Abstract
Aspergillus section Nigri comprises filamentous fungi relevant to biomedicine, bioenergy, health, and biotechnology. To learn more about what genetically sets these species apart, as well as about potential applications in biotechnology and biomedicine, we sequenced 23 genomes de novo, forming a full genome compendium for the section (26 species), as well as 6 Aspergillus niger isolates. This allowed us to quantify both inter- and intraspecies genomic variation. We further predicted 17,903 carbohydrate-active enzymes and 2,717 secondary metabolite gene clusters, which we condensed into 455 distinct families corresponding to compound classes, 49% of which are only found in single species. We performed metabolomics and genetic engineering to correlate genotypes to phenotypes, as demonstrated for the metabolite aurasperone, and by heterologous transfer of citrate production to Aspergillus nidulans. Experimental and computational analyses showed that both secondary metabolism and regulation are key factors that are significant in the delineation of Aspergillus species.
Similar content being viewed by others
Main
Species in the genus Aspergillus are of broad interest to medical1, applied2,3, and basic research4. Members of Aspergillus section Nigri (‘black aspergilli’) are prolific producers of native and heterologous proteins5,6, organic acids (in particular citric acid2,7,8), and secondary metabolites (including biopharmaceuticals and mycotoxins like ochratoxin A). Furthermore, the section members are generally very efficient producers of extracellular enzymes9,10; they are the production organisms for 49 out of 260 industrial enzymes11,12. Among the most important of these, in addition to A. niger, are A. tubingensis, A. aculeatus, and A. luchuensis (previously A. acidus, A. kawachii, and A. awamori13,14,15, respectively).
Members of Aspergillus section Nigri are also known as destructive degraders of foods and feeds, and some isolates produce the potent mycotoxins ochratoxin A16 and fumonisins17,18,19. In addition, some species in this section have been proposed to be pathogenic to humans and other animals20. It is thus of interest to further examine section Nigri for industrial exploitation, as well as prevention of food spoilage, toxin production, and pathogenicity caused by these fungi.
A combined phylogenetic and phenotypic approach has shown that section Nigri contains at least 27 species21,22,23,24,25. Recent results have shown that the section contains species with high diversity and may consist of two separate clades: the biseriate species and the uniseriate species26, which show differences in sexual states27, sclerotium formation28, and secondary metabolite production29. In the section, only six species have had their genome sequenced: A. niger2,8, A. luchuensis15,30, A. carbonarius31, A. aculeatus31, A. tubingensis31, and A. brasiliensis31.
This section, with its combination of species richness and fungal species with a diverse impact on humanity, is thus particularly interesting for studying the diversification of fungi into species. In this study, we have de novo-sequenced the genomes of 20 species of section Nigri, thus completing a genome compendium of 26 described species in the section. Further, we have genome-sequenced three additional A. niger isolates (including two previously described as species A. lacticoffeatus10 and A. phoenicis32), which in combination with the other analyses allows for inter- and intraspecies comparison of 32 isolates. The development of algorithms for comparative genomics, combined with experimental analysis of the species, allows us to track genetic diversity across genomes, from the protein level, over the evolution of biosynthetic gene clusters, to the groups of genes that define clades or individual species. The high resolution in genome sequences allows us to characterize both species diversification and variation within species.
Results
New genomes show high genetic diversity of section Nigri
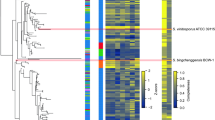
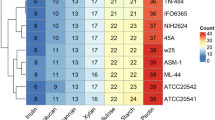
We present 23 whole-genome draft sequences: 20 genomes of section Nigri species previously unsequenced and 3 additional A. niger genomes for assessment of intraspecies diversity. All genomes were sequenced, assembled, and annotated using the Joint Genome Institute (JGI) fungal genome pipeline33,34 (Supplementary Table 1; genomes were sequenced by either Illumina or Pacific Biosciences sequencing). Figure 1 shows a phylogenetic tree as well as gene richness, number of scaffolds, and functional annotation (InterPro35,36). The tree supports previous proposals10,32 that A. lacticoffeatus and A. phoenicis are synonyms of A. niger.
Additional information is available in Supplementary Table 1. a, Phylogenetic tree created using RAxML49, MAFFT50, and gBlocks51, based on 2,022 conserved genes. Plate growth pictures are presented for each newly sequenced species. b, Colors indicate whether the organism is from this or another sequencing project. c, Five bubble plots of descriptive numbers for each genome. The bubble sizes have been scaled to the categories and are not comparable across categories.
In comparing key statistics of the genomes, we found that some traits are quite similar and others surprisingly variable. Many of the investigated species have around the average number of genes (11,900), but there is considerable variation from the smallest number of predicted genes (10,066) to the largest (13,687). The smallest number of predicted genes in section Nigri is found in A. saccharolyticus, which supports the previous observation37,38 that this species is quite atypical in section Nigri.
We further evaluated the annotation of the 23 genome sequences we generated. The percentage of complete genes (including a start and stop codon) is in the range of 94–98%, and 67% of the proteins could be assigned one or more InterPro domains. The number of scaffolds (average 166) varies from 47 in A. piperis to 518 in A. ellipticus. On average, 70% of the proteins had sequence homologs in Swiss-Prot (91% of proteins have homologs within section Nigri; see next section). This means that even though six members of section Nigri have already been sequenced, ~30% of the predicted gene models in each of the new genomes are not found to encode proteins with homologs in Swiss-Prot.
The pan- and core-genome shows genome flexibility
Given the genetic diversity in section Nigri, we were interested in examining the extent of genome diversification. For this analysis, we focused on three conceptual groups of genes:
-
(1)
The pan-genome: all genes present in one or more species.
-
(2)
The core-genome: genes present in all included species, including paralogs. This set is expected to encode cellular functions needed for all species.
-
(3)
Species-unique genes: genes found in only one species in our analysis, with or without paralogs. Included in these, we would expect to find genes involved in environmental adaptation. This group can also include annotation errors.
We first identified orthologs and paralogs with a BLASTp-based pipeline using reciprocal hits according to cut-offs specifically selected here for the Aspergillus genus (Methods). Groups of homologous proteins are referred to as families. Figure 2a–c shows the overall genetic diversity between 38 fungal strains (32 species) from closely related genera (Fig. 2a), within the Aspergillus genus (36 of the 38 strains; Fig. 2b), and from section Nigri (32 of the 38 strains; Fig. 2c).
a–c, Histogram representation of the total number of genes and families distributed over the pan-genome, core-genome, and unique genome for the 38 fungal genomes of this study (a), the 36 Aspergillus genus genomes (b), and the 32 genomes in section Nigri (c). d, Dendrogram of the phylogenetic relation between the 32 species in section Nigri. The black nodes represent homolog families found only in the species branching from the nodes. The white boxes represent the genes unique to the specific species. e, Stacked histograms of the gene count, the core-genome genes, and unique genes for each species; values shown are total numbers of genes. Dark colors, InterPro annotation; light colors, no annotation.
The Aspergillus genus pan-genome comprises 433,116 genes across the 36 Aspergillus genomes, and from this, 62,996 gene families were constructed. Of those families, 6% are found in all genomes (3,769 core families), while 9% are genes without orthologs in the other genomes (40,424 unique genes; 39,929 unique families) (Fig. 2b). We also found evidence of gene loss, duplication, and potential gene transfers between species of this section, as 23% of the pan-gene families are not present in groups of species fitting the phylogenetic tree (Supplementary Table 2). This is consistent with previous work reporting extensive horizontal gene transfer in Aspergillus39.
We further performed an analysis defining the number of core-gene families in section Nigri and in all sub-clades thereof (Fig. 2d). The core-genome of section Nigri is 32% larger than that of the genus (4,983 families relative to 3,769; Fig. 2b,c). Conversely, 9% are unique to a specific species (32,378 unique genes in 32,036 families; Fig. 2c). The fraction of genes unique to a species is similar within the section and across the genus, meaning that adding a new section Nigri genome adds as many new genes as adding a more distantly related Aspergillus (within the analyzed group of species). This is rather interesting and shows a generally high genetic diversity of genus Aspergillus. However, such a tendency could be also the result of overpredicting genes, considering the low rate of InterPro annotation in the unique genes (Fig. 2e).
The section Nigri core genome contains carbohydrate-active enzymes and secondary metabolism gene clusters
To associate biological functions to the pan-, core-, and unique genomes, and genes exclusive to only members of the black aspergilli, we employed the InterPro database36. An examination of the core-genome of 38 fungal genomes (Fig. 2a and Supplementary Fig. 1) revealed that only 4.5% of the genes lack InterPro domains (Supplementary Table 3a), indicating—as would be expected—that the core-genes across closely related fungal genera include generally known and conserved functions. For the pan-genomes of the 36 Aspergillus species compared with the section Nigri species, the percentages of unknown function are similar (32% compared with 33%; Supplementary Tables 3d and 4a and Supplementary Fig. 2), as are the corresponding percentages for the core-genomes (14% compared with 17%, Fig. 2e; Supplementary Tables 3d and 4a). General functions like transporters, regulators, organelle-specific proteins, primary metabolism, and structural domains were found as core features across all 36 aspergilli (Supplementary Table 3f), which supports the general validity of the method.
We expected the section Nigri core-genome (gene families found in all the species of section Nigri but not in any other aspergilli examined) to contain Nigri signature genes, and we found this to be the case. These 1,214 gene families contain 580 InterPro domains conserved to a varying degree, including a many genes involved in the saprotrophic lifestyle and secondary metabolism (Supplementary Table 5). It is hypothesized that these genes are defining for the section compared with other aspergilli and will encode functions related to the phenotypes of species in this section.
Unique secondary metabolism genes in Aspergillus species
The genetic diversity seen in section Nigri led us to investigate whether the unique genes for each species show common trends in function. While these genes by definition do not have homologs in other species investigated in this work, we can predict general functions using InterPro domains. Unique genes of species in section Nigri matched 1,334 different InterPro domains (Supplementary Table 6a–c). Within the unique genes, we searched the list of InterPro domains in all sets of genes unique to individual section Nigri species (excluding the six A. niger isolates, to remove intraspecies redundancy). Surprisingly, we identified only ten domains that were found in nearly all Nigri species (25–26 species). Notably, nine of those are related to functions involved in secondary metabolism, gene regulation (transcription factors), or protein regulation (protein kinases) (Supplementary Table 7). Finding these functions in nearly all sets of species-specific genes suggests that secondary metabolite production and regulatory proteins are commonly identified as the species-‘unique’ genes and are therefore critical differentiates for fungal species at the genetic level.
Intra- and interspecies genetic variations are similar
We were interested in comparing the diversity between isolates of the same species to the diversity among species in the same clade. We thus compared six A. niger isolates to the eight closely related species in the A. tubingensis clade (Fig. 2d). The A. niger isolates have a high degree of genetic homogeneity, as 80% of the A. niger pan-genome is conserved across the six isolates and only 6% is unique to any of the isolates (Supplementary Fig. 3a). The same scale is seen in the A. tubingensis clade (77% shared pan-genome, 7% unique; Supplementary Fig. 3b). Moreover, the percentage of genes with predicted functional domains within the two groups is similar to that within section Nigri as a whole (Supplementary Fig. 4 and Supplementary Tables 4a and 8a,d). The unique genes belonging to each of the two groups are largely of unknown function (A. tubingensis clade 82%, A. niger complex 86%; Supplementary Tables 8a,d and 9a,b). The functions of the A. niger core-genome (3,798 domains) are, not surprisingly, very similar to those of section Nigri as a whole (Supplementary Tables 4c and 8c). In summary, the interspecies variation in the A. tubingensis clade is of the same scale as the intraspecies variation in the A. niger isolates, showing that large genetic variation does not directly translate to the currently circumscribed species.
Extra citrate synthase genes confer increased citrate
As species of section Nigri are known organic-acid producers, the genes involved in central metabolism are of interest, particularly given that the cause of citric acid overproduction in several of the section members is still not identified40. We thus analyzed the number of paralogs in central carbon metabolism in our set of 38 fungal genomes using a curated version of an A. niger genome-scale metabolic model41 as a source of pathway annotation (Fig. 3).
Proteins putatively catalyzing the same metabolic step (that is, complex subunits or functionally related enzymes) have been grouped in gray boxes. Species are sorted according to the phylogeny in Fig. 1. a, Glycolysis isoenzymes. b, Tricarboxylic acid cycle isoenzymes. c, Growth of strains on CREA medium, which turns yellow with lowered pH. A. niger CBS 513.88 has been mutagenized2.
The analysis of paralogs in glycolysis shows very little variance across the 32 Nigri genomes, similar to the variation in the 6 other fungal species genomes (Fig. 3a). For the tricarboxylic acid cycle (Fig. 3b), it is evident that certain metabolic steps in the pathway are conserved throughout all species, while others vary in paralog numbers. The biseriates are particularly homogeneous. These, along with four uniseriates, are also the primary citric acid-producing species in the section (Fig. 3c).
Of particular interest is the citrate production phenotype, and thus citrate synthase. All biseriates have one extra citrate synthase, and the four acid-producing uniseriates have two extra. Sequence alignment identified three distinct types (Supplementary Fig. 5), two of which are mitochondrial and are found in all species. All extra citrate synthase paralogs are of the third type, predicted to be cytosolic. We identified the extra biseriate citrate synthase (citB42) in a conserved 30 kB gene cluster including two transcription factor genes, a transporter gene, and two putative fatty acid synthase genes. We performed heterologous expression of the A. niger citB gene cluster in A. nidulans (which has only the two mitochondrial citrate synthases) using two constitutive promoters to control the transcription factors. This expression increased citrate concentrations by 42–52% (Supplementary Table 10). We hypothesize that this gene cluster may have a particular role in citrate production and additional undescribed functions involving the fatty acid synthase-like genes.
Carbon utilization is not correlated with carbohydrate-active enzyme content
Aspergilli have a particularly broad ability to degrade and convert plant biomass31. It is thus essential to examine the species diversity of this trait at the genotype and phenotype levels. We predicted the carbohydrate-active enzyme (CAZyme) gene content of the genomes across section Nigri (17,903 CAZyme domains; Fig. 4 and Supplementary Table 11) and performed growth profiling on plant biomass-related carbon sources (Supplementary Fig. 6). Growth on d-glucose was used to evaluate relative growth, showing variation between species.
Details on CAZy families are available in Supplementary Table 11. Growth profiles are available in Supplementary Fig. 6. All black aspergilli grew well on pectin and have a highly conserved and extensive set of genes encoding pectin-active enzymes. Growth on other plant polysaccharides such as xylan, starch, and guar gum was more variable, despite the presence of highly conserved genes related to xyloglucan and starch degradation. The growth and genetic variability on inulin are particularly high: nine species showed reduced growth. Moreover, endoinulinase (GH32 INU) is only present in eight of the black aspergilli, while the remainder of inulin-related genes (GH32 INV and INX) are more commonly present (Supplementary Table 11). However, the growth phenotypes show no correlation with the gene content (Supplementary Fig. 6).
In a previous study10, enzyme levels were measured in several black aspergilli, and significant differences were found. However, differences in enzyme levels do not reflect the copy number differences seen here (Supplementary Table 11). Considering the relative uniformity of the CAZyme content (Fig. 4), no correlation between genome content and growth on plant biomass-related carbon sources (Supplementary Fig. 6) was observed for the black aspergilli, suggesting that the differences in capability for plant biomass degradation reflect gene expression levels in the individual fungus. This confirms a proteome study of less-related aspergilli, in which the different response to plant biomass appeared to be mainly at the regulatory level43. The data suggest that this is the case for section Nigri: species-specific phenotypes are driven not generally by CAZyme content in closely related species, but by regulation.
Secondary metabolism in section Nigri contains 455 families
Secondary metabolism is thought to be a component of chemical defense, virulence, toxicity, mineral uptake, and communication in fungi44 and has a wide range of potential medical applications. As we had identified it to be commonly unique to individual species, we examined the exometabolite diversity of 37 Aspergillus and Penicillium species according to predictions of secondary metabolism gene clusters (SMGCs) as well as chemical profiles of the species of section Nigri on multiple substrates.
We identified 2,717 SMGCs in the 37 genomes. This is an even higher number of SMGCs per species than a previous study found in 24 Penicillium genomes45. We were further interested in quantifying the actual diversity of the SMGCs in section Nigri and in analyzing presence patterns of SMGCs across species. We therefore defined SMGC ‘families’ as genetically similar SMGCs across genomes ( Methods). Each SMGC family is expected to produce the same or similar compounds. This clustering resulted in the definition of 455 SMGC families across the 37 genomes (Supplementary Fig. 7), indicating the potential production of 455 different chemical families. Most families (82%) are found in fewer than 10 organisms, and 49% contain only one gene cluster (Supplementary Fig. 8 shows examples). On average there are 8.75 unique clusters per species, despite the close phylogenetic distance of the section.
Phylogenetic examination shows dynamic content of SMGCs
To reveal more about how SMGCs evolve and differentiate between species, each of the 455 SMGC families was characterized by the type of backbone enzyme and analyzed according to the phylogeny (Fig. 5a,b). Only five out of all SMGCs were present in all analyzed species, including clusters for the non-ribosomal peptide synthetase (NRPS)-derived siderophore ferrichrome, the circular NRP fungisporin46/nidulanin A47, and pigment (YWA1) synthesis. Two shared SMGC families were false predictions, namely two fatty acid synthases.
a, Cladogram of 36 Aspergillus (32 Nigri) genomes and one Penicillium genome. Numbers at nodes show SMGC families shared by species in the branch. End-points show families unique to an organism. b, Total SMGC family profile of the organism classified by backbone enzyme type. c, Presence/absence map of detected secondary metabolites in section Nigri species (Methods). Strains with no available metabolite data are marked in light gray. DMAT, dimethylallyltransferase (prenyl transferases); Hybrid, gene containing domains from NRPS and PKS backbones; NRPS, non-ribosomal peptide synthetase; NRPS-like, non-ribosomal peptide synthetase-like containing at least two NRPS-specific domains and another domain or one NRPS A domain in combination with NAD_binding_4 domain or short chain dehydrogenase; PKS, polyketide synthase; PKS-like, polyketide synthase-like, containing at least two PKS-specific domains and another domain; TC, terpene cyclase.
Examining the dynamics of the families, only 32% and 19% of SMGCs found in two or three organisms, respectively, follow the whole-genome phylogeny and suggest intragenus horizontal gene transfer or SMGC loss to be relatively common. As an example, an SMGC is found in five distantly related species (Supplementary Fig. 8b). The cluster is found in all A. niger isolates as well as in A. homomorphus, A. welwitschiae, A. sclerotiicarbonarius, and A. brasiliensis.
As seen in Fig. 5a, the presence of unique SMGC families at every major branch point in the phylogenetic tree supports that SMGCs are a part of what sets the species apart: all biseriates share a previously undescribed polyketide synthase (PKS) and an NRPS-like protein, the A. carbonarius clade a terpene cyclase, and the remaining biseriates share another PKS and another NRPS-like protein. Furthermore, the A. niger complex and A. tubingensis clade each share unique PKS genes. Uniseriates share four unique previously undescribed SMGC families (Fig. 5a). Examinations of individual species reveal that every single section Nigri species has a unique combination of SMGCs (Fig. 3b). Furthermore, nearly all Nigri species genomes (with the exception of A. tubingensis, A. niger, A. brasiliensis, and A. vadensis) encode one or more unique SMGCs. These patterns show the existence of high diversity of SMGCs between species and of a homogeneous set of SMGCs within isolates from the same species.
Correlating secondary metabolisms with SMGC families links gene to function
As a further application of the constructed SMGC families, we hypothesized that we can correlate SMGC families to classes of compounds. We performed extensive exometabolome analysis of 27 of the sequenced strains and identified 35 compound families (Fig. 5c and Supplementary Table 12).
The most abundant group was naphtha-γ-pyrones, of which aurasperone B29 was identified in 14 of the isolates. We compared the presence patterns of SMGC families with the compound class (Fig. 5c) and combined it with a knowledge-based filtering of InterPro domains leaving one hit (Methods and Supplementary Fig. 8d). The candidate SMGC family is a nine-gene cluster found in 18 genomes—including the 14 where we detected the compound—and it contains all activities needed to synthesize aurasperone. In support of this identification, an SMGC for a closely related compound, aurofusarin, has been experimentally verified in Fusarium graminearum48. The aurasperone cluster shares six genes (one of which is a duplication) with the aurofusarin cluster. This finding supports the assignment of this family of SMGCs to the production of aurasperone B and conceptually justifies this approach for efficient linking of clusters to compounds. We see this correlation approach as highly useful for future elucidation of fungal metabolites.
Discussion
We have sequenced the genomes of a whole section of filamentous fungi, and a diverse set of A. niger isolates, and found that the species are highly diverse in some traits, in particular secondary metabolism and to a lesser extent regulatory proteins, and homogeneous in others, such as glycolytic metabolism and CAZymes. The presented data furthermore provide an extensive compendium of 24 new genomes, which adds substantial information on fungal genetic diversity. We further combined genome analysis with metabolite profiling and heterologous gene expression to identify the genetic basis of several phenotypes within primary and secondary metabolism.
Of particular interest was the finding that the species-specific genes in all species share functions within gene/protein regulation and secondary metabolism, showing that unique sets of these functions exist for all species in the investigated set.
URLs
Carbohydrate-Active enZYmes Database, http://www.cazy.org; RoerdamAndersenLab GitHub repositories of scripts for comparative genomics, https://github.com/RoerdamAndersenLab/; HGAP Assembly, http://files.pacb.com/software/smrtanalysis/2.2.0/doc/smrtportal/help/!SSL!/Webhelp/CS_Prot_RS_HGAP_Assembly3.htm; JGI fungal genome portal MycoCosm, http://jgi.doe.gov/fungi; JGI website, http://jgi.doe.gov; Joint BioEnergy Institute website, http://www.jbei.org; Technical University of Denmark (DTU) Bioinformatics TargetP web server, http://www.cbs.dtu.dk/services/TargetP/.
Methods
Fungal strains
Unless otherwise noted, the species examined were taken from the IBT Culture Collection of Fungi at DTU. Strains employed in this study are denoted in Supplementary Table 1.
Purification of DNA and RNA
For all sequences generated for this study (Supplementary Table 1), spores were defrosted from storage at −80 °C and inoculated onto solid CYA medium. Fresh spores were harvested after 7–10 d and suspended in a 0.1% Tween solution. Spores were stored in solution at 5 °C for up to 3 weeks. Biomass for all fungal strains was obtained from shake flasks containing 200 ml of complex medium, either CYA, MEAox, or CY20 depending on the strain (see Supplementary Table 1) cultivated for 5–10 d at 30 °C. Biomass was isolated by filtering through Miracloth (Millipore, 475855-1R), freeze dried, and stored at 80 °C. DNA isolation was performed using a modified version of the standard phenol extraction (ref. 52 and below) and checked for quality and concentration using a NanoDrop (BioNordika). RNA isolation was performed using the Qiagen RNeasy Plant Mini Kit according to the manufacturer’s instructions.
A sample of frozen biomass was subsequently used for RNA purification. First, hyphae were transferred to a 2 ml microtube together with a 5 mm steel bead (Qiagen), placed in liquid nitrogen, then lysed using the Qiagen TissueLyser LT at 45 Hz for 50 s. Then the Qiagen RNeasy Mini Plus Kit was used to isolate RNA. RLT Plus buffer (with 2-mercaptoethanol) was added to the samples, vortexed, and spun down. The lysate was then used in step 4 in the instructions provided by the manufacturer, and the protocol was followed from this step. For genomic DNA, a protocol based on Fulton et al.53 was used (See Supplementary Note).
DNA and RNA sequencing and assembly
All genomes in this study, except for those of A. heteromorphus, A. eucalypticola, and A. sclerotioniger, and all transcriptomes were sequenced with Illumina. The genomes of A. heteromorphus, A. eucalypticola, and A. sclerotioniger were sequenced with PacBio.
For all genomic Illumina libraries, 100 ng of DNA was sheared to 270 bp fragments using the Covaris LE220 (Covaris) and size selected using SPRI beads (Beckman Coulter). The fragments were treated with end-repair and A-tailing and ligated to Illumina-compatible adapters (IDT) using the KAPA-Illumina library creation kit (KAPA Biosystems).
For transcriptomes, stranded complementary DNA libraries were generated using the Illumina TruSeq Stranded Total RNA LT Sample Prep Kit. Messenger RNA (mRNA) was purified from 1 µg of total RNA using magnetic beads containing poly(T) oligos. mRNA was fragmented using divalent cations and high temperature. The fragmented RNA was reverse transcribed using random hexamers and SSII (Invitrogen) followed by second-strand synthesis. The fragmented complementary DNA was treated with end-pair, A-tailing, adapter ligation, and 10 cycles of PCR.
The prepared libraries were quantified using KAPA Biosystems’ next-generation sequencing library quantitative PCR kit and run on a Roche LightCycler 480 real-time PCR instrument. The quantified libraries were then multiplexed with other libraries, and library pools were prepared for sequencing on the Illumina HiSeq sequencing platform using a TruSeq paired-end cluster kit, v3, and Illumina’s cBot instrument to generate clustered flow cells for sequencing. Sequencing of the flow cells was performed on the Illumina HiSeq2000 sequencer using a TruSeq SBS sequencing kit, v3, following a 2 × 150 indexed run recipe.
After sequencing, the genomic FASTQ files were quality control-filtered to remove artifacts/process contamination and assembled using Velvet54. The resulting assemblies were used to create in silico long mate-pair libraries with inserts of 3,000 ± 90 bp, which were then assembled with the target FASTQ using AllPathsLG release version R4771055. Illumina transcriptome reads were assembled into consensus sequences using Rnnotator v3.3.256.
For the genomes of A. heteromorphus, A. eucalypticola, and A. sclerotioniger, amplified libraries were generated using a modified shearing version of the Pacific Biosciences standard template preparation protocol. To generate each library, 5 μg of genomic DNA was used. The DNA was sheared using a Covaris LE220 focused-ultrasonicator with their Red miniTUBES to generate fragments 5 kb in length. The sheared DNA fragments were then prepared according to the Pacific Biosciences protocol using their SMRTbell template preparation kit, where the fragments were treated with DNA damage repair (ends were repaired so that they were blunt ended and 5′ phosphorylated). Pacific Biosciences hairpin adapters were then ligated to the fragments to create the SMRTbell template for sequencing. The SMRTbell templates were then purified using exonuclease treatments and size-selected using AMPure PB beads.
Sequencing primer was then annealed to the SMRTbell templates, and version P4 sequencing polymerase was bound to them. The prepared SMRTbell template libraries were sequenced on a Pacific Biosciences RS II sequencer using version C2 chemistry and 2 h sequencing movie run times. The three Pacific Biosciences genome datasets were assembled using HGAP3 (see URLs).
All genomes were annotated using the JGI annotation pipeline33. Genome assembly and annotations are available at the JGI fungal genome portal MycoCosm33 (see URLs) and have been deposited in the DNA Data Bank of Japan (DDBJ)/European Molecular Biology Laboratory (EMBL)/GenBank under the following accession numbers: A. aculeatinus (PSTE00000000), A. brunneoviolaceus (PSTC00000000), A. costaricaensis (PSTH00000000), A. ellipticus (PSSY00000000), A. eucalypticola (MSFU00000000), A. fijiensis (PSTG00000000), A. heteromorphus (MSFL00000000), A. homomorphus (PSTJ00000000), A. ibericus (PSTI00000000), A. indologenus (PSTB00000000), A. japonicus (PSTF00000000), A. lacticoffeatus (MSFR00000000), A. neoniger (MSFP00000000), A. niger ATCC 13157 (A. phoenicis) (QQUR00000000), A. niger ATCC 13496 (QQZP00000000), A. piperis (PSTD00000000), A. saccharolyticus (MSFQ00000000), A. sclerotiicarbonarius (PSSZ00000000), A. sclerotioniger (MSFK00000000), A. uvarum (MSFT00000000), A. vadensis (MSFS00000000), A. violaceofuscus (PSTA00000000), and A. welwitschiae (QQZQ00000000).
See also the Nature Research Reporting Summary linked to this article.
Analysis of secondary metabolism
Cultivation for secondary metabolite analysis
Fungal strains were cultivated as three-point cultures on CYA, CYAS29, and YES media for 7 d in the dark at 25 °C. Three 6 mm inner diameter plugs taken across the cultures were then extracted using an (3:2:1) (ethylacetate–dichloromethane–methanol) mixture and dissolved in methanol57.
Extraction of fungal metabolites
Fungal metabolite extracts were prepared using one of the three following methods29: (1) chloroform–methanol–acetone–ethylacetate extraction, (2) micro-extraction using methanol–dichloromethane–ethylacetate, or (3) 75% methanol extraction.
Chemical analysis of secondary metabolites
All chemical analyses were done by reversed-phase ultrahigh-performance liquid chromatography (UHPLC) coupled to ultraviolet–visible diode array detection (DAD) combined with either fluorescence detection (FLD) or high-resolution mass spectrometry (HRMS). Three different methods were used:
Method 1
Pure UHPLC–DAD–FLD was performed using a rapid-separation liquid chromatography (RSLC) UltiMate 3000 system (Dionex) linked to an 1100 Series FLD (Agilent). The system was equipped with an Agilent Poroshell phenyl-hexyl column (150 × 2.1 mm, 2.6 μm) and was run using a linear gradient of water–acetonitrile starting at 10% acetonitrile and increasing to 100% (both containing 50 ppm trifluoroacetic acid) over 8 min, then using 100% acetonitrile for 2 min. The column temperature was 60 °C, the flow rate 0.8 ml min−1, and the injection volume was 1 µl. The ultraviolet spectra 200–640 nm were matched against our internal database29.
Method 2
UHPLC–DAD–HRMS was conducted on a Dionex RSLC UltiMate system linked to a maXis high-definition quadrupole-time-of-flight mass spectrometer (Q-TOF MS) (Bruker Daltonics). Separation was done on a Kinetex C18 column (100 × 2.1 mm, 2.6 μm), with a linear gradient consisting of water and acetonitrile (both buffered with 20 mM formic acid), starting at 10% acetonitrile and increasing to 100% over 10 min, where it was held for 2 min and returned (0.4 ml min−1, 40 °C). Injection volume, depending on sample concentration, typically varied between 0.1 and 1 µl. Some samples were analyzed in electrospray ionization (ESI)+ and some in ESI− full-scan mode, scanning m/z 100–1,250. Data were analyzed by aggressive dereplication55 using lists of compounds considered to be from black aspergilli only (~350); a list with all Aspergillus compounds (~2,450); and a list of 1,600 reference standards, of which 500 are known to come from Aspergillus. Unknown peaks were matched against Antibase2012 and dereplicated using accurate mass, isotope patterns, adduct patterns, log D, and ultraviolet–visible data58.
Method 3
UHPLC–DAD–HRMS was conducted on an Agilent Infinity 1290 UHPLC system coupled to an Agilent 6550 Q-TOF MS. Separation was obtained on an Agilent Poroshell 120 phenyl-hexyl column (2.1 × 250 mm, 2.7 µm) using a linear gradient of water and acetonitrile (both buffered with 20 mM formic acid), progressing from 10% to 100% acetonitrile over 15 min, where it was held for 2 min. The flow was 0.35 ml min−1 and the temperature 60 °C. Injection volume was between 0.1 and 1 µl, depending on the sample concentration.
Some samples were analyzed in ESI+ and some in ESI− full-scan mode, scanning m/z 100–1,700 and with automatic MS/MS enabled for ion counts above 100,000 and with a quarantine time of 0.06 min. MS/MS spectra were obtained at 10, 20, and 40 eV (ref. 59).
Full-scan data were analyzed as above in MassHunter59. MS/MS data were matched to our internal MS library (~1,700 compounds) of reference standards and tentatively identified compounds59.
Genome annotation and analysis
Genome annotation
All genomes were annotated based on the JGI annotation pipeline34 as previously described60.
Swiss-Prot comparison
Swiss-Prot comparisons were done using protein BLAST alignments with BLAST+ (v2.3.0), e-value cut-off 1 × 10−5, -max_target_seqs 100, -max_hsps 1, and locally optimal Smith-Waterman alignments (-use_sw_tback).
Whole-genome phylogeny
Protein sequences of all organisms were compared using BLASTp (e-value cut-off 1 × 10−5). Orthologous groups of sequences were constructed on the basis of best bidirectional hits. Two hundred groups with a member from each species were selected, and the sequences of each organism were concatenated into one long protein sequence. Concatenated sequences were aligned using MAFFT (thread 16), and well-aligned regions were extracted using gBlocks (−t = p; −b4 = 5; −b5 = h). Trees were then constructed using multithreaded RAxML, the PROTGAMMAWAG model, and 100 bootstrap replicates.
Prediction of SMGCs
For the prediction of SMGCs, we developed a command-line Python script roughly following the SMURF algorithm:
According to SMURF the following genes were predicted as a ‘backbone’ genes:
-
Genes that have at least three PFAM domains—ketoacyl-synt (PF00109), Ketoacyl-synt_C (PF02801), and Acyl_transf_1 (PF00698)—were predicted as ‘PKS’ genes.
-
Genes that have ketoacyl-synt (PF00109) and Ketoacyl-synt_C (PF02801) but not Acyl_transf_1 (PF00698) were predicted as ‘PKS-like’ genes.
-
Genes that have at least the three domains AMP-binding (PF00501), PP-binding (PF00550), and Condensation (PF00668) were predicted as ‘NRPS’ genes.
-
Genes that have an AMP-binding (PF00501) domain and at least one of the domains PP-binding (PF00550), Condensation (PF00668), NAD_binding_4 (PF07993), and Epimerase (PF01370) were predicted as ‘NRPS-like’ genes.
-
Genes that have both ‘PKS’ and ‘NRPS’ domains were predicted as ‘Hybrid’ genes.
-
Genes that have a Trp_DMAT domain were predicted as ‘DMAT’ genes.
-
Genes that have Terpene_synth (PF01397) or Terpene_synth_C (PF03936) domains were predicted as ‘Terpene cyclase/synthase’ genes.
Secondary metabolite-specific PFAM domains were taken from Supplementary Table 2 of the SMURF paper61.
As input, the program takes genomic coordinates and the annotated PFAM domains of the predicted genes. Based on the multidomain PFAM composition of identified ‘backbone’ genes, it can predict seven types of secondary metabolite clusters: (1) polyketide synthases (PKSs), (2) PKS-like, (3) non-ribosomal peptide-synthetases (NRPSs), (4) NRPS-like, (5) hybrid PKS-NRPS, (6) prenyltransferases (DMATS), and (7) terpene cyclases (TCs). Besides backbone genes, PFAM domains, which are enriched in experimentally identified secondary metabolite clusters (secondary metabolite-specific PFAMs), were used in determining the borders of gene clusters. The maximum allowed size of intergenic regions in a cluster was set to 3 kb, and each predicted cluster was allowed to have up to 6 genes without secondary metabolite-specific domains.
Prediction of secreted proteases
Secretome prediction was done using an in-house adaptation of SignalP62.
Gene-compound assignment
Identification of conserved or highly similar fungal gene clusters was performed on the basis of the gene cluster predictions above. The genomes were compared using the BLASTp function from the BLAST+ suite63. Presence/absence of an orthologous gene to a member in a gene cluster was based on a bidirectional best hit, with e < 1 × 10−100 and coverage of >90%. Presence/absence of a full gene cluster was based on the occurrence of the majority of the predicted members in a gene cluster, including the backbone synthetase in another species.
Detection of encoded CAZymes
Each Aspergillus protein model was compared using BLASTp to proteins listed in the Carbohydrate-Active enZYmes database (CAZy)64. Models with over 50% identity over the entire length of an entry in CAZy were directly assigned to the same family (or subfamily when relevant). Proteins with less than 50% identity to a protein in CAZy were all manually inspected, and conserved features, such as the catalytic residues, were searched whenever known. Because 30% sequence identity results in widely different e-values (from non-significant to highly significant), for CAZy family assignments, we examined sequence conservation (percentage identity over CAZy domain length). Sequence alignments with isolated functional domains were performed in the case of multimodular CAZymes. The same methods were used for Penicillium chrysogenum and Neurospora crassa.
Mapping of genes shared by groups of species
All predicted sets of protein sequences for the 38 genomes analyzed were aligned using the BLASTp function from the BLAST+ suite version 2.2.27 (e-value cut-off ≤1 × 10−10). These 1.444 whole-genome BLAST tables were analyzed to identify bidirectional hits in all pairwise comparisons. Using custom Python scripts, homologs were identified within and across the genomes and grouped into sequence-similar families using single linkage, if they met the following criterion: The sum of the alignment coverage between the pairwise sequences was >130%, the alignment identity between the pairwise sequences was >50%, and the hit must be found in both of the species’ BLAST output (reciprocal hits). Singletons were assigned a family having only one gene member. This allowed for identification of species-unique genes as well as genes shared by sections, clades, and sub-clades of species. All homologs were assigned functional and structural domains using InterPro version 4865 and checked for annotation and sequencing errors by investigating scaffold location and sequence identity.
For the analysis of the pan- and core-genomes of a subset of 38 fungal species used in this study, the orthologous and paralogous families were subsetted to include only the species of interest. Therefore, the genes representing the core and unique portion of the genomes will adjust relative to the accompanying species.
Identification of SMGC families
Our implementation of SMURF was run on genomic data from 37 Aspergillus strains. Proteins of the resulting SMGCs were compared with each other by alignment using BLASTp (BLAST+ suite version 2.2.27, e-value ≤1 × 10−10). Subsequently, a score based on BLASTp identity and shared proteins was created to determine the similarity between gene clusters as depicted in the formula below. Using these scores, we created a weighted network of SMGC clusters and used a random walk community detection algorithm (R version 3.3.2, igraph_1.0.166) to determine families of SMGC clusters. Finally, we ran another round of random walk clustering on the communities that contained more members than species in the analysis (ptailoring/pbackbone = sum of percentage BLAST alignment of tailoring/backbone enzymes, respectivly; ntailoring/nbackbone = number of tailoring/backbone enzymes with significant hits, respectivly; ttailoring/tbackbone = total number of tailoring/backbone enzymes):
To create a cluster similarity score, a combined score of tailoring and backbone enzymes was created. The sum of the BLASTp percent identity (ptailoring/pbackbone) of all hits for tailoring enzymes between two clusters was divided by the maximum amount of tailoring enzyme (ttailoring/tbackbone) and multiplied by 0.35. Then the score for the backbone enzymes was calculated in the same manner but multiplied by 0.65 to give more weight to the backbone enzymes. The scores were added to create an overall cluster similarity score:
Identification of shared SMGC families at nodes of the phylogenetic tree
A list containing organisms of each branch of the phylogenetic tree was created and compared with the list of organisms for each SMGC family. If all organisms of a family matched, the count on the corresponding node was increased by one.
Prediction of the aurasperone B gene cluster
Lists of organisms for all SMGC families were compared with the lists of aurasperone B-producing species and filtered for InterPro annotations containing the terms ‘cytochrome P450’ or ‘methyltransferase’.
Primary metabolism
Copy numbers were assessed using the homologous protein families generated during the analysis of genome diversity. The gene pathway associations were taken from the A. niger genome-scale model41. All proteins in the respective protein families were considered putative isozymes and were included in the copy number analyses.
Comparing the putative isoenzymes in the different species, gene sequences were aligned and clustered using neighbor-joining with MUSCLE v3.8.3168. Resulting trees were visualized and edited for publication using the Python ETE Toolkit67. Subcellular localizations for the genes included in the analysis were predicted using the TargetP69 web server (see URLs).
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Code availability
Code for generation of gene families of homologs and for generation of SMGC families is available from GitHub (see URLs).
Data availability
All genomes used in the study are available from Joint Genome Institute fungal genome portal MycoCosm (http://jgi.doe.gov/fungi). All new genomes published in the study have been deposited in the DNA Data Bank of Japan (DDBJ)/European Molecular Biology Laboratory (EMBL)/GenBank under the following accessions: A. aculeatinus (PSTE00000000), A. brunneoviolaceus (PSTC00000000), A. costaricaensis (PSTH00000000), A. ellipticus (PSSY00000000), A. eucalypticola (MSFU00000000), A. fijiensis (PSTG00000000), A. heteromorphus (MSFL00000000), A. homomorphus (PSTJ00000000), A. ibericus (PSTI00000000), A. indologenus (PSTB00000000), A. japonicus (PSTF00000000), A. lacticoffeatus (MSFR00000000), A. neoniger (MSFP00000000), A. niger ATCC 13157 (A. phoenicis) (QQUR00000000), A. niger ATCC 13496 (QQZP00000000), A. piperis (PSTD00000000), A. saccharolyticus (MSFQ00000000), A. sclerotiicarbonarius (PSSZ00000000), A. sclerotioniger (MSFK00000000), A. uvarum (MSFT00000000), A. vadensis (MSFS00000000), A. violaceofuscus (PSTA00000000), and A. welwitschiae (QQZQ00000000).
References
Nierman, W. C. et al. Genomic sequence of the pathogenic and allergenic filamentous fungus Aspergillus fumigatus. Nature 438, 1151–1156 (2005).
Pel, H. J. et al. Genome sequencing and analysis of the versatile cell factory Aspergillus niger CBS 513.88. Nat. Biotechnol. 25, 221–231 (2007).
Machida, M. et al. Genome sequencing and analysis of Aspergillus oryzae. Nature 438, 1157–1161 (2005).
Galagan, J. E. et al. Sequencing of Aspergillus nidulans and comparative analysis with A. fumigatus and A. oryzae. Nature 438, 1105–1115 (2005).
Papagianni, M. Advances in citric acid fermentation by Aspergillus niger: biochemical aspects, membrane transport and modeling. Biotechnol. Adv. 25, 244–263 (2007).
Punt, P. J. et al. Filamentous fungi as cell factories for heterologous protein production. Trends Biotechnol. 20, 200–206 (2002).
Currie, J. The citric acid fermentation of Aspergillus niger. J. Biol. Chem. 31, 15–37 (1917).
Andersen, M. R. et al. Comparative genomics of citric-acid-producing Aspergillus niger ATCC 1015 versus enzyme-producing CBS 513.88. Genome Res. 21, 885–897 (2011).
Wösten, H., Scoltmeijer, K. & de Vries, R. in Food Mycology: A Multifaceted Approach to Fungi and Food (eds. Dijksterhuis, J. & Samson, R.) 183–196 (CRC Press, Boca Raton, FL, USA, 2007).
Meijer, M., Houbraken, J. A. M. P., Dalhuijsen, S., Samson, R. A. & de Vries, R. P. Growth and hydrolase profiles can be used as characteristics to distinguish Aspergillus niger and other black aspergilli. Stud. Mycol. 69, 19–30 (2011).
List of Commercial Enzymes (Association of Manufacturers and Formulators of Enzyme Products, Brussels, 2009).
Workman, M., Andersen, M. R. & Thykaer, J. Integrated approaches for assessment of cellular performance in industrially relevant filamentous fungi. Ind. Biotechnol. 9, 337–344 (2013).
Hong, S.-B. et al. Aspergillus luchuensis, an industrially important black Aspergillus in East Asia. PLoS ONE 8, e63769 (2013).
Perrone, G. et al. Aspergillus niger contains the cryptic phylogenetic species A. awamori. Fungal Biol. 115, 1138–1150 (2011).
Futagami, T. et al. Genome sequence of the white koji mold Aspergillus kawachii IFO 4308, used for brewing the Japanese distilled spirit shochu. Eukaryot. Cell 10, 1586–1587 (2011).
Abarca, M. L., Bragulat, M. R., Castella, G. & Cabanes, F. J. Ochratoxin A production by strains of Aspergillus niger var. niger. Appl. Environ. Microbiol. 60, 2650–2652 (1994).
Frisvad, J. C., Smedsgaard, J., Samson, R. A., Larsen, T. O. & Thrane, U. Fumonisin B2 production by Aspergillus niger. J. Agric. Food Chem. 55, 9727–9732 (2007).
Frisvad, J. C. et al. Fumonisin and ochratoxin production in industrial Aspergillus niger strains. PLoS ONE 6, e23496 (2011).
Perrone, G. et al. Biodiversity of Aspergillus species in some important agricultural products. Stud. Mycol. 59, 53–66 (2007).
Monod, M. et al. Secreted proteases from pathogenic fungi. Int. J. Med. Microbiol. 292, 405–419 (2002).
Jurjević, Z. et al. Two novel species of Aspergillus section Nigri from indoor air. IMA Fungus 3, 159–173 (2012).
Varga, J. et al. New and revisited species in Aspergillus section Nigri. Stud. Mycol. 69, 1–17 (2011).
Samson, R. A., Houbraken, J. A. M. P., Kuijpers, A. F. A., Frank, J. M. & Frisvad, J. C. New ochratoxin A or sclerotium producing species in Aspergillus section Nigri. Stud. Mycol. 50, 45–61 (2004).
Samson, R. A. et al. Diagnostic tools to identify black aspergilli. Stud. Mycol. 59, 129–145 (2007).
Samson, R. A. et al. Phylogeny, identification and nomenclature of the genus Aspergillus. Stud. Mycol. 78, 141–173 (2014).
Visagie, C. M. et al. Aspergillus, Penicillium and Talaromyces isolated from house dust samples collected around the world. Stud. Mycol. 78, 63–139 (2014).
Rajendran, C. & Muthappa, B. N. Saitoa, a new genus of Plectomycetes. Proc. Plant Sci. 89, 185–191 (1980).
Frisvad, J. C. et al. Formation of sclerotia and production of indoloterpenes by Aspergillus niger and other species in section Nigri. PLoS ONE 9, e94857 (2014).
Nielsen, K. F., Mogensen, J. M., Johansen, M., Larsen, T. O. & Frisvad, J. C. Review of secondary metabolites and mycotoxins from the Aspergillus niger group. Anal. Bioanal. Chem. 395, 1225–1242 (2009).
Yamada, O. et al. Genome sequence of Aspergillus luchuensis NBRC 4314. DNA Res. 23, 507–515 (2016).
de Vries, R. P. et al. Comparative genomics reveals high biological diversity and specific adaptations in the industrially and medically important fungal genus Aspergillus. Genome Biol. 18, 28 (2017).
Kozakiewicz, Z. et al. Proposals for nomina specifica conservanda and rijicienda in Aspergillus and Penicillium (Fungi). Taxon 41, 109–113 (1992).
Grigoriev, I. V. et al. MycoCosm portal: gearing up for 1000 fungal genomes. Nucleic Acids Res. 42, 699–704 (2014).
Grigoriev, I. V., Martinez, D. A. & Salamov, A. A. Fungal genomic annotation. Appl. Mycol. Biotechnol. 6, 123–142 (2006).
Hunter, S. et al. InterPro: the integrative protein signature database. Nucleic Acids Res. 37, D211–D215 (2008).
Mitchell, A. et al. The InterPro protein families database: the classification resource after 15 years. Nucleic Acids Res. 43, D213–D221 (2015).
Sørensen, A., Lübeck, P. S., Lübeck, M., Teller, P. J. & Ahring, B. K. β-Glucosidases from a new Aspergillus species can substitute commercial β-glucosidases for saccharification of lignocellulosic biomass. Can. J. Microbiol. 57, 638–650 (2011).
Sørensen, A. et al. Identifying and characterizing the most significant β-glucosidase of the novel species Aspergillus saccharolyticus. Can. J. Microbiol. 58, 1035–1046 (2012).
Szöllősi, G. J., Davín, A. A., Tannier, E., Daubin, V. & Boussau, B. Genome-scale phylogenetic analysis finds extensive gene transfer among fungi. Philos. Trans. R. Soc. B Biol. Sci. 370, 20140335 (2015).
Karaffa, L. & Kubicek, C. P. Aspergillus niger citric acid accumulation: do we understand this well working black box? Appl. Microbiol. Biotechnol. 61, 189–196 (2003).
Andersen, M. R., Nielsen, M. L. & Nielsen, J. Metabolic model integration of the bibliome, genome, metabolome and reactome of Aspergillus niger. Mol. Syst. Biol. 4, 178 (2008).
Hossain, A. H. et al. Rewiring a secondary metabolite pathway towards itaconic acid production in Aspergillus niger. Microb. Cell Fact. 15, 130 (2016).
Benoit, I. et al. Closely related fungi employ diverse enzymatic strategies to degrade plant biomass. Biotechnol. Biofuels 8, 107 (2015).
Fox, E. M. & Howlett, B. J. Secondary metabolism: regulation and role in fungal biology. Curr. Opin. Microbiol. 11, 481–487 (2008).
Nielsen, J. C. et al. Global analysis of biosynthetic gene clusters reveals vast potential of secondary metabolite production in Penicillium species. Nat. Microbiol. 2, 17044 (2017).
Ali, H. et al. A non-canonical NRPS is involved in the synthesis of fungisporin and related hydrophobic cyclic tetrapeptides in Penicillium chrysogenum. PLoS ONE 9, e98212 (2014).
Andersen, M. R. et al. Accurate prediction of secondary metabolite gene clusters in filamentous fungi. Proc. Natl Acad. Sci. USA 110, E99–E107 (2013).
Frandsen, R. J. N. et al. The biosynthetic pathway for aurofusarin in Fusarium graminearum reveals a close link between the naphthoquinones and naphthopyrones. Mol. Microbiol. 61, 1069–1080 (2006).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Katoh, K. & Standley, D. M. MAFFT: multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Talavera, G. & Castresana, J. Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst. Biol. 56, 564–577 (2007).
Sambrook, J. & W Russell, D. Molecular Cloning: A Laboratory Manual. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 2012).
Fulton, T. M., Chunwongse, J. & Tanksley, S. D. Microprep protocol for extraction of DNA from tomato and other herbaceous plants. Plant Mol. Biol. Rep. 13, 207–209 (1995).
Zerbino, D. R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Gnerre, S. et al. High-quality draft assemblies of mammalian genomes from massively parallel sequence data. Proc. Natl Acad. Sci. USA 108, 1513–1518 (2011).
Martin, J. et al. Rnnotator: an automated de novo transcriptome assembly pipeline from stranded RNA-Seq reads. BMC Genomics 11, 663 (2010).
Smedsgaard, J. Micro-scale extraction procedure for standardized screening of fungal metabolite production in cultures. J. Chromatogr. A 760, 264–270 (1997).
Klitgaard, A. et al. Aggressive dereplication using UHPLC-DAD-QTOF: screening extracts for up to 3000 fungal secondary metabolites. Anal. Bioanal. Chem. 406, 1933–1943 (2014).
Kildgaard, S. et al. Accurate dereplication of bioactive secondary metabolites from marine-derived fungi by UHPLC-DAD-QTOFMS and a MS/HRMS library. Mar. Drugs 12, 3681–3705 (2014).
Kis-Papo, T. et al. Genomic adaptations of the halophilic Dead Sea filamentous fungus Eurotium rubrum. Nat. Commun. 5, 3745 (2014).
Khaldi, N. et al. SMURF: genomic mapping of fungal secondary metabolite clusters. Fungal Genet. Biol. 47, 736–741 (2010).
Bendtsen, J., Nielsen, H., von Heijne, G. & Brunak, S. Improved prediction of signal peptides: SignalP 3.0. J. Mol. Biol. 340, 783–795 (2004).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Li, L. et al. The Carbohydrate-Active EnZymes database (CAZy): an expert resource for glycogenomics. Nucleic Acids Res. 37, D233–D238 (2014).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Csárdi, G. & Nepusz, T. The igraph software package for complex network research. InterJournal Complex Syst. 1695, 1–9 (2006).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Huerta-Cepas, J., Dopazo, J. & Gabaldón, T. ETE: a python environment for tree exploration. BMC Bioinformatics 11, 24 (2010).
Emanuelsson, O., Nielsen, H., Brunak, S. & von Heijne, G. Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J. Mol. Biol. 300, 1005–1016 (2000).
Acknowledgements
T.C.V., J.L.N., S.T., and M.R.A. acknowledge funding from The Villum Foundation/Villum Fonden, grant VKR023437. J.B. and M.R.A. acknowledge funding from the Novo Nordisk Foundation, grant NNF13OC0004831. M.R.A. and T.C.V. acknowledge funding from the Danish National Research Foundation (DNRF137) for the Center for Microbial Secondary Metabolites. J.C.F. acknowledges funding from the Novo Nordisk Foundation, grant NNF13OC0005201. Work performed at the US Department of Energy (DOE) Joint BioEnergy Institute is supported by the US DOE, Office of Science, Office of Biological and Environmental Research, through Contract No. DE-AC02-05CH11231 between Lawrence Berkeley National Laboratory and the US DOE. The work conducted by the US DOE Joint Genome Institute, a DOE Office of Science User Facility, is supported by the Office of Science of the US DOE under Contract No. DE-AC02-05CH11231 between Lawrence Berkeley National Laboratory and the US DOE. S. Mondo is acknowledged for GenBank submission of 20 of the genomes. I. Kjærbølling is acknowledged for critical and constructive feedback on the manuscript and figures.
Author information
Authors and Affiliations
Contributions
J.L.N. and S.T. analyzed data, designed and generated algorithms, contributed to design of research, and wrote parts of the manuscript. J.C.F. contributed to design of research, contributed analytical tools and data for species selection and verification, wrote parts of the manuscript, and analyzed data. T.O.L. and K.F.N. generated data on secondary metabolism, analyzed data, and wrote parts of the manuscript. J.B.H. and M.T.N. engineered strains, analyzed data, and wrote parts of the manuscript. J.B. analyzed data on primary metabolism and wrote parts of the manuscript. A.S., R.R., A.K., and S.H. annotated genomes and analyzed data. J.M.G. and J.K.M. contributed to design of research and contributed analytical tools. P.P. contributed analytical tools. E.K.L. and M.E.K. contributed to design of research, developed methods, conducted experiments, wrote parts of the manuscript, and analyzed data. C.C. and L.S. sequenced RNA and DNA. M.N., A.L., K.L., and A.C. assembled the genomes. M.H., E.D., and B.H. contributed analytical tools and analyzed CAZyme data. A.W. performed part of the experiments. M.R.M. analyzed data and wrote part of the manuscript. R.P.d.V. analyzed data and wrote part of the manuscript. A.T., K.S., and E.M. contributed to design of research, contributed analytical tools and data, and analyzed data. K.B. and I.V.G. coordinated the DNA and RNA sequencing and annotation. B.A.S. contributed to design of research. U.H.M. designed parts of the experiments and developed methods. S.E.B. conceived the overall project, analyzed data, contributed to design of research, and contributed to writing and editing the manuscript. T.C.V. and M.R.A. conceived the overall project, analyzed data, contributed to design of research, designed algorithms, wrote parts of the manuscript, and coordinated the project. All authors commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Note
Supplementary Table 1
Genome statistics of sequencing and annotation
Supplementary Table 4
InterPro domains for the core genome specific to section Nigri
Supplementary Table 5
InterPro domains in the unique gene set
Supplementary Table 6
InterPro domains potentially involved in general speciation within section Nigri without A. niger isolates
Supplementary Table 7
InterPro coverage of the Niger clade species
Supplementary Table 8
InterPro domains in the unique gene set
Supplementary Table 9
Overview of areas under the curve for citrate production using HPLC quantification
Supplementary Table 10
Overviews of the number of glycoside hydrolases (GH), glycosyl transferases (GT), polysaccharide lyases (PL), carbohydrate esterases (CE), carbohydrate-binding modules (CBM) and auxiliary activities (AA) in 37 genomes included in the study
Supplementary Table 11
Overview of SM families detected in 27 different Aspergillus isolates
Supplementary Table 12
All hfams, which do not follow the branching of the phylogenetic tree
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vesth, T.C., Nybo, J.L., Theobald, S. et al. Investigation of inter- and intraspecies variation through genome sequencing of Aspergillus section Nigri. Nat Genet 50, 1688–1695 (2018). https://doi.org/10.1038/s41588-018-0246-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0246-1
This article is cited by
-
Two near-chromosomal-level genomes of globally-distributed Macroascomycete based on single-molecule fluorescence and Hi-C methods
Scientific Data (2024)
-
Co-production of sugars and aroma compounds from tobacco waste using biomass-degrading enzymes produced by Aspergillus brunneoviolaceus Ab-10
Archives of Microbiology (2024)
-
Fabrication of Biofunctionalized Protease-Based Chitosan/Collagen Composite Membranes and Efficient Biodegradation Using Recombinant Aspergillus Fumigatus
Journal of Polymers and the Environment (2023)
-
Whole-genome sequencing of biofilm-forming and chromium-resistant mangrove fungus Aspergillus niger BSC-1
World Journal of Microbiology and Biotechnology (2023)
-
Space exposure enhanced pectin-degrading enzymes expression and activity in Aspergillus costaricaensis
World Journal of Microbiology and Biotechnology (2023)