Abstract
We expanded GWAS discovery for type 2 diabetes (T2D) by combining data from 898,130 European-descent individuals (9% cases), after imputation to high-density reference panels. With these data, we (i) extend the inventory of T2D-risk variants (243 loci, 135 newly implicated in T2D predisposition, comprising 403 distinct association signals); (ii) enrich discovery of lower-frequency risk alleles (80 index variants with minor allele frequency <5%, 14 with estimated allelic odds ratio >2); (iii) substantially improve fine-mapping of causal variants (at 51 signals, one variant accounted for >80% posterior probability of association (PPA)); (iv) extend fine-mapping through integration of tissue-specific epigenomic information (islet regulatory annotations extend the number of variants with PPA >80% to 73); (v) highlight validated therapeutic targets (18 genes with associations attributable to coding variants); and (vi) demonstrate enhanced potential for clinical translation (genome-wide chip heritability explains 18% of T2D risk; individuals in the extremes of a T2D polygenic risk score differ more than ninefold in prevalence).
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Summary-level data are available at the DIAGRAM consortium website http://diagram-consortium.org/ and Accelerating Medicines Partnership T2D portal http://www.type2diabetesgenetics.org/.
References
Scott, R. A. et al. An expanded genome-wide association study of type 2 diabetes in Europeans. Diabetes 66, 2888–2902 (2017).
Zhao, W. et al. Identification of new susceptibility loci for type 2 diabetes and shared etiological pathways with coronary heart disease. Nat. Genet. 49, 1450–1457 (2017).
Mahajan, A. et al. Refining the accuracy of validated target identification through coding variant fine-mapping in type 2 diabetes. Nat. Genet. 50, 559–571 (2018).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283 (2016).
Jónsson, H. et al. Whole genome characterization of sequence diversity of 15,220 Icelanders. Sci. Data 4, 170115 (2017).
Flannick, J. & Florez, J. C. Type 2 diabetes: genetic data sharing to advance complex disease research. Nat. Rev. Genet. 17, 535–549 (2016).
Voight, B. F. et al. Twelve type 2 diabetes susceptibility loci identified through large-scale association analysis. Nat. Genet. 42, 579–589 (2010).
Morris, A. P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Kooner, J. S. et al. Genome-wide association study in individuals of South Asian ancestry identifies six new type 2 diabetes susceptibility loci. Nat. Genet. 43, 984–989 (2011).
Cho, Y. S. et al. Meta-analysis of genome-wide association studies identifies eight new loci for type 2 diabetes in east Asians. Nat. Genet. 44, 67–72 (2011).
Lotta, L. A. et al. Integrative genomic analysis implicates limited peripheral adipose storage capacity in the pathogenesis of human insulin resistance. Nat. Genet. 49, 17–26 (2017).
Magi, R., Lindgren, C. M. & Morris, A. P. Meta-analysis of sex-specific genome-wide association studies. Genet. Epidemiol. 34, 846–853 (2010).
Small, K. S. et al. Identification of an imprinted master trans regulator at the KLF14 locus related to multiple metabolic phenotypes. Nat. Genet. 43, 561–564 (2011).
Teslovich, T. M. et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature 466, 707–713 (2010).
Maller, J. B. et al. Bayesian refinement of association signals for 14 loci in 3 common diseases. Nat. Genet. 44, 1294–1301 (2012).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Fuchsberger, C. et al. The genetic architecture of type 2 diabetes. Nature 536, 41–47 (2016).
Gradwohl, G., Dierich, A., LeMeur, M. & Guillemot, F. Neurogenin3 is required for the development of the four endocrine cell lineages of the pancreas. Proc. Natl. Acad. Sci. USA 97, 1607–1611 (2000).
Rubio-Cabezas, O. et al. Permanent neonatal diabetes and enteric anendocrinosis associated with biallelic mutations in NEUROG3. Diabetes 60, 1349–1353 (2011).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Uchida, T. et al. Deletion of Cdkn1b ameliorates hyperglycemia by maintaining compensatory hyperinsulinemia in diabetic mice. Nat. Med. 11, 175–182 (2005).
Pasquali, L. et al. Pancreatic islet enhancer clusters enriched in type 2 diabetes risk-associated variants. Nat. Genet. 46, 136–143 (2014).
Varshney, A. et al. Genetic regulatory signatures underlying islet gene expression and type 2 diabetes. Proc. Natl. Acad. Sci. USA 114, 2301–2306 (2017).
Thurner, M. et al. Integration of human pancreatic islet genomic data refines regulatory mechanisms at Type 2 diabetes susceptibility loci. eLife 7, e31977 (2018).
Gaulton, K. J. et al. Genetic fine mapping and genomic annotation defines causal mechanisms at type 2 diabetes susceptibility loci. Nat. Genet. 47, 1415–1425 (2015).
Fogarty, M. P., Cannon, M. E., Vadlamudi, S., Gaulton, K. J. & Mohlke, K. L. Identification of a regulatory variant that binds FOXA1 and FOXA2 at the CDC123/CAMK1D type 2 diabetes GWAS locus. PLoS. Genet. 10, e1004633 (2014).
Dimas, A. S. et al. Impact of type 2 diabetes susceptibility variants on quantitative glycemic traits reveals mechanistic heterogeneity. Diabetes 63, 2158–2171 (2014).
Wood, A. R. et al. A genome-wide association study of IVGTT-based measures of first-phase insulin secretion refines the underlying physiology of type 2 diabetes variants. Diabetes 66, 2296–2309 (2017).
Pickrell, J. K. Joint analysis of functional genomic data and genome-wide association studies of 18 human traits. Am. J. Hum. Genet. 94, 559–573 (2014).
Plenge, R. M., Scolnick, E. M. & Altshuler, D. Validating therapeutic targets through human genetics. Nat. Rev. Drug. Discov. 12, 581–594 (2013).
van der Harst, P. & Verweij, N. Identification of 64 novel genetic loci provides an expanded view on the genetic architecture of coronary artery disease. Circ. Res. 122, 433–443 (2018).
van de Bunt, M. et al. Transcript expression data from human islets links regulatory signals from genome-wide association studies for type 2 diabetes and glycemic traits to their downstream effectors. PLoS. Genet. 11, e1005694 (2015).
Prokopenko, I. et al. A central role for GRB10 in regulation of islet function in man. PLoS. Genet. 10, e1004235 (2014).
Kaburagi, T., Kizuka, Y., Kitazume, S. & Taniguchi, N. The inhibitory role of α2,6-sialylation in adipogenesis. J. Biol. Chem. 292, 2278–2286 (2017).
Locke, A. E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Shungin, D. et al. New genetic loci link adipose and insulin biology to body fat distribution. Nature 518, 187–196 (2015).
Lizio, M. et al. Mapping mammalian cell-type-specific transcriptional regulatory networks using KD-CAGE and ChIP-seq data in the TC-YIK cell line. Front. Genet. 6, 331 (2015).
Scott, L. J. et al. The genetic regulatory signature of type 2 diabetes in human skeletal muscle. Nat. Commun. 7, 11764 (2016).
McCarthy, M. I., Rorsman, P. & Gloyn, A. L. TCF7L2 and diabetes: a tale of two tissues, and of two species. Cell. Metab. 17, 157–159 (2013).
Gaulton, K. J. et al. A map of open chromatin in human pancreatic islets. Nat. Genet. 42, 255–259 (2010).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Meigs, J. B., Cupples, L. A. & Wilson, P. W. Parental transmission of type 2 diabetes: the Framingham Offspring Study. Diabetes 49, 2201–2207 (2000).
Meigs, J. B. et al. Genotype score in addition to common risk factors for prediction of type 2 diabetes. N. Engl. J. Med. 359, 2208–2219 (2008).
Weedon, M. N. et al. Combining information from common type 2 diabetes risk polymorphisms improves disease prediction. PLoS. Med. 3, e374 (2006).
Euesden, J., Lewis, C. M. & O’Reilly, P. F. PRSice: Polygenic Risk Score software. Bioinformatics 31, 1466–1468 (2015).
Gatineau, M. et al. Adult obesity and type 2 diabetes (Public Health England, London, 2014). https://www.gov.uk/government/uploads/system/uploads/attachment_data/file/338934/Adult_obesity_and_type_2_diabetes_.pdf.
Zheng, J. et al. LD Hub: a centralized database and web interface to perform LD score regression that maximizes the potential of summary level GWAS data for SNP heritability and genetic correlation analysis. Bioinformatics 33, 272–279 (2017).
Kang, H. M. et al. Variance component model to account for sample structure in genome-wide association studies. Nat. Genet. 42, 348–354 (2010).
Loh, P. R. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Cook, J. P., Mahajan, A. & Morris, A. P. Guidance for the utility of linear models in meta-analysis of genetic association studies of binary phenotypes. Eur. J. Hum. Genet. 25, 240–245 (2017).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Ioannidis, J. P., Patsopoulos, N. A. & Evangelou, E. Heterogeneity in meta-analyses of genome-wide association investigations. PLoS One 2, e841 (2007).
Pulit, S. L., de With, S. A. & de Bakker, P. I. Resetting the bar: statistical significance in whole-genome sequencing-based association studies of global populations. Genet. Epidemiol. 41, 145–151 (2017).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Wakefield, J. A Bayesian measure of the probability of false discovery in genetic epidemiology studies. Am. J. Hum. Genet. 81, 208–227 (2007).
Denny, J. C. et al. PheWAS: demonstrating the feasibility of a phenome-wide scan to discover gene-disease associations. Bioinformatics 26, 1205–1210 (2010).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Stegle, O., Parts, L., Piipari, M., Winn, J. & Durbin, R. Using probabilistic estimation of expression residuals (PEER) to obtain increased power and interpretability of gene expression analyses. Nat. Protoc. 7, 500–507 (2012).
Ongen, H., Buil, A., Brown, A. A., Dermitzakis, E. T. & Delaneau, O. Fast and efficient QTL mapper for thousands of molecular phenotypes. Bioinformatics 32, 1479–1485 (2016).
Hormozdiari, F. et al. Colocalization of GWAS and eQTL signals detects target genes. Am. J. Hum. Genet. 99, 1245–1260 (2016).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Frazer, K. A. et al. A second generation human haplotype map of over 3.1 million SNPs. Nature 449, 851–861 (2007).
Acknowledgements
This work was supported primarily by the NIDDK as part of the Accelerating Medicines Partnership-T2D, funded by U01DK105535 (M.I.M.), U01DK062370 (M.B.), and U01DK078616 (J.M.) grants. Part of this work was conducted using the UK Biobank resource under application number 9161. A full list of acknowledgements appears in the Supplementary Note.
Author information
Authors and Affiliations
Contributions
Project coordination: A. Mahajan, A.P.M., M.B., and M.I.M. Writing: A. Mahajan, D.T., A.P.M., M.B., and M.I.M. Core analyses: A. Mahajan, D.T., M.T., J.M.T., A.J.P., A.P.M., M.B., and M.I.M. DIAMANTE analysis group: A. Mahajan, J.E.B., D.W.B., J.C.C., Y.J.K., M.C.Y.N., L.E.P., X.S., W.Z., A.P.M., M.B., and M.I.M. Statistical analysis in individual studies: A. Mahajan, D.T., N.R.R., N.W.R., V.S., R.A.S., N.G., J.P.C., E.M.S., M.W., C. Sarnowski, J.N., S.T., C. Lecoeur, M.H.P., B.P.P., X.G., L.F.B., J.B.-J., M.C., K.L., C.-T.L., A.E.L., J’a.L., C. Schurmann, L.Y., G.T., and A.P.M. Genotyping and phenotyping: A. Mahajan, R.A.S., R.M., C.G., S.T., K.-U.E., K.F., S.L.R.K., F.K., I.N., C.M.B., C. Schurmann, E.P.B., I.B., C.C., G.D., I.F., V.G., M.I., M.E.J., S.L., A.L., V.L., V.M., A.D.M., G.N., N.S., A.S., D.R.W., S.S., E.P.B., S.H., C.H., J. Kriebel, T.M., A.P., B.T., A.D., A.K., G.R.A., C. Langenberg, N.J.W., A.P.M., M.B., and M.I.M. Islet annotations: M.T., J.M.T., A.J.B., V.N., A.L.G., and M.I.M. Individual study design and principal investigators: E.P.B., J.C.F., O.H.F., T.M.F., A.T.H., M.A.I., T.J., J. Kuusisto, C.M.L., K.L.M., J.S.P., K. Strauch, K.D.T., U.T., J.T., J.D., P.A.P., E.Z., R.J.F.L., P.F., E.I., L.L., L.G., M.L., F.S.C., J.W.J., C.N.A.P., H.G., A. Metspalu, A.D., A.K., G.R.A., J.B.M., J.I.R., J.M., O.P., T.H., C. Langenberg, N.J.W., K. Stefansson, A.P.M., M.B., and M.I.M.
Corresponding authors
Ethics declarations
Competing interests
J.C.F. has received consulting honoraria from Merck and from Boehringer-Ingelheim. O.H.F. works at ErasmusAGE, a center for aging research across the course of life, funded by Nestlé Nutrition (Nestec Ltd.), Metagenics Inc., and AXA. E.I. is a scientific advisor for Precision Wellness and Olink Proteomics for work unrelated to the present project. A.D. has received consultancy fees and research support from Metagenics Inc. (outside the scope of the present work). T.M.F. has consulted for Boeringer Ingelheim and Sanofi-Aventis on the genetics of diabetes and has an MRC CASE studentship with GSK. G.R.A. is a consultant for 23andMe, Regeneron, Merck, and Helix. R.A.S. is an employee of and shareholder in GlaxoSmithKline. N.S. is working with Boehringer-Ingelheim on a genetics project but has received no remuneration. M.I.M. has served on advisory panels for NovoNordisk and Pfizer, and has received honoraria from NovoNordisk, Pfizer, Sanofi-Aventis, and Eli Lilly. The companies named above had no role in the design or conduct of this study; collection, management, analysis, and interpretation of the data, or in the preparation, review, or approval of the manuscript. Authors affiliated with deCODE (V.S., G.T., U.T. and K.S.) are employed by deCODE Genetics/Amgen, Inc.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Sex-differentiated analyses.
(a) Manhattan plot (top panel) of genome-wide association results for T2D (without BMI adjustment) from female-specific meta-analysis of up to 30,053 cases and 434,336 controls. The association p-value (on -log10 scale) for each SNP (y-axis) is plotted against the genomic position (NCBI Build 37; x-axis). Association signals that reached genome-wide significance (p < 5×10−8) in sex-combined analysis are shown in purple or yellow, if novel. (b) Manhattan plot (bottom panel) of genome-wide association results for T2D without BMI adjustment from male-specific meta-analysis of up to 41,846 cases and 383,767 controls. (c) Z-score for each of the 403 distinct signals from male-specific analysis (y-axis) is plotted against the z-score from the female-specific analysis (y-axis). Colour of each point varies with –log10 gender heterogeneity p-value and diameter of the circle is proportional to sex-combined -log10 p-value.
Supplementary Figure 2 Distributions of the allele frequency, imputation score, and posterior probability of association.
Distribution of the risk allele frequencies for all variants having >1% posterior probability of association in genetic credible set (x-axis) plotted against average imputation quality (y-axis). Diameter varies with the posterior probability of association assigned to each variant.
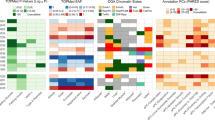
Supplementary Figure 3 Islet annotation overlap of the variant with the highest probability in genetic credible sets.
Number of variants with posterior probability of association >1% (x-axis) plotted against the highest posterior probability of association (y-axis) assigned to a variant in the credible set. Points are colour coded according to (a) islet epigenome states and (b) overlap with transcription factor binding sites.
Supplementary Figure 4 Enrichment of cross-tissue epigenetic states in T2D GWAS data.
fGWAS log2 fold enrichment (based on joint model for each tissue) including 95% confidence intervals (x-axis) of all chromatin states (y-axis) genome-wide. Analyses are based on the Varshney et al.1 data which combined standard epigenomic annotations for the four principal tissues of interest. These analyses performed separately for each tissue show some enrichment for enhancers and/or promoters in all tissues with strongest and most consistent enrichment observed in islets. The universally enriched “transcript” category refers to coding sequence which is by definition represented by the same sequence in each “tissue-specific” analysis. 1Varshney, A. et al. Genetic regulatory signatures underlying islet gene expression and type 2 diabetes. Proc Natl Acad Sci U S A 114, 2301-2306 (2017).
Supplementary Figure 5 Enrichment of islet epigenetic states in T2D GWAS data.
fGWAS log2 fold enrichment including 95% confidence intervals (x-axis) of all chromatin states (y-axis) genome-wide.
Supplementary Figure 6 Epigenome landscape of the ST6GAL1 locus.
For variants included in 99% credible set (PPA>1%) of each distinct signal at ST6GAL1 locus, following information is shown: genomic position of each variant (colour coded for each distinct signal; variant with highest PPA in bold); whole genome bisulphite methylation data (black), 4 human islet ATAC-seq tracks (green, middle), islet chromatin states (from Thurner et al.1, Pasquali et al.2, and Varshney et al.3); and adipose, liver and skeletal muscle chromatin states from Varshney et al.3. 1 Thurner, M. et al. Integration of human pancreatic islet genomic data refines regulatory mechanisms at Type 2 Diabetes susceptibility loci. Elife 7(2018). 2 Pasquali, L. et al. Pancreatic islet enhancer clusters enriched in type 2 diabetes risk-associated variants. Nat Genet 46, 136-143 (2014). 3 Varshney, A. et al. Genetic regulatory signatures underlying islet gene expression and type 2 diabetes. Proc Natl Acad Sci U S A 114, 2301-2306 (2017).
Supplementary Figure 7 Epigenome landscape of the ANK1 locus.
For variants included in 99% credible set (PPA>1%) of each distinct signal at ANK1 locus, following information is shown: genomic position of each variant (colour coded for each distinct signal; variant with highest PPA in bold); whole genome bisulphite methylation data (black), 4 human islet ATAC-seq tracks (green, middle), islet chromatin states (from Thurner et al.1, Pasquali et al.2, and Varshney et al.3); and adipose, liver and skeletal muscle chromatin states from Varshney et al.3. 1 Thurner, M. et al. Integration of human pancreatic islet genomic data refines regulatory mechanisms at Type 2 Diabetes susceptibility loci. Elife 7(2018). 2 Pasquali, L. et al. Pancreatic islet enhancer clusters enriched in type 2 diabetes risk-associated variants. Nat Genet 46, 136-143 (2014).3 Varshney, A. et al. Genetic regulatory signatures underlying islet gene expression and type 2 diabetes. Proc Natl Acad Sci U S A 114, 2301-2306 (2017).
Supplementary Figure 8 Epigenome landscape of the TCF7L2 locus.
For variants included in 99% credible set (PPA>1%) of each distinct signal at TCF7L2 locus, following information is shown: genomic position of each variant (colour coded for each distinct signal; variant with highest PPA in bold); whole genome bisulphite methylation data (black), 4 human islet ATAC-seq tracks (green, middle), islet chromatin states (from Thurner et al.1, Pasquali et al.2, and Varshney et al.3); and adipose, liver and skeletal muscle chromatin states from Varshney et al.3. 1 Thurner, M. et al. Integration of human pancreatic islet genomic data refines regulatory mechanisms at Type 2 Diabetes susceptibility loci. Elife 7(2018). 2 Pasquali, L. et al. Pancreatic islet enhancer clusters enriched in type 2 diabetes risk-associated variants. Nat Genet 46, 136-143 (2014). 3 Varshney, A. et al. Genetic regulatory signatures underlying islet gene expression and type 2 diabetes. Proc Natl Acad Sci U S A 114, 2301-2306 (2017).
Supplementary Figure 9 Heritability estimates.
Chip heritability estimates for T2D (on the liability scale) at different empirical estimates of population- and sample-level T2D prevalence.
Supplementary Figure 10 Polygenic risk scores.
Genome-wide polygenic risk score (PRS) identifies individuals with significantly increased risk of T2D. a) PRS in UK Biobank individuals is normally distributed with a shift towards right, observed for T2D cases. PRS is plotted on the x-axis, with values scaled to a mean of 0 and standard deviation of 1. b) Individuals were binned into 40 groups based on PRS, with each grouping representing 2.5% of population. c) BMI distribution in T2D cases, within each PRS bin.
Supplementary Figure 11 Genetic correlations between T2D and biomedically relevant traits, estimated by LD-score regression implemented in LDHub.
Genetic correlations (z-score) between T2D (y-axis) and range of metabolic and anthropometric traits (x-axis) as estimated using LD Score regression. The genetic correlation estimates are colour coded according to phenotypic area. Allelic direction of effect is aligned to increased T2D risk. Size of the circle denotes the significance level for the correlation.
Supplementary Figure 12 Effect of BMI adjustment on genetic correlation estimates between various traits and T2D.
Genetic correlations (z-score) between range of metabolic and anthropometric traits and T2D without BMI adjustment (x-axis) and T2D with BMI adjustment (y-axis) as estimated using LD Score regression. The genetic correlation estimates are colour coded according to phenotypic area. Allelic direction of effect is aligned to increased T2D risk. Size of the circle denotes the significance level for the correlation.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12 and Supplementary Note
Rights and permissions
About this article
Cite this article
Mahajan, A., Taliun, D., Thurner, M. et al. Fine-mapping type 2 diabetes loci to single-variant resolution using high-density imputation and islet-specific epigenome maps. Nat Genet 50, 1505–1513 (2018). https://doi.org/10.1038/s41588-018-0241-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0241-6
This article is cited by
-
Genome-wide association study and trans-ethnic meta-analysis identify novel susceptibility loci for type 2 diabetes mellitus
BMC Medical Genomics (2024)
-
DNA methylation may partly explain psychotropic drug-induced metabolic side effects: results from a prospective 1-month observational study
Clinical Epigenetics (2024)
-
The role of mitochondrial DNA copy number in cardiometabolic disease: a bidirectional two-sample mendelian randomization study
Cardiovascular Diabetology (2024)
-
A genome-wide association meta-analysis implicates Hedgehog and Notch signaling in Dupuytren’s disease
Nature Communications (2024)
-
Comprehensive genetic study of the insulin resistance marker TG:HDL-C in the UK Biobank
Nature Genetics (2024)