Abstract
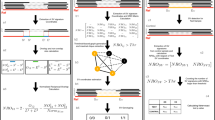
Whole-genome sequencing of DNA from single cells has the potential to reshape our understanding of mutational heterogeneity in normal and diseased tissues. However, a major difficulty is distinguishing amplification artifacts from biologically derived somatic mutations. Here, we describe linked-read analysis (LiRA), a method that accurately identifies somatic single-nucleotide variants (sSNVs) by using read-level phasing with nearby germline heterozygous polymorphisms, thereby enabling the characterization of mutational signatures and estimation of somatic mutation rates in single cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Code availability
LiRA is available at https://github.com/parklab/LiRA.
Data availability
LiRA was applied to single-neuron and bulk sequencing data collected from the postmortem brain, heart (UMB1465 and UMB4638), and liver (UMB4643) tissue of three individuals. These data were acquired as part of a previous study5 and are available in the NCBI SRA under accession nos. SRP041470 (UMB1465) and SRP061939 (UMB4638 and UMB4643). The neuron counts by individual were: UMB1465 (16); UMB4638 (10); and UMB4643 (10).
References
Leung, M. L., Wang, Y., Waters, J. & Navin, N. E. SNES: single nucleus exome sequencing. Genome Biol. 16, 55 (2015).
Xu, X. et al. Single-cell exome sequencing reveals single-nucleotide mutation characteristics of a kidney tumor. Cell 148, 886–895 (2012).
Hou, Y. et al. Single-cell exome sequencing and monoclonal evolution of a JAK2-negative myeloproliferative neoplasm. Cell 148, 873–885 (2012).
Baslan, T. et al. Genome-wide copy number analysis of single cells. Nat. Protoc. 7, 1024–1041 (2012).
Lodato, M. A. et al. Somatic mutation in single human neurons tracks developmental and transcriptional history. Science 350, 94–98 (2015).
Wang, Y. et al. Clonal evolution in breast cancer revealed by single nucleus genome sequencing. Nature 512, 155–160 (2014).
Zong, C., Lu, S., Chapman, A. R. & Xie, X. S. Genome-wide detection of single-nucleotide and copy-number variations of a single human cell. Science 338, 1622–1626 (2012).
Dean, F. B. et al. Comprehensive human genome amplification using multiple displacement amplification. Proc. Natl Acad. Sci. USA 99, 5261–5266 (2002).
Chen, C. et al. Single-cell whole-genome analyses by Linear Amplification via Transposon Insertion (LIANTI). Science 356, 189–194 (2017).
Huang, L., Ma, F., Chapman, A., Lu, S. & Xie, X. S. Single-cell whole-genome amplification and sequencing: methodology and applications. Annu. Rev. Genomics Hum. Genet. 16, 79–102 (2015).
Gawad, C., Koh, W. & Quake, S. R. Single-cell genome sequencing: current state of the science. Nat. Rev. Genet. 17, 175–188 (2016).
de Bourcy, C. F. A. et al. A quantitative comparison of single-cell whole genome amplification methods. PLoS ONE 9, e105585 (2014).
Esteban, J. A., Salas, M. & Blanco, L. Fidelity of phi 29 DNA polymerase: comparison between protein-primed initiation and DNA polymerization. J. Biol. Chem. 268, 2719–2726 (1993).
Fryxell, K. J. & Zuckerkandl, E. Cytosine deamination plays a primary role in the evolution of mammalian isochores. Mol. Biol. Evol. 17, 1371–1383 (2000).
Lindahl, T. & Nyberg, B. Heat-induced deamination of cytosine residues in deoxyribonucleic acid. Biochemistry 13, 3405–3410 (1974).
Frederico, L. A., Kunkel, T. A. & Shaw, B. R. A sensitive genetic assay for the detection of cytosine deamination: determination of rate constants and the activation energy. Biochemistry 29, 2532–2537 (1990).
Usuyama, N. et al. HapMuC: somatic mutation calling using heterozygous germ line variants near candidate mutations. Bioinformatics 30, 3302–3309 (2014).
Freed, D. & Pevsner, J. The contribution of mosaic variants to autism spectrum disorder. PLoS Genet. 12, e1006245 (2016).
Ju, Y. S. et al. Somatic mutations reveal asymmetric cellular dynamics in the early human embryo. Nature 543, 714–718 (2017).
Ramu, A. et al. DeNovoGear: de novo indel and point mutation discovery and phasing. Nat. Methods 10, 985–987 (2013).
Dong, X. et al. Accurate identification of single-nucleotide variants in whole-genome-amplified single cells. Nat. Methods 14, 491–493 (2017).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Hoang, M. L. et al. Genome-wide quantification of rare somatic mutations in normal human tissues using massively parallel sequencing. Proc. Natl Acad. Sci. USA 113, 9846–9851 (2016).
Zafar, H., Wang, Y., Nakhleh, L., Navin, N. & Chen, K. Monovar: single-nucleotide variant detection in single cells. Nat. Methods 13, 505–507 (2016).
Koboldt, D. C. et al. VarScan: variant detection in massively parallel sequencing of individual and pooled samples. Bioinformatics 25, 2283–2285 (2009).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Chen, L., Liu, P., Evans, T. C. Jr. & Ettwiller, L. M. DNA damage is a pervasive cause of sequencing errors, directly confounding variant identification. Science 355, 752–756 (2017).
Lodato, M. A. et al. Aging and neurodegeneration are associated with increased mutations in single human neurons. Science 359, 555–559 (2018).
Li, Y. et al. Single-cell sequencing analysis characterizes common and cell-lineage-specific mutations in a muscle-invasive bladder cancer. Gigascience 1, 12 (2012).
Ho, J. R. et al. Deregulation of Rab and Rab effector genes in bladder cancer. PLoS ONE 7, e39469 (2012).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Delaneau, O. et al. Integrating sequence and array data to create an improved 1000 Genomes Project haplotype reference panel. Nat. Commun. 5, 3934 (2014).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Acknowledgements
This work was mainly supported by the training grant in Bioinformatics and Integrative Genomics from the National Human Genome Research Institute (grant no. T32HG002295 to C.L.B., A.R.B., L.J.L., and V.V.), a Brain Somatic Mosaicism Network grant from the National Institute of Mental Health (grant no. U01MH106883 to P.J.P., C.A.W.), and Ludwig Center at Harvard Medical School (P.J.P.). I.C.-C. received funding from the European Union (Marie Curie Skłodowska-Curie grant agreement no. 703543).
Author information
Authors and Affiliations
Contributions
C.L.B. and M.A.L. conceived the project and P.J.P. supervised it. C.L.B. developed the algorithm. C.L.B., A.R.B., M.K., and A.G. generated the alignment and performed the variant calling. A.R.B., M.A.L., R.E.R., L.J.L., V.V., D.C.G., I.C.-C., M.A.S., M.K., M.E.C., and C.A.W. suggested impactful improvements to LiRA and aided in evaluating its performance. C.L.B. wrote the manuscript supervised by P.J.P., with input from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Bohrson, C.L., Barton, A.R., Lodato, M.A. et al. Linked-read analysis identifies mutations in single-cell DNA-sequencing data. Nat Genet 51, 749–754 (2019). https://doi.org/10.1038/s41588-019-0366-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0366-2
This article is cited by
-
Analyzing somatic mutations by single-cell whole-genome sequencing
Nature Protocols (2024)
-
Computational immunogenomic approaches to predict response to cancer immunotherapies
Nature Reviews Clinical Oncology (2024)
-
Somatic mutations in aging and disease
GeroScience (2024)
-
Long-read whole-genome analysis of human single cells
Nature Communications (2023)
-
Comprehensive benchmarking and guidelines of mosaic variant calling strategies
Nature Methods (2023)