypogaea that were not detected in one or both of the ancestral diploids (Supplementary Fig. 27). This indicates that after hybridization and polyploidy, somatic transposable element regulation was not heavily disturbed and no new large-scale mobilization of transposable elements occurred. Comparisons of genome sequences of the ancestral species and A. hypogaea support this; we could not identify any large-scale insertions. Consistent with previous findings35, most newly inserted elements are MUTATOR-like elements (Supplementary Fig. 28).
Inversions
Comparisons of Tifrunner subgenomes showed three more major chromosome inversions than were observed when comparing the sequenced accessions of the two ancestral diploid species: two in the A subgenome, on Arahy.05 and Arahy.07, and one in the B subgenome, on Arahy.11 (Supplementary Figs. 1, 5 and 8). We consider it likely that at least two of these three extra inversions were already present in the diploid ancestors. The alternative chromosomal arrangement of Arahy.07 is indicated by a genetic map derived from a cross of two different A. duranensis accessions (see the genetic map of a previously published study36, which is presented relative to the sequenced genome of A. duranensis V14167 in the supplementary dataset of another study6). Furthermore, significantly higher DNA identity between Arahy.07 and five A. duranensis accessions (including the closest ones to the A subgenome ancestor; see below) is observed when compared to others (Supplementary Tables 8 and 9). Similarly, for Arahy.05, markedly higher identities to three A. duranensis accessions may indicate the presence of the inversion in some representatives of A. duranensis, possibly including the ancestral A subgenome donor (Supplementary Table 8 and 9).
We previously reported that inversions move repeat-rich DNA to more distal chromosome regions where DNA is lost by recombination, thus reducing genome size (although regions moved to more proximal positions gain DNA, this effect is smaller)6,37. Following this pattern, the inverted region in Arahy.05 has shrunk relative to A. duranensis V14167 (tetraploid size/diploid size = 0.89; Supplementary Table 10). We found that removal of LTR retrotransposons is the predominant cause of this reduction (Supplementary Fig. 29). Furthermore, the presence of repeats in A. duranensis, at the ends of the regions, which are missing in A. hypogaea, clearly implicate unequal intrastrand recombination in about 20% of cases (107 out of 502 regions). By contrast, there is little difference in relative sizes of the inversions on Arahy.07 and Arahy.11.
Observations of independent polyploidy events
We used allotetraploids derived by colchicine treatment of hybrids of the peanut’s ancestral diploid species38 to investigate genome changes that followed independent polyploidy events. We studied 37 different lineages from two independent induced polyploidy events. Genetic exchange between subgenomes occurred in large blocks and interspersed alleles along chromosome segments; these events seem at least partly stochastic, and were different between different lineages and from A. hypogaea. Spontaneous changes in flower color in some lineages (Fig. 3) could be ascribed to genetic exchange between subgenomes; the A genome region that confers the yellow flower color had been replaced by the homeologous B genome region that confers orange flower color (dataset 6 in ref. 25). This provides a simple demonstration of phenotypic change as a consequence of genetic exchange between subgenomes.
The initial allotetraploid, A. ipaensis × A. duranensis (2n = 4× = 40) has yellow flowers (left), as expected. However, after several generations some lineages spontaneously began to bear orange flowers (right). By genotyping, this could be assigned to homeologous recombination, where in alleles that confer yellow flowers (from A. duranensis) are replaced by alleles that confer orange flowers (from A. ipaensis; see dataset 6 in ref. 25). Scale bar, 5 mm.
A closer representative of the A subgenome ancestor
Because their seeds develop underground, wild Arachis populations are unusually static over time5. In addition, they typically have very high rates of self-pollination. This, and a serendipitous collection by pioneering botanical collectors, enabled our previous discovery that the sequenced A. ipaensis K30076 was very likely a descendant of the same population that donated the B subgenome to A. hypogaea6. Here we endeavored to identify the extant A. duranensis population that is closest to the A subgenome donor. We characterized 55 accessions, representative of all known major populations of A. duranensis, by sequencing DNA enriched for genic regions (using exome capture methods). A selection of these accessions was chosen for whole-genome re-sequencing. The A. duranensis accessions that were most similar to the Tifrunner A subgenome were from Rio Seco (Argentina), a location previously indicated as the likely origin of the A subgenome ancestor on the basis of chloroplast and ribosomal DNA haplotypes39 (Fig. 4 and Supplementary Tables 8, 9 and 11). However, in some cases, the ranking of similarity changed by chromosome (especially for Arahy.05), possibly reflecting variations in chromosomal arrangements in different accessions of A. duranensis (as discussed above; Supplementary Table 9). Comparisons of the Tifrunner A subgenome with the whole-genome sequences of A. duranensis accessions indicated median DNA identities of 99.76% for the Rio Seco accessions (KGBSPSc 30065, PI 468201 and KGBSPSc 30067, PI 468202); 99.61% for the sequenced V14167 (ref. 6); and 98.23% for PI 475845 from the northern range of the species and with a partially assembled genome40 (Supplementary Table 8 and Supplementary Fig. 30; dataset 7 in ref. 25).
Genomic DNAs of 55 accessions, representing all known major populations of A. duranensis, were compared to the A subgenome of Tifrunner. Similarity is strongly influenced by hydrographic basins. Accessions with the highest similarity (in red) are concentrated around Rio Seco, a tributary of the Rio San Francisco; next in similarity (in orange) are accessions concentrated around Jujuy, a region that drains into the Rio San Francisco and the Lerma valleys; followed by accessions from the Rio Juramento (in light brown), a region that receives water from the Lerma valley. Following these in similarity are accessions from the endorreic basins that occasionally drain in the Bermejo River (northwest Argentina and south Bolivia) (in yellow) followed by accessions in the basins of the Rio Pilcomayo (in light green) and finally accessions from the Rio Parapetí basin, Izozog Swamps and West Paraguay sand dunes (in dark green). Outliers to this general pattern are likely to represent populations that have resulted from the occasional human movement of seeds among basins (most of these movements are likely to have occurred long ago). The maps were generated using Natural Earth.
The A subgenome chromosomes are, in general, less similar to their A. duranensis counterparts than the B subgenome chromosomes are to their A. ipaensis counterparts. This is consistent with the greater flow of alleles from the B subgenome into the A subgenome than vice versa (as described above, see also a previously published study6).
Discussion
A genome sequence is a landmark for the research of the biology of a crop. It provides a catalog of gene content, with chromosomal context and a unified framework for biological investigations and cross-species comparisons. In the case of peanut, a polyploid of recent hybrid origin, the previous sequencing of very close representatives of its diploid ancestors provides the opportunity to investigate more generally applicable principles regarding the genetics of polyploidy and its importance to crop domestication.
Polyploidy has long been recognized as an important feature of plant evolution; it has occurred multiple times during the evolution of almost all flowering plants. Following each polyploidy event, over tens of millions of years, deletions, divergence of duplicated genes and rearrangements return the genome to a diploid state. The recurrence of these ‘wondrous cycles’ is thought to have played an important part in diversification and adaptation during plant evolution41,42,43. It has also long been recognized that many crop plants are recent polyploids; and, although the matter has generated decades of debate, it does seem that polyploids are favored for domestication1,2.
We consider the evidence that polyploid A. hypogaea was favored for domestication over its diploid relatives very persuasive. Archaeological remains and remnant populations of Arachis species far from their natural distributions, and the existence of a diploid domesticated species (A. villosulicarpa) testify to widespread and large-scale cultivation of at least four diploid species (Supplementary Note 1). Indeed, the hybridization that gave rise to A. hypogaea was only possible because of human transport of A. ipaensis into the range of A. duranensis6. It seems important that, in spite of higher genetic diversity of the diploid species and their cultivation having started earlier, it was—in fact—the allotetraploid A. hypogaea that became the crop of worldwide importance.
Following trends seen in many plants, Arachis allotetraploids are larger than their diploid progenitors. The tetraploids also have different transpiration characteristics44 and produce more photosynthetic pigments45. These traits—or other ploidy-related changes—may have been advantageous; however, contrary to common expectations, the seeds of the allotetraploid ancestor of peanut seem likely to have been similar size to those of its diploid progenitors45. The increased number of alleles associated with being a ‘fixed hybrid’ would have increased heterosis and therefore probably adaptability. However, the extreme genetic bottleneck that accompanied the polyploid origin may have been expected to reduce variability on which artificial selection could act. We investigated genome changes after polyploidy that could have generated variation. We found no evidence for widespread mobilization of transposable elements (Supplementary Fig. 27). However, we could identify some mobile element insertion polymorphisms and some of these are likely to have influenced gene activity (Supplementary Fig. 28). In addition, variable deletions, especially in proximal chromosome regions, have occurred since polyploidy and these also must have generated variation. However, it was a different genetic phenomenon, associated with harboring full chromosome complements from two species, that most drew our attention: genetic exchange between subgenomes3,6,15,16,26,28,29.
We identified two patterns of homeologous recombination. One involves the transfer of chromosome segments between distal collinear regions of chromosomes mostly resulting in tetrasomic genome structures (AAAA and, to a lesser extent, BBBB; Figs. 1 and 2 and Supplementary Figs. 1–12, 25). The other involves transfer of dispersed alleles that has occurred throughout the chromosomes; it is strongly biased, with much more transfer of alleles from B subgenome to A subgenome (Supplementary Tables 6 and 7). Overall, the genetic flux seems to have caused a greater erosion of similarity of the A subgenome to its progenitor A. duranensis than of the B subgenome to its progenitor A. ipaensis (even though the distal regions of the B chromosomes are more invaded by segments of the A genome than vice versa). Collections from Rio Seco were the closest representatives of the A subgenome ancestor, although several accessions from Salta (including the sequenced V14167 (ref. 6)) showed quite similar degrees of identity (Fig. 4, Supplementary Tables 8, 9 and 11 and Supplementary Fig. 30).
On the whole-genome scale, the effects of homeologous recombination appear similar in diverse peanut accessions. Most of the tetrasomic structures were present in all A. hypogaea and A. monticola analyzed; furthermore, fingerprint-like fine-scale patterns of interspersed homeologous alleles within the distal tetrasomic regions were also found to be uniform (Fig. 1; datasets 4a and 5 in ref. 25). By contrast, homeologous recombination patterns in allotetraploid hybrids were completely distinct (Fig. 1; dataset 4a,b in ref. 25). This emphasizes the close relationship of A. hypogaea and A. monticola, and favors a single polyploid origin of both species. However, when observed on a finer scale in other genome regions, it becomes apparent that homeologous recombination in A. hypogaea has generated new diversity (Fig. 2). Some tetrasomic regions differ in different accessions of A. hypogaea; in certain genome regions some peanut accessions have an AAAA genome structure, whereas others have BBBB (Fig. 2 and Supplementary Fig. 25). Our observation for flower color, although a simple trait, provides a proof-of-principle link between homeologous recombination and generation of phenotypic diversity (Fig. 3; dataset 6 in ref. 25).
In summary, we determined the genome sequence of one reference peanut cultivar, and surveyed the genome structures of a diverse sample of landraces and cultivars. The genome structure of peanut is segmental allotetraploid (as defined by Stebbins46). We suggest that genetic deletions and exchange between the subgenomes generated variation that helped to favor the domestication of A. hypogaea over its diploid relatives. These results highlight a possible wider importance of these genetic mechanisms in accounting for the higher than expected frequency of polyploids in domesticated plants.
Methods
Plant material for genome sequencing
To generate the reference genome, we used A. hypogaea cv. Tifrunner22, a runner-type peanut adapted for the southeast of the United States (CV-93, PI 644011). Phenotypically, Tifrunner would be classified as A. hypogaea ssp. hypogaea var. hypogaea, although, like almost all runner peanuts that are currently grown in the southeast United States, it has cultivars termed ‘Spanish’ in its pedigree (Supplementary Fig. 31). Plants grown in an isolation plot in 2005 were genotyped with 146 simple sequence repeat DNA markers positioned on all 20 chromosomes. A line with no detected heterozygosity was used as the founder of the peanut genome project stock.
Sequencing of the reference tetraploid genome
We sequenced Arachis hypogaea cv. Tifrunner using a whole-genome shotgun sequencing strategy and standard sequencing protocols. Illumina and PacBio reads were produced at USDA ARS GBRU and the HudsonAlpha Institute. Illumina reads were produced using the Illumina HiSeq platform and the PacBio reads were generated on the RSII platform. Two 800-bp insert 2× 250 Illumina fragment libraries were obtained for a total of 63.09× coverage. Before use, all Illumina reads were screened for mitochondria, chloroplast and PhiX contamination. Reads composed of >95% simple sequences were removed. Illumina reads that were <75 bp after trimming for adapter and quality (Q < 20) were removed. An additional deduplication step was performed on the Illumina mate pairs that identifies and retains only one copy of each PCR duplicate. These two Illumina libraries were used in the final polishing of homozygous SNPs and insertions and deletions (indels) in the consensus sequence. For the PacBio sequencing, high-molecular weight DNA was isolated at the Arizona Genomics Institute (https://www.genome.arizona.edu), a total of 301 chips (P6C4 chemistry) were sequenced with a total yield of 207.2 Gb (76.74×) and after error correction a total of 130.27 Gb (48.25×) was used in the assembly (Supplementary Tables 12 and 13).
Genome assembly and construction of pseudomolecule chromosomes
The 17,747,748 PacBio reads (76.74× sequence coverage) were assembled using MECAT47. This produced 7,692 contigs with an N50 of 696.6 kb, 4,778 larger than 100 kb and a total genome size of 2,502.6 Mb (Supplementary Table 14). The resulting assembly was polished using Quiver48. Three genetic maps (see below; dataset 8 in ref. 25) were used to identify potential misjoined regions in the MECAT assembly. Synteny with A. duranensis and A. ipaensis diploid references was then used to pinpoint breakpoints. A total of 856 potential misjoined regions were identified and broken.
Hi-C scaffolding
The broken assembly was then scaffolded with Hi-C data using the 3D-DNA pipeline20. We prepared two in situ Hi-C libraries as previously described49 and sequenced them (library 1: 62,762,161 of PE85 and 114,895,839 of PE150 reads; library 2: 228,896,977 of PE150; Supplementary Table 12). The Hi-C reads were aligned to the broken assembly using the Juicer pipeline50,51. The 3D-DNA pipeline was run with the following parameters: --editor-saturation-centile 10 --editor-coarse-resolution 100000 --editor-coarse-region 400000 --editor-repeat-coverage 50. The results were polished using the Juicebox Assembly Tools—an assembly-specific module in the Juicebox visualization system52. The Hi-C scaffolding resulted in 20 chromosome-length scaffolds.
Construction of pseudomolecule chromosomes
An additional set of six breaks were made after scaffolding. Scaffolds were then oriented, ordered and joined together into 20 chromosomes. A total of 17 joins were made during this process, these chromosome joins were padded with 10,000 N characters. Telomeric sequences were identified using the (TTTAGGG)n repeat and care was taken to ensure correct orientation. Chromosomes were named according to subgenome, with the A subgenome being denoted as Arahy.01–Arahy.10 and the B subgenome denoted as Arahy.11–Arahy.20. Seven regions at the ends of chromosomes had autotetraploid-like genome structures and, as expected, were represented only once instead of twice. These were identified by twofold higher mapping densities on one subgenome of Tifrunner Illumina sequence reads (dataset 9 in ref. 25). These seven regions, covering 16.6 Mb, were identified (Supplementary Table 15), duplicated and added into the other subgenome. A total of 99.30% of the assembled sequence is represented in the chromosomes.
The combined assembly was then screened for contamination. Homozygous SNPs and indels were corrected in the release sequence using ~60× of Illumina reads (2× 250, 800-bp insert size) by aligning the reads using BWA MEM53 and identifying homozygous SNPs and indels with the GATK’s UnifiedGenotyper tool54. A total of 90 homozygous SNPs and 134,361 homozygous indels were corrected in the release sequence. The final version contains 2,552.5 Mb of sequence, consisting of 384 scaffolds (4,037 contigs) with a contig N50 of 1.5 Mb and a total of 99.3% of assembled bases in chromosomes. A Hi-C contact map visualization of the completed assembly is shown in Supplementary Fig. 32.
Assessment of assembly accuracy
A set of 223 bacterial artificial chromosome (BAC) clones (20.8 Mb) were sequenced in order to assess the accuracy of the assembly. DNAs were extracted individually and then pooled into sets of 96 BACs. PACBIO RS II was used for sequencing with a targeted 100× depth. Assembly of the pools was performed using HGAP3 (version 2.3.0) followed by consensus sequence calling with Quiver (version 2.1). Vectors were identified and trimmed, clones recircularized and repolished with Quiver to obtain the final BAC contigs. A range of variants were detected in the comparison of the BAC clone contigs and the genome assembly. Two of the BAC contigs were excluded, because they aligned to highly repetitive pericentromeric regions, and 46 of the contigs were excluded based on length (≤20 kb), leaving 175 contig alignments for analysis. Of these, a total of 79 alignments were of high quality (<0.1% bp error; Supplementary Fig. 33); dot plots were generated using Gepard55. The next 86 BAC contigs indicate a higher error rate, which was mainly due to their placement in more repetitive regions (Supplementary Fig. 34). The final ten BAC contigs indicate putative overlaps on adjacent contigs within a chromosome (Supplementary Fig. 35). The overall bp error rate (including marked gap bases) in the BAC clone contigs is 1 error per 33,510 bp (431 discrepant bp out of 14,442,956).
Mapping populations, genotyping and linkage maps
The A. hypogaea cv. Tifrunner × A. hypogaea GT-C20 population was composed of 91 F8 individuals derived by single-seed descent and was used for mapping, whole-genome sequencing and marker calling as previously described56. Joinmap 5.0 was used for genetic map construction after selecting markers without segregation distortion (χ2 test; P > 0.05; 1:1 ratio of alleles), using the Kosambi mapping function and a minimum logarithm of odds score for linkage of 10.
The A. hypogaea cv. Runner IAC 886 × (A. ipaensis K30076 × A. duranensis V14167)(2n = 4× = 40) population consists of 89 F6 individuals that were derived by single-seed descent. The linkage map has been described previously6.
The A. hypogaea cv. Runner IAC 886 × (A. batizocoi K9484 × A. stenosperma V10309)(2n = 4× = 40) consists of a population of 196 F2 individuals. Genotyping for SNPs was done using the Affymetrix genotyping array16,57. Maps were constructed using the Kosambi function in Mapdisto58 version 2.0; 20% of missing data was allowed, with a minimum logarithm of odds score of 20 and a maximum recombination frequency of 0.30.
Identification of repetitive DNA
Mobile elements were identified using a number of homology and de novo structural pattern-finding algorithms and manual curation; see Supplementary Note 2.
Structural comparisons of chromosomes
Structural comparisons between chromosomes were generated and visualized using the MUMmer suite of alignment tools59.
Assembly of transcripts and gene annotation
A transcriptome assembly to support annotation was generated from more than 6.4 billion cleaned sequence reads from A. hypogaea ssp. hypogaea genotypes (Supplementary Table 16). Libraries were constructed and 100- or 125-bp paired-end sequences generated following recommendations of the manufacturer (Illumina). Assembly was carried out with Trinity using the tetraploid genome as a guide. Read redundancy was first reduced with Trinity in silico normalization, with --max_cov 100, giving 97 million normalized reads. The normalized reads were aligned to the Tifrunner genome assembly using gsnap60 and then assembled using Trinityrnaseq61 version 2.5.0, with maximum intron size of 10,000, and k-mer minimum coverage of 3. After filtering transcript assemblies using Kallisto62 (transcripts per million of 1.5; 90,519 assembled transcripts were retained, with an average size of 1,540 nucleotides.
The A. hypogaea cv. Tifrunner genome was annotated using the MAKER pipeline63 version 2.31.9 (specifically, the dockerized image maker-2.31.9-3.img run under singularity 2.4). The genome sequence was hard-masked for ‘complex repeats’ (for example, transposable elements) using RepeatMasker and a library of repeat sequences identified in A. duranensis and A. ipaensis6 and A. hypogaea (this manuscript). Simple repeats were soft-masked by MAKER, allowing them to be accessible for gene annotation in some cases. Ab initio gene prediction methods used within MAKER included SNAP64 version 2006-07-28 and AUGUSTUS65 version 3.2.3. Arachis-specific model parameters for the ab initio predictors were obtained initially from gene model calls made against chromosomes Arahy.01 and Arahy.11 (representing contributions from the two diploid progenitor species) by using only the highest-confidence gene models produced in a first iteration of the pipeline (annotated edit distance ≤ 0.25); this subset was used to train the predictors for the model parameters used in subsequent iterations of the full annotation process (four iterations in total). Protein sequences used as queries for homology-based predictions consisted of the Uniprot Fabaceae protein set (retrieved December 2017). Nucleotide sequences used as queries for homology-based predictions consisted of two transcriptome assemblies generated from A. hypogaea Tifrunner: the genome-guided transcriptome assembly described above, and the 22-tissue transcriptome assembly that has been described previously66. Provisional functional assignments for the gene models were produced using InterProScan67 and BLASTP68 against annotated proteins from Arabidopsis thaliana, Glycine max and Medicago truncatula, with outputs processed using AHRD (https://github.com/groupschoof/AHRD), for lexical analysis and selection of the best functional descriptor of each gene product.
Comparison of gene expression in subgenomes
Paired-end sequencing data from expressed RNA66 was quality trimmed (Q ≥ 25) and reads shorter than 50 bp after trimming were discarded. Sequences were then aligned to the A. hypogaea cv. Tifrunner genome and counts of reads uniquely mapping to annotated genes were obtained using STAR69 version 2.5.3a. Outliers among the individual experimental samples were verified based on the Pearson correlation coefficient, r2 ≥ 0.85. Fragments per kilobase of exon per million fragments mapped values were calculated for each gene by normalizing the read count data to both the length of the gene and the total number of mapped reads in the sample and considered as the metric for estimating gene expression levels70. Normalized count data was obtained using the relative log expression (RLE) method in DESeq2 (ref. 71) version 1.14.1. Genes with low expression were filtered out, by requiring ≥2 RLE-normalized counts in at least two samples for each gene.
High-confidence homeologous gene pairs were initially identified by their reciprocal highest scores in similarity searches (BLAT) of all annotated genes in each Tifrunner subgenome versus the other. We also applied the criteria of a minimum of 80% nucleotide identity and 80% sequence length coverage and only considered gene pairs that reside on homeologous chromosomes and established reciprocal translocations (dataset 1a in ref. 25). We performed differential expression analysis between the genes in homeologous pairs for each tissue and pod developmental stage using DESeq2 (version 1.14.1) with log2-transformed expression ratio ≥1 and Benjamini–Hochberg-adjusted P < 0.05 as the statistical cut-off for asymmetrically expressed genes. We used Gene Ontology for functional analysis of asymmetrically expressed homeolgous gene pairs. To determine overrepresented Gene Ontology categories across biological processes, cellular component and molecular function domains, topGO72,73, an R Bioconductor package was used. Enrichment of Gene Ontology terms was tested using Fisher’s exact test with P < 0.05 considered as significant. Statistical analyses and visualizations were performed using the R version 3.4.1 statistical software (R Development Core Team 2011).
DNA methylation
Genomic DNA was isolated from whole young unexpanded leaves using the DNeasy Plant Mini Kit (Qiagen). MethylC-sequencing libraries were constructed as previously described74. In brief, approximately 1 μg of genomic DNA spiked with about 10 ng of unmethylated lambda DNA was sonicated to around 200 bp using a Covaris S-2. Size selection was performed using magnetic purification beads. The End-It DNA End-Repair Kit (Epicentre) was used to perform end repair on the fragmented DNA. A-tails were added to blunt-end fragments using Klenow 3′–5′ exonuclease and dA-Tailing Buffer (New England Biolabs). Methylated NEXTflex DNA adapters (Bio Scientific) were then ligated onto the DNA using T4 DNA ligase (New England Biolabs). Bisulfite conversion was done using the MethylCode Bisulfite Conversion Kit (Invitrogen). Finally, eight rounds of PCR using Kapa HiFi Uracil and Hotstart DNA polymerase (Kapa Biosystems) was used to amplify the libraries. Between each reaction, magnetic purification beads were used to clean up the DNA. Libraries were sequenced on an Illumina HiSeq 2500.
Quality-trimmed reads were aligned to the A. hypogaea cv. Tifrunner genome using Bismark75 version 0.7.0. Multiple mapped reads and clonal reads that corresponded to potential bias from PCR amplification were discarded. The first and last 5 bp of each read where masked before methylation calling to remove biases in methylation levels introduced during the end-repairing step of library preparation. Cytosine methylation levels were calculated using the binomial distribution as previously described76. The bisulfite non-conversion rate was calculated by mapping the unmapped reads to the unmethylated lambda genome. Only cytosines covered by at least three reads in at least one of the two replicates were retained and the two replicates were then merged for further analysis.
Small RNAs
Low-molecular-weight RNAs were separated from total cellular RNAs extracted using Direct-zol RNA MiniPrep (Zymo Research) as previously described77. Libraries were prepared with TruSeq Small RNA Library Preparation Kit (Illumina) and sequenced using NextSeq (Illumina).
Small RNA reads from three replicates were trimmed for adapters and quality using cutadapt78 and merged into a single non-redundant small RNA library. Small RNA reads of 21-, 22- and 24-nucleotide length were then mapped to the A. hypogaea cv. Tifrunner genome using Bowtie2. For each read, all alignments where reported using the -a option in Bowtie279. Only perfectly matched reads were kept for further analysis. Unique small RNA reads were defined as reads that perfectly aligned to a single location in the reference genome.
Diverse genotypes for analysis of genome structures
A diverse panel of more than 200 tetraploid genotypes that represent the wild A. monticola, all six botanical varieties of the cultivated A. hypogaea, modern varieties and induced allotetraploid hybrids of peanut’s ancestral species A. duranensis (V14167) and A. ipaensis (K30076) were sequenced using Illumina short (100–250 bp) paired-end sequencing (dataset 2 in ref. 25).
Investigating genetic exchange between subgenomes
Genetic exchange between the subgenomes was inferred by different methods: observing mapping densities of Illumina whole-genome sequences onto the combined sequenced diploid ancestral species genomes6; and by analysis of SNPs that consistently differentiate representatives of A and B genome diploid species.
Mapping densities onto the combined diploid ancestral species genomes
The methodology that uses mapping densities takes advantage of the diploid genome sequences being very similar to the corresponding subgenomes of A. hypogaea6. After quality filtering, sequences were assigned to A or B genomes by mapping to the combined chromosomal pseudomolecule sequences of A. duranensis V14167 and A. ipaensis K30076 (ref. 6) using Bowtie2 version 2.2.9 with the -sensitive -local option. Mpileup files were generated using SamTools version 1.3, which were parsed to create average mapping densities for defined windows dividing the chromosomes. Three types of windows were used. First were windows defined by the annotated start and stop positions of 17,373 high-confidence orthologous gene pairs from the two diploids (dataset 4a in ref. 25). Second were pairs of windows defined using syntenous regions of the diploid genomes identified using DAGchainer80; taking into account the relative orientation of the blocks and any difference in size, syntenous blocks were divided into corresponding A–B windows of approximately 10 kb (dataset 4b in ref. 25). Third were fixed-size 10-kb windows (dataset 4c in ref. 25). Normalized mapping densities (and for the first and second windows, log2-normalized values of ratios of mapping densities) when displayed graphically across chromosomal sequences allow the genome compositions of tetraploid genotypes to be visualized.
SNPs that consistently differentiate A and B genomes
We chose Arachis species and accessions within the A and B genome groups with well-defined relationships: A. ipaensis K30076, A. magna K30097 Krapov., W.C. Gregory & C.E. Simpson, A. valida V9157 Krapov. & W.C. Gregory, A. duranensis V14167, A. duranensis K36003, A. duranensis K30077 and A. cardenasii GKP10017 Krapov. & W.C. Gregory. For an overview of their relationships, see Supplementary Fig. 24.
Whole-genome Illumina sequences from these diploid species/accessions were mapped separately onto the Tifrunner A subgenome chromosomes and the B subgenome chromosomes. Variants were called using SamTools mpileup. SNPs that are diagnostic of the A and B genomes were identified as sites that differentiated all detected A species from the B species. For analysis of the diverse tetraploid genotypes, Illumina whole-genome sequences from each tetraploid accession were mapped to the A subgenome chromosomes and the B subgenome chromosomes separately. The alignment files were used to count the alleles at each diagnostic site using the pysam module in biopython.
Homeologous exchange in polyploid hybrids made from peanut’s ancestral species
We investigated homeologous exchange in induced allotetraploid A. duranensis × A. ipanesis (2n = 4× = 40) individuals by genotyping using the Affymetrix Axiom_Arachis Array16. By reference to the diploid parental controls, alleles could be assigned to the A and B genomes, enabling identification of genotype calls that represented AABB, AAAA and BBBB genome structures (dataset 6 in ref. 25).
Circular DNA and active transposable elements
For descriptions of the methods used for identifying active transposable elements, see Supplementary Note 2.
Identification of the A subgenome ancestor
For this analysis, we used representatives from every known major population of A. duranensis. To compile these, we drew on knowledge of the distributions and characteristics of populations that has been built up during botanical expeditions and research over more than 50 years (much of this documented in a previous study by Krapovickas and Gregory5). Representatives of most populations were available from the USDA National Genetic Resources Program (USDA Germplasm Resources Information Network; https://www.ars-grin.gov), these were supplemented with DNA samples from accessions held at the Instituto de Botánica del Nordeste, Embrapa Recursos Genéticos e Biotecnologia and the International Crops Research Institute for the Semi-Arid Tropics. In total, DNA from 55 accessions, plus some control species were used (Supplementary Table 11).
An exome-capture bait set (SeqCap EZ Developer; Nimblegen/Roche) was designed for 30,460 genes, including untranslated regions, annotated in the A. ipaensis genome assembly6 and 6,993 A. hypogaea SNPs, the majority of which had been identified previously by genotyping by sequencing. The bait set represents a capture region of about 50.14 Mb in diploid peanuts.
Barcode-indexed sequencing libraries were generated from genomic DNA samples sheared on an E220 Focused Ultrasonicator (Covaris), For each sample, 1 μg of sheared DNA was converted to sequencing libraries using a Kapa Hyper Library Preparation Kit (Kapa Biosystems/Roche). The exome-capture analysis was carried out with SeqCap EZ capture reagents according to the recommendations of the manufacturer (Nimblegen/Roche). Subsequently, 18 libraries were pooled before exome capture and sequenced on one Illumina HiSeq 4000 (Illumina) lane with paired-end 150-bp reads.
Sequencing data were mapped to the Tifrunner A subgenome chromosomes using BWA MEM with default parameters. Variants were called using SamTools mpileup and bcftools call (bcftools call -vc). Only variants that were called ‘homozygous alternative’ were considered. To normalize for missing data among lines across observed variant sites, similarity to the Tifrunner A subgenome was calculated as observed sites at which there is read coverage and a genotype score as homozygous reference divided by the total number of observed sites with read coverage. This strategy controls for differences in covered sites among the accessions.
Statistical analysis
For a description of the statistical analyses, see Supplementary Note 3.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The datasets generated during and/or analyzed for this study are available in the public repository of the National Center for Biotechnology Information (NCBI; https://www.ncbi.nlm.nih.gov) and/or the open access internet sites PeanutBase (https://peanutbase.org/). Genome assemblies and annotations, identified transposable elements, transcript assemblies, base methylation states and map data are available at Peanutbase (https://peanutbase.org/peanut_genome). A. hypogaea cv. Tifrunner sequence reads are archived in the NCBI under BioProject accession number PRJNA419393, the genome assembly has GenBank accession numbers CM009801–CM009820. Small RNA sequences are deposited in the NCBI Sequence Read Archive (SRA) under accession numbers SAMN06658954, SAMN06658955 and SAMN06658956. Datasets 1–9, as cited in manuscript, are deposited at https://doi.org/10.25739/hb5x-wx74, Cyverse (http://datacommons.cyverse.org/browse/iplant/home/shared/commons_repo/curated/Bertioli_Arachis_genome_supplement_TVDM_Mar2019) and PeanutBase (https://peanutbase.org/data/public/Arachis_hypogaea/Tifrunner.esm.TVDM/). Whole-genome sequencing data of diverse accessions are deposited in the NCBI under BioProject accession numbers PRJNA525866, PRJNA511155 and PRJNA490832.
References
Hilu, K. W. Polyploidy and the evolution of domesticated plants. Am. J. Bot. 80, 1494–1499 (1993).
Salman-Minkov, A., Sabath, N. & Mayrose, I. Whole-genome duplication as a key factor in crop domestication. Nat. Plants 2, 16115 (2016).
Husted, L. Cytological studies of the peanut Arachis. II. Chromosome number, morphology and behavior, and their application to the problem of the origin of the cultivated forms. Cytologia 7, 396–423 (1936).
Fernández, A. & Krapovickas, A. Cromosomas y evolución en Arachis (Leguminosae). Bonplandia 8, 187–220 (1994).
Krapovickas, A. & Gregory, W. C. Taxonomy of the genus Arachis (Leguminosae). Bonplandia 16, 1–205 (2007).
Bertioli, D. J. et al. The genome sequences of Arachis duranensis and Arachis ipaensis, the diploid ancestors of cultivated peanut. Nat. Genet. 48, 438–446 (2016).
Kochert, G. et al. RFLP and cytogenetic evidence on the origin and evolution of allotetraploid domesticated peanut, Arachis hypogaea (Leguminosae). Am. J. Bot. 83, 1282–1291 (1996).
Seijo, G. et al. Genomic relationships between the cultivated peanut (Arachis hypogaea, Leguminosae) and its close relatives revealed by double GISH. Am. J. Bot. 94, 1963–1971 (2007).
Moretzsohn, M. C. et al. A study of the relationships of cultivated peanut (Arachis hypogaea) and its most closely related wild species using intron sequences and microsatellite markers. Ann. Bot. 111, 113–126 (2013).
Bertioli, D. J. et al. An overview of peanut and its wild relatives. Plant Genet. Resour. 9, 134–149 (2011).
Krapovickas, A., Vanni, R. O., Pietrarelli, J. R., Williams, D. E. & Simpson, C. E. Las Razas de Maní de Bolivia. Bonplandia 18, 95–189 (2009).
Ramos, M. L. et al. Chromosomal and phylogenetic context for conglutin genes in Arachis based on genomic sequence. Mol. Genet. Genomics 275, 578–592 (2006).
Bertioli, D. J. et al. The repetitive component of the A genome of peanut (Arachis hypogaea) and its role in remodelling intergenic sequence space since its evolutionary divergence from the B genome. Ann. Bot 112, 545–559 (2013).
Nielen, S. et al. Matita, a new retroelement from peanut: characterization and evolutionary context in the light of the Arachis A–B genome divergence. Mol. Genet. Genomics 287, 21–38 (2012).
Leal-Bertioli, S. et al. Tetrasomic recombination is surprisingly frequent in allotetraploid Arachis. Genetics 199, 1093–1105 (2015).
Clevenger, J. et al. Genome-wide SNP genotyping resolves signatures of selection and tetrasomic recombination in peanut. Mol. Plant 10, 309–322 (2017).
Nguepjop, J. R. et al. Evidence of genomic exchanges between homeologous chromosomes in a cross of peanut with newly synthetized allotetraploid hybrids. Front. Plant Sci. 7, 1635 (2016).
Leal-Bertioli, S. C. M. et al. Segmental allopolyploidy in action: increasing diversity through polyploid hybridization and homoeologous recombination. Am. J. Bot. 105, 1053–1066 (2018).
Eid, J. et al. Real-time DNA sequencing from single polymerase molecules. Science 323, 133–138 (2008).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95 (2017).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Holbrook, C. C. & Culbreath, A. K. Registration of ‘Tifrunner’ peanut. J. Plant Regist. 1, 124 (2007).
Samoluk, S. S., Chalup, L., Robledo, G. & Seijo, J. G. Genome sizes in diploid and allopolyploid Arachis L. species (section Arachis). Genet. Resour. Crop Evol. 62, 747–763 (2015).
Dhillon, S. S., Rake, A. V. & Miksche, J. P. Reassociation kinetics and cytophotometric characterization of peanut (Arachis hypogaea L.) DNA. Plant Physiol. 65, 1121–1127 (1980).
Bertioli, D. Supplementary material for "The genome sequence of segmental allotetraploid peanut Arachis hypogaea". CyVerse Data Commons https://doi.org/10.25739/hb5x-wx74 (2019).
Chalhoub, B. et al. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 345, 950–953 (2014).
Zhang, T. et al. Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat. Biotechnol. 33, 531–537 (2015).
Gaeta, R. T. & Pires, C. J. Homoeologous recombination in allopolyploids: the polyploid ratchet. New Phytol. 186, 18–28 (2010).
Hurgobin, B. et al. Homoeologous exchange is a major cause of gene presence/absence variation in the amphidiploid Brassica napus. Plant Biotechnol. J. 16, 1265–1274 (2018).
Robledo, G., Lavia, G. I. & Seijo, G. Species relations among wild Arachis species with the A genome as revealed by FISH mapping of rDNA loci and heterochromatin detection. Theor. Appl. Genet. 118, 1295–1307 (2009).
Robledo, G. & Seijo, G. Species relationships among the wild B genome of Arachis species (section Arachis) based on FISH mapping of rDNA loci and heterochromatin detection: a new proposal for genome arrangement. Theor. Appl. Genet. 121, 1033–1046 (2010).
Moretzsohn, M. C. et al. A linkage map for the B-genome of Arachis (Fabaceae) and its synteny to the A-genome. BMC Plant Biol. 9, 40 (2009).
Levinson, G. & Gutman, G. A. Slipped-strand mispairing: a major mechanism for DNA sequence evolution. Mol. Biol. Evol. 4, 203–221 (1987).
Lanciano, S. et al. Sequencing the extrachromosomal circular mobilome reveals retrotransposon activity in plants. PLoS Genet. 13, e1006630 (2017).
Shirasawa, K. et al. Characterization of active miniature inverted-repeat transposable elements in the peanut genome. Theor. Appl. Genet. 124, 1429–1438 (2012).
Nagy, E. D. et al. A high-density genetic map of Arachis duranensis, a diploid ancestor of cultivated peanut. BMC Genomics 13, 469 (2012).
Ren, L., Huang, W., Cannon, E. K. S., Bertioli, D. J. & Cannon, S. B. A mechanism for genome size reduction following genomic rearrangements. Front. Genet. 19, 454 (2018).
Fávero, A. P., Simpson, C. E., Valls, F. M. J. & Velo, N. A. Study of evolution of cultivated peanut through crossability studies among Arachis ipaensis, A. duranensis and A. hypogaea. Crop Sci. 46, 1546–1552 (2006).
Grabiele, M., Chalup, L., Robledo, G. & Seijo, G. Genetic and geographic origin of domesticated peanut as evidenced by 5S rDNA and chloroplast DNA sequences. Plant Syst. Evol. 298, 1151–1165 (2012).
Chen, X. et al. Draft genome of the peanut A-genome progenitor (Arachis duranensis) provides insights into geocarpy, oil biosynthesis, and allergens. Proc. Natl Acad. Sci. USA 113, 6785–6790 (2016).
Blanc, G. & Wolfe, K. H. Widespread paleopolyploidy in model plant species inferred from age distributions of duplicate genes. Plant Cell 16, 1667–1678 (2004).
Soltis, D. E., Soltis, P. S. & Tate, J. A. Advances in the study of polyploidy since plant speciation. New Phytol. 161, 173–191 (2004).
Wendel, J. F. The wondrous cycles of polyploidy in plants. Am. J. Bot. 102, 1753–1756 (2015).
Leal-Bertioli, S. C. et al. The effect of tetraploidization of wild Arachis on leaf morphology and other drought-related traits. Environ. and Exp. Bot 84, 17–24 (2012).
Leal‐Bertioli, S. C. et al. Phenotypic effects of allotetraploidization of wild Arachis and their implications for peanut domestication. Am. J. Bot. 104, 379–388 (2017).
Stebbins, G. L. Types of polyploids: their classification and significance. Adv. Genet. 1, 403–429 (1947).
Xiao, C. L. et al. MECAT: fast mapping, error correction, and de novo assembly for single-molecule sequencing reads. Nat. Methods 14, 1072–1074 (2017).
Chin, C. S. et al. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat. Methods 10, 563–569 (2013).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98 (2016).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3, 99–101 (2016).
Dudchenko, O. et al. The Juicebox Assembly Tools module facilitates de novo assembly of mammalian genomes with chromosome-length scaffolds for under $1000. Preprint at bioRxiv https://doi.org/10.1101/254797 (2018).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Krumsiek, J., Arnold, R. & Rattei, T. Gepard: a rapid and sensitive tool for creating dotplots on genome scale. Bioinformatics 23, 1026–1028 (2007).
Agarwal, G. et al. High-density genetic map using whole-genome resequencing for fine mapping and candidate gene discovery for disease resistance in peanut. Plant Biotechnol. J. 16, 1954–1967 (2018).
Pandey, M. K. et al. Development and evaluation of a high density genotyping ‘Axiom_Arachis’ array with 58 K SNPs for accelerating genetics and breeding in groundnut. Sci. Rep. 7, 40577 (2017).
Lorieux, M. MapDisto: fast and efficient computation of genetic linkage maps. Mol. Breed. 30, 1231–1235 (2012).
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004).
Wu, T. D. & Nacu, S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26, 873–881 (2010).
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512 (2013).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Campbell, M. S. et al. MAKER-P: a tool kit for the rapid creation, management, and quality control of plant genome annotations. Plant Physiol. 164, 513–524 (2014).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59 (2004).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, W435–W439 (2006).
Clevenger, J., Chu, Y., Scheffler, B. & Ozias-Akins, P. A developmental transcriptome map for allotetraploid Arachis hypogaea. Front. Plant Sci. 7, 1446 (2016).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421–430 (2009).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Alexa, A., Rahnenführer, J. & Lengauer, T. Improved scoring of functional groups from gene expression data by decorrelating GO graph structure. Bioinformatics 22, 1600–1607 (2006).
Alexa, A. & Rahnenfuhrer, J. topGO: enrichment analysis for gene ontology. R version 2 2240 (2016); https://www.r-project.org
Urich, M. A., Nery, J. R., Lister, R., Schmitz, R. J. & Ecker, J. R. MethylC-seq library preparation for base-resolution whole-genome bisulfite sequencing. Nat. Protoc. 10, 475–483 (2015).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Lu, C., Meyers, B. C. & Green, P. J. Construction of small RNA cDNA libraries for deep sequencing. Methods 43, 110–117 (2007).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Haas, B. J., Delcher, A. L., Wortman, J. R. & Salzberg, S. L. DAGchainer: a tool for mining segmental genome duplications and synteny. Bioinformatics 20, 3643–3646 (2004).
Acknowledgements
We thank G. Birdsong, V. Nwosu, J. Elder, D. Smyth, H. Valentine, F. Luo, D. Hoisington, H. Shapiro, D. Ward, S. Knapp, R. Wilson and S. Brown for their support of, and work for, the Peanut Genome Initiative. Major financial contributors for this work were from Mars-Wrigley Confectionary, US peanut sheller associations, the National Peanut Board and other industry groups. A full list can be downloaded at https://peanutbase.org/IPGI. D.J.B. thanks the Georgia Peanut Commission and the Georgia Research Alliance for support. The genome sequencing was funded from grant 04-852-14 from The Peanut Foundation to J.S. and B.E.S., characterization of diverse genotypes was funded from grant 04-805-17 to D.J.B. The work conducted by the US Department of Energy Joint Genome Institute is supported by the Office of Science of the US Department of Energy under contract number DE-AC02-05CH11231. The work done at the DNA Technologies and Expression Analysis Cores at the UC Davis Genome Center was supported by NIH Shared Instrumentation Grant 1S10OD010786-01. We thank the US National Science Foundation for support from grant number 1339194 to S.A.J. This research was funded in part by the US Department of Agriculture Agricultural Research Service, projects 5030-21000-069-00-D, 6048-21000-028-00-D, 6048-21000-029-00-D, 6066-21310-005-00-D and NIFA Award no. 2018-67013-28139. We also grateful for funding granted to X.Z. and Z.Z. from the Henan Province Open Cooperation Project of Science and Technology (172106000007), the Henan Science and Technology Major Project of the Ministry of Science and Technology of China (161100111000), the China Agriculture Research System (CARS-13), and Henan Agriculture Research System (S2012-5). We thank the Indian Council of Agricultural Research, National Agricultural Science Funds, Government of India and the CGIAR Research Program on Grain Legumes and Dryland Cereals for grants to R.K.V. and M.K.P. ICRISAT is a member of the CGIAR. S.S. was supported by the Ramalingwaswami Re-entry Grant (BT/RLF/Re-entry/41/2013) from the Ministry of Science and Technology, India. We thank D. Kudrna at the University of Arizona for high-molecular-weight DNA extractions, S. Simpson of USDA ARS GBRU for valuable support with PacBio sequencing and the USDA National Plant Germplasm System for Arachis seeds.
Author information
Authors and Affiliations
Contributions
Project planning and coordination: J.S., S.A.J., B.E.S., S.B.C., P.O.-A., D.J.B., C.C.H., J.G., E.K.S.C., R.K.V., R.M., S.C.M.L.-B., L.F. and A.D.F. Production of base genetic material (including mapping populations and allotetraploid hybrids): C.C.H., Y.C., P.O.-A., S.C.M.L.-B., D.J.B. and M.C.M. Tifrunner genome sequencing (including BACs and quality control): J.J., A.S., J.S., J.G. and B.E.S. Hi-C libraries and sequencing: O.D., C.G.L., M.K.P. and E.L.A. Genome assembly: J.J., J.S., O.D. and D.J.B. Transcriptome assembly, gene, mobile element and repeat annotation: D.G., J. Campbell, C. Cameron, S.D., A.D.F., N.T.W., P.O.-A., D.J.B. and S.A.J. Comparison of gene expression in subgenomes: A.S. and J.S. Extra chromosomal circular DNAs: M.M. and S.L. Small RNAs and methylation: K.D.K., J.H.S., M.E.B., S.A.J., S.C.M.L.-B. and D.J.B. Structural analysis of Tifrunner genome: L.R., S.B.C., E.K.S.C., D.G., D.J.B., J. Clevenger and B.A. Data preparation and data basing, visualizations and expression analysis: S.B.C., E.K.S.C. and W.H. Genotyping and linkage mapping: C.B.-T., C. Chavarro, Y.C., J. Clevenger, S.C.M.L.-B., G.A., B.G., P.O.-A., S.A.J. and D.J.B. Analysis of homeologous recombination: B.A., J. Clevenger, S.C.M.L.-B. and D.J.B. Diverse tetraploid samples and data: S.C.M.L.-B., X.Z., Z.Z., Z.S., A.C., M.K.P., R.K.V., K.S., P.O.-A., S.A.J. D.J.B. and C. Chavarro. Analysis of variations in tetraploid genome structure: B.A., J. Clevenger, W.K. and D.J.B. Curation and sequencing of A. duranensis accessions: S.S.S., G.S., S.C.M.L.-B. and D.J.B. Exome capture and analysis: L.F., R.M., S.S., S.S.S., G.S., J. Clevenger, S.C.M.L.-B. and D.J.B. Biogeography: G.S., S.S.S. and D.J.B. Manuscript: D.J.B., G.S., S.B.C, M.M., J.J., P.O.-A., J. Campbell, J. Clevenger, S.C.M.L.-B, R.M., D.G., M.E., S.S.S., L.R. and S.A.J.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Notes 1–3, Supplementary Tables 1–16 and Supplementary Figs. 1–35
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bertioli, D.J., Jenkins, J., Clevenger, J. et al. The genome sequence of segmental allotetraploid peanut Arachis hypogaea. Nat Genet 51, 877–884 (2019). https://doi.org/10.1038/s41588-019-0405-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0405-z
This article is cited by
-
Homologous mapping yielded a comprehensive predicted protein–protein interaction network for peanut (Arachis hypogaea L.)
BMC Plant Biology (2024)
-
Genome-wide association study and development of molecular markers for yield and quality traits in peanut (Arachis hypogaea L.)
BMC Plant Biology (2024)
-
High-throughput diagnostic markers for foliar fungal disease resistance and high oleic acid content in groundnut
BMC Plant Biology (2024)
-
Genome wide analysis of carotenoid cleavage oxygenases (CCO) gene family in Arachis hypogaea (peanut) under biotic stress
BMC Genomics (2024)
-
Contribution of homoeologous exchange to domestication of polyploid Brassica
Genome Biology (2024)