Abstract
We report full-length draft de novo genome assemblies for 16 widely used inbred mouse strains and find extensive strain-specific haplotype variation. We identify and characterize 2,567 regions on the current mouse reference genome exhibiting the greatest sequence diversity. These regions are enriched for genes involved in pathogen defence and immunity and exhibit enrichment of transposable elements and signatures of recent retrotransposition events. Combinations of alleles and genes unique to an individual strain are commonly observed at these loci, reflecting distinct strain phenotypes. We used these genomes to improve the mouse reference genome, resulting in the completion of 10 new gene structures. Also, 62 new coding loci were added to the reference genome annotation. These genomes identified a large, previously unannotated, gene (Efcab3-like) encoding 5,874 amino acids. Mutant Efcab3-like mice display anomalies in multiple brain regions, suggesting a possible role for this gene in the regulation of brain development.
Similar content being viewed by others
Main
Inbred laboratory strains of mice are broadly organized into two groups, classical and wild-derived strains1, that can be used to model the variation observed in human populations2,3. Inbred laboratory strains of wild-derived origin represent a rich source of phenotypic responses and genetic diversity not present in classical strains of mice4,5,6. Wild-derived strains have been crossed with classical strains to create powerful resources such as the Collaborative Cross (CC) and Diversity Outbred Cross (DO) in which genetic traits have been mapped7,8,9,10.
The generation and assembly of a reference genome for C57BL/6J accelerated the discovery of the genetic landscape underlying phenotypic variation11. Using this reference, genome-wide variation catalogs (single nucleotide polymorphisms (SNPs), short indels, and structural variation) for 36 laboratory mouse strains were generated12,13. However, reliance on mapping next-generation sequencing reads to C57BL/6J has meant that the true extent of strain-specific variation is unknown. At some loci, the genetic difference between the reference and sequenced strain genomes is comparable to that between humans and chimpanzees, making it hard to distinguish whether a read is mismapped or highly divergent. De novo genome assembly methods address this issue by allowing unbiased assessments of the differences between genomes.
We have completed the first draft de novo assemblies and strain-specific gene annotation for 12 classical inbred laboratory mouse strains (129S1/SvImJ, A/J, AKR/J, BALB/cJ, C3H/HeJ, C57BL/6NJ, CBA/J, DBA/2J, FVB/NJ, LP/J, NZO/HlLtJ, and NOD/ShiLtJ) and 4 wild-derived strains representing the backgrounds Mus musculus castaneus (CAST/EiJ), M. m. musculus (PWK/PhJ), M. m. domesticus (WSB/EiJ), and M. spretus (SPRET/EiJ). This collection comprises a large and diverse array of laboratory strains, including those closely related to commonly used mouse cell lines (BALB/3T3 and L929, derived from BALB/c and C3H related strains, respectively), embryonic stem cell-derived gene knockouts (historically 129 related strains)14, mouse models of human disease (such as NOD-related nude mice)15, gene knockout background strains (C57BL/6NJ)16, the founders of commonly used recombinant inbred lines such as the AKXD, BXA, BXD, CXB, and CC17, and outbred mapping populations such as the DO and the heterogeneous stock18.
Results
Sequence assemblies and genome annotation
Chromosome-scale assemblies were produced for 16 laboratory mouse strains using a mixture of Illumina paired-end (40–70×), mate-pairs (3, 6, 10 kilobases (kb)), fosmid, and BAC end sequences (Supplementary Table 1), and Dovetail Genomics Chicago libraries19. Pseudochromosomes were produced in parallel utilizing cross-species synteny alignments resulting in genome assemblies of between 2.254 (WSB/EiJ) and 2.328 gigabases (Gb) (AKR/J) excluding unknown gap bases. Approximately 0.5–2% of total genome length per strain was unplaced and is composed of unknown gap bases (18–49%) and repeat sequences (61–79%) (Supplementary Table 2), with between 89 and 410 predicted genes per strain (Supplementary Table 3). Mitochondrial genome (mtDNA) assemblies for 14 strains supported previously published sequences20, although a small number of high-quality novel sequence variants in AKR/J, BALB/cJ, C3H/HeJ, and LP/J conflicted with GenBank entries (Supplementary Table 4). Novel mtDNA haplotypes were identified in PWK/PhJ and NZO/HlLtJ. Notably, NZO/HlLtJ contained 55 SNPs (33 shared with the wild-derived strains) and appears distinct compared to the other classical inbred strains (Supplementary Fig. 1). Previous variation catalogs have indicated high concordance (>97% shared SNPs) between NZO/HlLtJ and another inbred laboratory strain NZB/BlNJ21.
We assessed the base accuracy of the strain chromosomes relative to two versions of the C57BL/6J reference genome (MGSCv311 and GRCm382) by first realigning all of the paired-end sequencing reads from each strain back to their respective genome assemblies, then using these alignments to identify SNPs and indels. The combined SNP and indel error rate was 0.09–0.1 errors per kb, compared to 0.334 for MGSCv3 and 0.02 for GRCm38 (Supplementary Table 5). Next, we used a set of 612 polymerase chain reaction (PCR) primer pairs previously used to validate structural variant calls in eight strains22. The assemblies had 4.7–6.7% primer pairs showing incorrect alignments compared to 10% for MGSCv3 (Supplementary Table 6). Finally, alignment of PacBio long-read complementary DNA sequences from liver and spleen of C57BL/6J, CAST/EiJ, PWK/PhJ, and SPRET/EiJ showed that the GRCm38 reference genome had the highest proportion of correctly aligned cDNA reads (99% and 98%, respectively) and the strains and MGSCv3 were 1–2% lower (Supplementary Table 7). The representation of known mouse repeat families in the assemblies shows that the short repeat (<200 base pairs (bp)) content was comparable to GRCm38 (Supplementary Fig. 2a,b). The total number of long repeats (>200 bp) is consistent across all strains; however, the total sequence lengths are consistently shorter than GRCm38 (Supplementary Fig. 2c).
Strain-specific consensus gene sets were produced using the GENCODE C57BL/6J annotation and strain-specific RNA sequencing (RNA-Seq) from multiple tissues23 (Supplementary Table 8 and Supplementary Fig. 3). The consensus gene sets contain over 20,000 protein coding genes and over 18,000 non-coding genes (Fig. 1a and Supplementary Table 1). For the classical laboratory strains, 90.2% of coding transcripts (88.0% in wild-derived strains) and 91.2% of non-coding transcripts (91.4% in wild-derived strains) present in the GRCm38 reference gene set were comparatively annotated. Gene predictions from strain-specific RNA-Seq (Comparative Augustus24) added an average of 1,400 new isoforms to wild-derived and 1,207 new isoforms to classical strain gene annotation sets. Gene prediction based on PacBio cDNA sequencing introduced an average of 1,865 further new isoforms to CAST/EiJ, PWK/PhJ, and SPRET/EiJ. Putative novel loci are defined as spliced genes that were predicted from strain-specific RNA-Seq and did not overlap any genes projected from the reference genome. On average, 37 genes were putative novel loci (Supplementary Data 1) in wild-derived strains and 22 in classical strains. Most often these appear to result from gene duplication events. Additionally, an automated pseudogene annotation workflow, Pseudopipe25, alongside manually curated pseudogenes lifted over from the GRCm38 reference genome, identified an average of 11,000 (3,317 conserved between all strains) pseudogenes per strain (Supplementary Fig. 4) that appear to have arisen either through retrotransposition (~80%) or gene duplication events (~20%).
a, Summary of the strain-specific gene sets showing the number of genes broken down by GENCODE biotype. b, Heterozygous SNP (hSNP) density for a 50 Mb interval on chromosome 11 in 200 kb windows for 17 inbred mouse strains based on sequencing read alignments to the C57BL/6J (GRCm38) reference genome (top). Labels indicate genes overlapping the densest regions. SNPs visualized in CAST/EiJ and WSB/EiJ for 71.006–71.170 Mb on GRCm38 (bottom), including Derl2 and Mis12 (upper panel) and Nlrp1b (lower panel). Grey indicates the strain base agrees with the reference, other colors indicate SNP differences and height corresponds to sequencing depth. c, Total amount of sequence and protein-coding genes in regions enriched for hSNPs (relative to the GRCm38 reference genome) per strain. d, Top PantherDB categories of coding genes in regions enriched for hSNPs based on protein class (left). Intersection of genes in the defence and immunity category for the wild-derived and classical inbred strains (right). e, Box plot of sequence divergence (%)for LTRs, LINEs, and SINEs within and outside of hSNP regions. Sequence divergence is relative to a consensus sequence for the transposable element type (n = number of repeats in GRCm38, *** indicated P < 0.001 using Welch’s two-sample t-test. Box plots show 25th and 75th percentiles, and the median value.
Regions of the mouse genome with extreme allelic variation
Inbred laboratory mouse strains are characterized by at least 20 generations of inbreeding and are genetically homozygous at almost all loci1. Despite this, previous SNP variation catalogs have identified high-quality heterozygous SNPs (hSNPs) when reads were aligned to the C57BL/6J reference genome12. The presence of higher densities of hSNPs may indicate copy number changes, or novel genes that are not present in the reference assembly, forced to partially map to a single locus in the reference12,21. Thus, their identification is a powerful tool for finding errors in genome assemblies. We identified between 116,439 (C57BL/6NJ) and 1,895,741 (SPRET/EiJ) high-quality hSNPs from the MGP variation catalog v521 (Supplementary Table 9). Focusing our analysis on the top 5% most hSNP-dense regions (windows ≥ 71 hSNPs per 10 kb sliding window) identified the majority of known polymorphic regions among the strains (Supplementary Fig. 5) and accounted for ~49% of all hSNPs (Supplementary Table 9 and Supplementary Fig. 6a). After applying this cut-off to all strain-specific hSNP regions and merging overlapping or adjacent windows, between 117 (C57BL/6NJ) and 2,567 (SPRET/EiJ) hSNP regions remained per strain (Supplementary Table 9), with an average size of 18–20 kb (Supplementary Fig. 6b). Many hSNP clusters overlap immunity (for example, MHC, NOD-like receptors, and AIM-like receptors), sensory (for example, olfactory and taste receptors), reproductive (for example, pregnancy-specific glycoproteins and sperm-associated E-rich proteins), and neuronal- and behavior-related genes (for example, itch receptors26 and γ-protocadherins27) (Fig. 1b and Supplementary Fig. 5). All of the wild-derived strain hSNP regions contained gene and coding sequence (CDS) base-pair counts larger than any classical inbred strain (≥503 and ≥0.36 megabases (Mb), respectively; Supplementary Table 9). The regions identified in C57BL/6J and C57BL/6NJ (117 and 141, respectively; 145 combined) intersect known GRCm38 assembly issues including gaps, unplaced scaffolds, or centromeric regions (107/145, 73.8%). The remaining candidate regions include large protein families (15/145, 10.3%) and repeat elements (17/145, 11.7%) (Supplementary Data 2).
We examined protein classes present in the hSNP regions by identifying 1,109 PantherDB matches, assigned to 26 protein classes from a combined set of all genes in hSNP dense regions (Supplementary Data 3). Defence and immunity was the largest represented protein class (155 genes, Supplementary Data 4), accounting for 13.98% of all protein class hits (Supplementary Table 10). This was a five-fold enrichment compared to an estimated genome-wide rate (Fig. 1d). Notably, 89 immune-related genes were identified in classical strains, 84 of which were shared with at least one of the wild-derived strains (Fig. 1d). SPRET/EiJ contributed the largest number of strain-specific gene hits (22 genes).
Many paralogous gene families were represented among the hSNP regions (Supplementary Data 3), including genes with functional human orthologs. Several prominent examples include apolipoprotein L alleles, variants of which may confer resistance to Trypanosoma brucei, the primary cause of human sleeping sickness28,29; IFI16 (interferon gamma inducible protein 16, a member of AIM2-like receptors), a DNA sensor required for death of lymphoid CD4 T cells abortively infected with human immunovirus (HIV)30; NAIP (NLR family apoptosis inhibitory protein) in which functional copy number variation is linked to increased cell death upon Legionella pneumophila infection31; and secretoglobins (Scgb members), which may be involved in tumor formation and invasion in both human and mouse32,33. Large gene families in which little functional information is known were also identified. A cluster of approximately 50 genes, which includes hippocalcin-like 1 (Hpcal1) and its homologs, were identified (chromosome 12: 18–25 Mb). Hpcal1 belongs to the neuronal calcium sensors expressed primarily in retinal photoreceptors, neurons, and neuroendocrine cells34. This region is enriched for hSNPs in all strains except C57BL/6J and C57BL/6NJ. Interestingly, within this region, Cpsf3 (21.29 Mb) is located on an island of high conservation in all strains and a homozygous C57BL/6NJ knockout produces subviable offspring35. Additional examples include another region on chromosome 12 (87–88 Mb) containing approximately 20 eukaryotic translation initiation factor 1A (eIF1a) homologs and on chromosome 14 (41–45 Mb) containing approximately 100 Dlg1-like genes. Genes within all hSNP candidate regions have been identified and annotated (Supplementary Fig. 5).
We examined retrotransposon content in hSNP dense regions on GRCm38 compared to an estimated null distribution (one million simulations) and found a significant enrichment of both LTRs (empirical P < 1 × 10–7) and long interspersed nuclear elements (LINEs) (empirical P < 1 × 10−7) (Supplementary Tables 11 and 12). Gene retrotransposition has long been implicated in the creation of gene family diversity36, novel alleles conferring positively selected adaptations37. Once transposed, transposable elements accumulate mutations over time as the sequence diverges38,39. For LTRs, LINEs and short interspersed nuclear elements (SINEs), the mean percentage sequence divergence was significantly lower (P < 1 × 10−22) within hSNP regions compared to the rest of the genome (Fig. 1e). The largest difference in mean sequence divergence was between LTRs within and outside of hSNP dense regions. Examining only repeat elements with less than 1% divergence, we found these regions are significantly enriched for LTRs (empirical P < 1 × 10−7) and LINEs (empirical P = 0.047).
De novo assembly of complex gene families
Our data elucidated copy number variation previously unknown in mouse strain genomes and uncovered gene expansions, contractions, and novel alleles (<80% sequence identity). For example, 23 distinct clusters of olfactory receptors were identified, indicating substantial variation among inbred strains. In mouse, phenotypic differences, particularly in diet and behavior, have been linked to distinct olfactory receptor repertoires40,41. To this end, we have characterized the CAST/EiJ olfactory receptor repertoire using our de novo assembly and identified 1,249 candidate olfactory receptor genes (Supplementary Data 5). Relative to the reference strain (C57BL/6J), CAST/EiJ has lost 20 olfactory receptors and gained 37 gene family members: 12 novel and 25 supported by published predictions based on messenger RNA (mRNA) derived from CAST/EiJ whole olfactory mucosa (Fig. 2a and Supplementary Table 13)42.
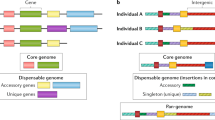
a, Olfactory receptor genes on chromosome 11 of CAST/EiJ. Gene gain/ loss and similarity are relative to C57BL/6J. Novel members are named after their most similar homologs. b, Gene order across Raet1/H60 locus in the Collaborative Cross parental strains (A/J, NOD/ShiLtJ and 129S1/SvImJ share the same haplotype at this locus, represented by NOD/ShiLtJ). Strain name in black/red indicates Aspergillus fumigatus resistant/susceptible. Dashed box indicates unconfirmed gene order. c, Novel protein-coding alleles of the Nlrp1 gene family in the wild-derived strains and two classical inbred strains. Colors represent the phylogenetic relationships (top, amino acid neighbor joining tree of NBD domain) and the relative gene order across strains (bottom). d, A regional dot plot of the Nlrp1 locus in PWK/PhJ compared to the C57BL/6J GRCm38 reference (color-coded same as panel c). Grey blocks indicate repeats and transposable elements.
We discovered novel gene members at several important immune loci regulating innate and adaptive responses to infection. For example, chromosome 10 (22.1–22.4 Mb) on C57BL/6J contains Raet1 alleles and minor histocompatibility antigen members of H60. Raet1 and H60 are important ligands for NKG2D, an activating receptor of natural killer cells43. NKG2D ligands are expressed on the surface of infected44 and metastatic cells45 and may participate in allograft autoimmune responses46. From the de novo assembly, six different Raet1/H60 haplotypes were identified among the eight CC founder strains; three of the haplotypes identified are shared among the classical inbred CC founders (A/J, 129S1/SvImJ and NOD/ShiLtJ have the same haplotype) and three different Raet1/H60 haplotypes were identified in each of the wild-derived inbred strains (CAST/EiJ, PWK/PhJ and WSB/EiJ) (Fig. 2b and Supplementary Figs. 7 and 8). The CAST/EiJ haplotype encodes only a single Raet1 family member (Raet1e) and no H60 alleles, while the classical NOD/ShiLtJ haplotype has four H60 and three Raet1 alleles. The Aspergillus-resistant locus 4 (Asprl4), one of several quantitative trait loci (QTLs) that mediate resistance against Aspergillus fumigatus infection, overlaps this locus and comprises of a 1 Mb (~10% of QTL) interval that, compared to other classical strains, contains a haplotype unique to NZO/HlLtJ (Supplementary Fig. 7). Strain-specific haplotype associations with Asprl4 and survival have been reported for CAST/EiJ and NZO/HlLtJ, both of which exhibit resistance to A. fumigatus infection47 and they are also the only strains to have lost H60 alleles at this locus.
We examined three immunity-related loci on chromosome 11, IRG (GRCm38: 48.85–49.10 Mb), Nlrp1 (71.05–71.30 Mb), and Slfn (82.9–83.3 Mb) because of their polymorphic complexity and importance for mouse survival48,49,50. The Nlrp1 locus (NOD-like receptors, pyrin domain-containing) encodes inflammasome components that sense endogenous microbial products and metabolic stresses, thereby stimulating innate immune responses51. In the house mouse, Nlrp1 alleles are involved in sensing Bacillus anthracis lethal toxin, leading to inflammasome activation and pyroptosis of macrophages52,53. We discovered seven distinct Nlrp1 family members by comparing six strains (CAST/EiJ, PWK/PhJ, WSB/EiJ, SPRET/EiJ, NOD/ShiLtJ, and C57BL/6J). Each strain has a unique haplotype of Nlrp1 members, highlighting the extensive sequence diversity at this locus across inbred mouse strains (Fig. 2c). Each of the three M. m. domesticus strains (C67BL/6J, NOD/ShiLtJ, and WSB/EiJ) carries a different combination of Nlrp1 family members; Nlrp1d–1f are novel strain-specific alleles that were previously unknown. Diversity between different Nlrp1 alleles is higher than sequence divergence between mouse and rat alleles. For example, C57BL/6J contains Nlrp1c, which is not present in the other two strains, while Nlrp1b2 is present in both NOD/ShiLtJ and WSB/EiJ but not C57BL/6J. In PWK/PhJ (M. m. musculus), the Nlrp1 locus is almost double in size relative to the GRCm38 reference genome and contains novel Nlrp1 homologs (Fig. 2c), whereas in M. spretus (also wild-derived) this locus is much shorter than in any other mouse strain. Approximately 90% of intergenic regions in the PWK/PhJ assembly of the Nlrp1 locus is composed of transposable elements (Fig. 2d).
The wild-derived PWK/PhJ (M. m. musculus) and CAST/EiJ (M. m. castaneus) strains share highly similar haplotypes; however, PWK/PhJ macrophages are resistant to pyroptotic cell death induced by anthrax lethal toxin while CAST/EiJ macrophages are not54. It has been suggested that Nlrp1c may be the causal family member mediating resistance; Nlrp1c can be amplified from cDNA from PWK/PhJ macrophages but not CAST/EiJ54. In the de novo assemblies, both mouse strains share the same promoter region for Nlrp1c; however, when transcribed, the cDNA of Nlrp1c_CAST could not be amplified with previously designed primers54 due to SNPs at the primer binding site (5′...CACT-3′ → 5′...TACC-3′). The primer binding site in PWK/PhJ is the same as that in C57BL/6J, however Nlrp1c is a predicted pseudogene. We found an 18 amino acid mismatch in the nucleotide-binding domain (NBD) between Nlrp1b_CAST and Nlrp1b_PWK. These divergent profiles suggest that Nlrp1c is not the sole mediator of anthrax lethal toxin resistance in the mouse but several other members may be involved. Newly annotated members Nlrp1b2 and Nlrp1d appear functionally intact in CAST/EiJ but were both predicted as pseudogenes in PWK/PhJ due to the presence of stop codons or frameshift mutations. In C57BL/6J, three splicing isoforms of Nlrp1b (SV1, SV2, and SV3) were reported54. A dot-plot between PWK/PhJ and the C57BL/6J reference illustrates the disruption of co-linearity at the PWK/PhJ Nlrp1b2 and Nlrp1d alleles (Fig. 2d). All of the wild-derived strains we sequenced contain full-length Nlrp1d and exhibit a similar disruption of co-linearity at these alleles relative to C57BL/6J (Supplementary Data 6). The SV1 isoform in C57BL/6J is derived from truncated ancestral paralogs of Nlrp1b and Nlrp1d, indicating that Nlrp1d was lost in the C57BL/6J lineage. The genome structure of the Nlrp1 locus in PWK/PhJ, CAST/EiJ, WSB/EiJ, and NOD/ShiLtJ was confirmed using Fiber-FISH (Supplementary Fig. 9).
The assemblies also showed extensive diversity at each of the other loci examined: immunity-related GTPases (IRGs) and Schlafen family (Slfn). IRG proteins belong to a subfamily of interferon-inducible GTPases present in most vertebrates55. In mouse, IRG protein family members contribute to the adaptive immune system by conferring resistance against intracellular pathogens such as Chlamydia trachomatis, Trypanosoma cruzi, and Toxoplasma gondii56. Our de novo assembly is concordant with previously published data for CAST/EiJ48. For the first time, it shows the order, orientation, and structure of three highly divergent haplotypes present in WSB/EiJ, PWK/PhJ, and SPRET/EiJ, including novel annotation of rearranged promoters, inserted processed pseudogenes, and a high frequency of LINE repeats (Supplementary Data 6).
The Schlafen (chromosome 11: 82.9–83.3 Mb) family of genes are reportedly involved in immune responses, cell differentiation, proliferation and growth, cancer invasion, and chemotherapy resistance. In humans, SLFN11 was reported to inhibit HIV protein synthesis by a codon-usage-based mechanism57 and in non-human primates positive selection on the gene Slfn11 has been reported58. In mouse, embryonic death may occur between strains carrying incompatible Slfn haplotypes59. Assembly of Slfn for the three CC founder strains of wild-derived origin (CAST/EiJ, PWK/PhJ, and WSB/EiJ) showed, for the first time, extensive variation at this locus. Members of group 4 Slfn genes50, Slfn8, Slfn9, and Slfn10, show significant sequence diversity among these strains. For example, Sfln8 is a predicted pseudogene in PWK/PhJ but is protein coding in the other strains; the CAST/EiJ allele contains 78 amino acid mismatches compared to the C57BL/6J reference (Supplementary Fig. 10). Both CAST/EiJ and PWK/PhJ contain functional copies of Sfln10, which is a predicted pseudogene in C57BL/6J and WSB/EiJ. A novel start codon upstream of Slfn4, which causes a 25 amino acid N-terminal extension, was identified in PWK/PhJ and WSB/EiJ. Another member present in the reference, Slfn14, is conserved in PWK/PhJ and CAST/EiJ but is a pseudogene in WSB/EiJ (Supplementary Fig. 10).
Reference genome updates informed by the strain assemblies
There are currently 11 genes in the GRCm38 reference assembly (C57BL/6J) that are incomplete due to a gap in the sequence. First, these loci were compared to the respective regions in the C57BL/6NJ assembly and used to identify contigs from public assemblies of the reference strain previously omitted due to insufficient overlap. Second, C57BL/6J reads aligned to the regions of interest in the C57BL/6NJ assembly were extracted for targeted assembly, leading to the generation of contigs covering sequences currently missing from the reference. Both approaches resulted in the completion of ten new gene structures (for example, Supplementary Fig. 11 and Supplementary Data 7) and the near-complete inclusion of the Sts gene that was previously missing.
Improvements to the reference genome, coupled with pan-strain gene predictions, were used to provide updates to the existing reference genome annotation, maintained by the GENCODE consortium60. We examined the strain-specific RNA-Seq (Comparative Augustus) gene predictions containing 75% novel introns compared to the existing reference annotation (Table 1) (GENCODE M8, chromosomes 1–12). Of the 785 predictions investigated, 62 led to the annotation of new loci, including 19 protein-coding genes and 6 pseudogenes (Supplementary Table 14 and Supplementary Data 8). In most cases where a new locus was predicted on the reference genome, we identified pre-existing, but often incomplete, annotation. For example, the Nmur1 gene was extended at its 5′ end and made complete on the basis of evidence supporting a prediction that spliced to an upstream exon containing the previously missing start codon. The Mroh3 gene, which was originally annotated as an unprocessed pseudogene, was updated to a protein-coding gene due to the identification of a novel intron that permitted extension of the CDS to full length. The previously annotated pseudogene model has been retained as a nonsense-mediated decay (NMD) transcript of the protein- coding locus. At the novel bicistronic locus, Chml_Opn3, the original annotation was a single exon gene, Chml, that was extended and found to share its first exon with the Opn3 gene.
We discovered a novel 188-exon gene on chromosome 11 that significantly extends the existing gene Efcab3 spanning between Itgb3 and Mettl2 (Fig. 3a). This Efcab3-like gene was manually curated, validated according to HAVANA guidelines61 and identified in GENCODE releases M11 onwards as Gm11639. Efcab3/Efcab13 encode calcium-binding proteins and the new gene primarily consists of repeated EF-hand protein domains (Supplementary Fig. 12). Analysis of synteny and genome structure showed that the Efcab3 locus is largely conserved across other mammals, including most primates. Comparative gene prediction identified the full-length version in orangutan, rhesus macaque, bushbaby, and squirrel monkey. However, the locus contains a breakpoint at the common ancestor of chimpanzee, gorilla, and human (Homininae) due to a ~15 Mb intrachromosomal rearrangement that also deleted many of the internal EF-hand domain repeats (Fig. 3b and Supplementary Fig. 13). Analysis of Genotype-Tissue Expression (GTEx) data62 in humans showed that the EFCAB13 locus is expressed across many tissue types, with the highest expression measured in testis and thyroid. In contrast, the EFCAB3 locus only has low-level measurable expression in testis. This is consistent with the promoter of the full-length gene being present upstream from the EFCAB13 version, which is supported by H3K4me3 analysis (Supplementary Fig. 14). In mice, the gene Efcab3 is specifically expressed during development throughout many tissues with high expression in the upper layers of the cortical plate (see URLs) and is located in the immediate vicinity of the genomic 17q21.31 syntenic region linked to brain structural changes in both mice and humans63. We used CRISPR (clustered regularly interspaced short palindromic repeats) to create Efcab3-like mutant mice (Efcab3em1(IMPC)Wtsi, see Methods) and recorded 188 primary phenotyping measures (Supplementary Data 9). We also measured 40 brain parameters across 22 distinct brain structures as part of a high-throughput neuro-anatomical screen (Supplementary Tables 15 and 16, see Methods). Notably, brain size anomalies were identified in Efcab3-like mutant mice when compared to matched wild-type controls (Fig. 3c). Interestingly, the lateral ventricle was one the most severely affected brain structures exhibiting an enlargement of 65% (P = 0.007). The pontine nuclei were also increased in size by 42% (P = 0.001) and the cerebellum by 27% (P = 0.02); these two regions are involved in motor activity (Fig. 3d and Supplementary Fig. 15). The thalamus was also larger by 19% (P = 0.007). As a result, the total brain area parameter was enlarged by 7% (P = 0.006). Taken together, these results suggest a potential role of the Efcab3-like gene to regulate brain development and brain size from the forebrain to the hindbrain.
a, Comparative Augustus identified an unannotated 188 exon gene (Efcab3-like, red tracks). RNA-Seq splicing from two tissues (B = Brain, L = Liver, blue tracks) and five strains are displayed. Manual annotation extended this gene to 188 exons (lower red track). b, Evolutionary history of Efcab3-like in vertebrates including genome structure and surrounding genes. The mRNA structure of each gene is shown with white lines on the blue blocks. Novel coding sequence discovered in this study is shown in yellow. Notably, Efcab13 and Efcab3 are fragments of the novel gene Efcab3-like. A recombination event happened in the common ancestor of subfamily Homininae, which disrupted Efcab3-like in gorilla (G. gorilla) and human (H. sapiens). c, Schematic representation of 22 brain regions plotted in sagittal plane for Efcab3-like mutant male mice (16 weeks of age, n = 3) according to P values (two-tailed equal variance t-test, left). Corresponding brain regions are labelled with a number that is described below the panel (Supplementary Table 15). White coloring indicates a P value > 0.05 and grey indicates that the brain region could not be confidently tested due to missing data. Histograms showing the neuroanatomical features as percentage increase or decrease of the assessed brain regions in Efcab3-like mutant mice compared to matched controls (right). d, Representative sagittal brain images of matched controls (left) and Efcab3-like mutant (right), showing a larger cerebellum, enlarged lateral ventricle and increased size of the pontine nuclei (n = 3, see Supplementary Fig. 15).
Discussion
The completion of the mouse reference genome, based on the classical inbred strain C57BL/6J, generated a transformative resource for human and mouse genetics. Here we generate the first chromosome-scale genome assemblies for 12 classical and 4 wild-derived inbred strains, thus revealing at unprecedented resolution the striking strain-specific allelic diversity that encompasses 0.5–2.8% (14.4–75.5 Mb, excluding C57BL/6NJ) of the mouse genome. Accessing shared and distinct genetic information across the Mus lineage in parallel during assembly and gene prediction leads to the placement of novel alleles, the accurate annotation of many strain-specific gene family haplotypes and the detection of genes lowly expressed but partially supported in all strains (Fig. 3a).
Genetic diversity at gene loci, particularly those related to defence and immunity, is often the result of selection that, if retained, can lead to the rise of divergent alleles in a population64. We used the presence of dense clusters of hSNPs on the C57BL/6J reference genome as a marker for extreme polymorphism and examined the de novo assembly to explore the underlying genomic architecture. Examining the hSNPs in C57BL/6J and C57BL/6NJ, we find that the vast majority can be explained as occurring in remaining gaps or problematic regions of the reference genome. However, we are left with six loci (57 kb) enriched for hSNPs in C57BL/6J and C57BL/6NJ that do not have an obvious explanation and could be attributed to residual heterozygosity. Across all strains, hSNP regions account for 1.5–5.5% of protein-coding genes (Fig. 1c) and are over-represented with genes associated with immunity, sensory, sexual reproduction and behavioral phenotypes (Fig. 1d). Genes related to immunological processes, particularly gene families involved in mediating innate immune responses (for example Raet1 and Nlrp1), exhibit great diversity among the strains, reflecting strain-specific disease associations, responses and susceptibility. Interestingly, regions of strain haplotype diversity appear enriched for recent LINEs and LTRs (Fig. 1e). We observed several innate immunity gene families in mice with a high density of retrotransposons, which is the likely mechanism for diversification at these loci (for example, Nlrp1, Fig. 2d).
The challenge of generating multiple closely related mammalian genomes and annotation required new approaches to whole-genome alignment65, comparative creation of whole-chromosome scaffolds66, and comparative approaches to simultaneous genome annotation within a clade23,24. Mus is the first mammalian lineage to have multiple chromosome-scale genomes. Simultaneous access to many rodent species assemblies, in parallel with individual-level gene predictions, expression and long-read data, facilitated the accurate prediction of many strain-specific haplotypes and gene isoforms. This approach identified previously unannotated genes, including Efcab3-like, one of the largest known mouse genes (5,874 amino acids) that also appears conserved in mammals. Interestingly, the previously unannotated Efcab3-like gene is very close to the 17q21.31 syntenic region associated in humans to the Koolen–de Vries microdeletion syndrome (KdVS). Both mouse deletion models of this syntenic interval67, containing four genes (Crhr1, Spplc2, Mapt, and Kansl1; Fig. 3a) and an Efcab3-like knockout, showed analogous brain phenotypes, suggesting common cis-acting regulatory mechanisms as shown previously in the context of the 16p11.2 microdeletion syndrome68. Efacb3-like is conserved in orangutan but reversed in gorilla and appears to have split into two separate protein-coding genes, EFCAB3 and EFCAB13, in the Homininae lineage. Many novel genes and transcripts were identified across all of the strains, highlighting unexplored sequence variation across the Mus lineage. The addition of these genomes, in particular C57BL/6NJ, enabled the resolution of GRCm38 reference assembly issues and the improvement of several reference gene annotations. The assembly and alignment of a variety of haplotypes at loci heterogenous amongst the laboratory strains allows for analysis of regions previously not placed in the reference assembly. These regions are often of variable copy number between various haplotypes69. In particular, the wild-derived strains represent a rich resource of novel target sites, resistance alleles, genes and isoforms not present in the reference strain, or indeed many classical strains. For the first time, the underlying sequence at these loci is represented in strain-specific assemblies and gene predictions from across the inbred mouse lineage, which should facilitate increased dissection of complex traits.
URLs
A digital atlas of gene expression patterns in the mouse: http://www.genepaint.org
A pipeline used to comparatively annotate the mouse strains for the Mouse Genomes Project: https://github.com/ucsc-mus-strain-cactus/MouseGenomesAnnotationPipeline
SGA – String Graph Assembler – a de novo genome assembler: https://github.com/jts/sga
SNAP – Scalable Nucleotide Alignment Program – a new sequence aligner: http://snap.cs.berkeley.edu
ImageJ – an image processing toolkit: https://imagej.nih.gov/ij/
Methods
Sequencing
All DNA was obtained from the Jackson Laboratories from female mice (Supplementary Table 17). For the paired-end libraries, 1–3 μg DNA was sheared to 100–1,000 bp using a Covaris E210 or LE220 and size selected (350–450 bp) using magnetic beads (Ampure XP). Sheared DNA was subjected to Illumina paired-end DNA library preparation and PCR-amplified for six cycles. Amplified libraries were sequenced using the Illumina HiSeq platform as paired-end 100 base reads according to the manufacturer's protocol. Illumina sequencing compatible Mate Pair libraries were created at 3 and 6 kb according to the Sanger method70. The 10 kb Illumina Nextera libraries were prepared according to the manufacturer’s instructions (Illumina Nextera Sample Preparation Guide) with the addition of a size-selection step on the BluePippin (Sage Science).
For CAST/EiJ, PWK/PhJ, and SPRET/EiJ, a Chicago library was prepared as described previously19. Briefly, for each library, 500 ng of high molecular weight genomic DNA (>50 kb mean fragment size) was reconstituted into chromatin in vitro and fixed with formaldehyde. Fixed chromatin was then digested with restriction enzyme Mbo I, the 5′ overhangs were filled in with biotinylated nucleotides and then free blunt-ends were ligated. After ligation, cross-links were reversed and the DNA was purified from protein. Purified DNA was treated to remove biotin that was not internal to ligated fragments. The DNA was sheared to ~350 bp mean fragment size and sequencing libraries were generated using NEBNext Ultra enzymes and Illumina-compatible adapters. Biotin-containing fragments were then isolated using streptavidin beads before PCR enrichment of each library. The libraries were sequenced on an Illumina HiSeq to produce 2× 125 bp read pairs. The number of read pairs produced and fold physical coverage (1–50 kb pairs) for each genome was: 374 million, 34× for PWK/PhJ; 373 million, 41× for SPRET/EiJ; and 380 million, 77× for CAST/EiJ. Every sequencing lane was genotype checked against the mouse Hapmap SNP calls71 using the Samtools/Bcftools v1.1 'gtcheck' command.
De novo assembly
The initial contigs were assembled from the paired-end sequencing reads using SGA v0.9.43 (see URLs)72. Parameters for assembly are listed in Supplementary Table 18.
All of the mate-pair reads were aligned to GRCm38 with BWA-MEM v0.7.5, and duplicate fragments were removed with GATK MarkDuplicates v3.4. The subsequent reads were used as input to SOAPdenovo273 r240 to produce genome scaffolds (parameters given in Supplementary Table 19). To detect potential scaffold misjoins, we realigned the mate-pair library reads onto the SOAP2 scaffolds with BWA-MEM v0.7.5, walked along each scaffold (greater than 10 kb in size) in 5 kb step intervals and counted the number of 10 kb and 40 kb (where available) spanning fragments at each interval. Scaffolds were broken in locations where there was not a minimum number of 10 kb and 40 kb (where available) fragments that spanned the join. Scaffold break parameters are shown in Supplementary Table 20.
For CAST/EiJ, PWK/PhJ, and SPRET/EiJ, we further scaffolded the assemblies with Dovetail Genomics long-range libraries. Each input genome assembly, along with its associated Chicago library read pairs in FASTQ format, were used as input data for HiRise, a software pipeline designed specifically to scaffold genomes using Chicago library data2. Shotgun and Chicago library sequences were aligned to the draft input assembly using a modified SNAP read mapper (see URLs). The separations of Chicago read pairs mapped within draft scaffolds were analyzed by HiRise to produce a likelihood model for genomic distance between read pairs. The model was used to identify putative misjoins and score prospective joins. After scaffolding, shotgun sequences were used to close gaps between contigs.
Pseudochromosomes
The scaffolds were assembled into chromosome-scale scaffolds using Ragout v2.0. Ragout identifies large conserved regions between genomes (hierarchical synteny blocks) by combining the whole genome alignment with a de Bruijn graph simplification algorithm66. Assembly scaffolds are further joined into chromosomes so as to minimize the number of structural differences (such as inversions or chromosomal translocations) between references and a target genome. We used the C57BL/6J GRCm38 sequence as a single reference and found that 95% adjacent synteny block pairs from the assemblies were also adjacent in C57BL/6J reference.
Each of the genomes was assembled into a complete set of chromosomes with less than 5% of unlocalized sequence (Supplementary Data 10). On average, 10% of synteny block adjacencies in the assembled genomes were not presented in C57BL/6J reference. Ragout classified 38% of them as valid rearrangements and the rest as misassemblies (which were removed).
Gene prediction and annotation
Three techniques were used to produce the gene annotation for each mouse strain. First, whole-genome alignments produced by Progressive Cactus65 were used as input to transMap, producing an initial set of orthologs. These initial orthologs, along with strain-specific RNA-Seq (Supplementary Table 8), were input to AUGUSTUS74 one at a time to apply local strain-specific refinement. A consensus-finding algorithm was employed to decide between possible versions of an orthologous transcript. We also created a de novo set of strain-specific genes and isoforms from Comparative Augustus (AugustusCGP)24 using the strain-specific RNA-Seq and the progressive Cactus alignment. A subsequent round of consensus finding was employed to incorporate these transcripts into the final consensus annotation set.
The progressiveCactus whole genome alignments were used to project annotations from GENCODE VM860 onto each of the strain-specific assemblies using transMap75. These transMapped transcript alignments were evaluated by a series of binary classifiers that attempt to diagnose differences between the parent and target genome. These classifiers include evaluating if a transcript maps multiple times, the proportion of unknown bases, splice site validity, both frameshifting and non-frameshifting indels and small alignment gaps. These comparative transcripts were given to the gene-finding tool AUGUSTUS13 as strong hints (external evidence) in conjunction with weaker hints derived from all available RNA-Seq data for the given strain. The RNA-Seq hints were generated for each of the novel strains by aligning RNA-Seq reads to the native genome with the spliced aligner STAR16. The resulting read alignments were quality filtered by coverage (≥80%), identity (≥90%) and uniqueness; that is, when a read mapped to multiple loci, the best alignment for that read was only kept if the alignment score of the second best was considerably worse. For the remaining reads (approximately 70%), strain-specific exonpart and intron hints were generated. The transcripts resulting from transMap as well as AUGUSTUS were evaluated by a consensus-finding algorithm that attempts to use a combination of fidelity to the reference and a series of binary classifiers to construct a consensus gene set. See the Mouse Genomes Annotation pipeline documentation for details on this process (see URLs).
For each transMapped transcript alignment t, one way to identify its structure was a pipeline component we here refer to as AugustusTMR (TM = transMap, R = RNA-Seq). The aim was to try to produce all splice forms from the reference (parent) genome that probably also exist in the target genome. In the genomic region around t, AUGUSTUS was set to predict a gene structure without alternative splicing, using evidence from t itself as well as from all RNA-Seq alignments in that region. Thereby, the evidence from t on the location of exons, introns and start and stop codons was given a much higher weight in order to produce the original splice form, also in cases where the majority of target RNA-Seq suggests a different major splice form. However, when part of a transcript structure was unclear, for example an unalignable transcript part, RNA-Seq evidence could help fill in missing parts.
By design, AugustusTMR restricts gene finding to regions that align to a reference gene, and thus is not able to predict genes missing in the reference annotation or genes in unaligned regions. To find novel splice forms and genes, Augustus is run in comparative gene prediction (CGP) mode, a recent extension24 that takes a whole-genome alignment of related species or strains and simultaneously predicts coding genes in all input genomes. In AugustusCGP the same types of evidence can be incorporated for either a subset or all species/ strains. With the genome alignment, evidence is transferred across genomes. This makes it possible to exploit the combined evidence for gene finding and to discover genes that, for example, are only weakly expressed and partially supported in the reference strain but that have a high expression in other strains. In this application, two different types of evidence are used: the RNA-Seq hints for each of the novel strains from above; and annotation evidence from GENCODE VM8 for the C57BL/6J reference strain. For the latter, CDS and intron hints were generated from the GENCODE VM8 protein-coding gene set for the reference strain.
The resulting AugustusCGP gene sets were quality filtered based on how well the exon-intron structure of a transcript was supported by the combined RNA-Seq evidence (≥80% of the introns with splice junction support and ≥80% of CDS exons with a read coverage of at least ten reads per kilobase of mRNA). One of the challenges of gene finders is to distinguish coding genes from pseudogenes and expressed non-coding genes that contain partial open reading frames. All AugustusCGP transcripts that partially aligned to a reference transcript annotated as pseudogene or non-coding gene were also discarded.
The AugustusCGP transcripts were incorporated into the consensus gene set through a subsequent round of consensus finding. Based on coordinate intersections, each transcript was assigned a putative parent gene, if possible. If multiple assignments were created, attempts to resolve them were made by finding if any gene had a Jaccard distance 0.2 greater than any other; otherwise, they were discarded. After parent assignment, they were aligned with BLAT to each coding transcript associated with the parent gene. For each AugustusCGP transcript, if it had a better match to the CDS of any of the assigned transcripts than the current consensus transcript, the latter was replaced. If the AugustusCGP transcript introduced new intron junctions supported by RNA-Seq, then it was incorporated as a new isoform of that gene. Finally, if the AugustusCGP transcript was not assigned to any gene, it was incorporated as a putative novel gene. This process allows for the rescue of genes lost in the first round of filtering and consensus finding, as well as the discovery of polymorphic pseudogenes in the laboratory mouse lineage.
For the strains with AugustusPB transcripts, they were combined with the AugustusCGP transcripts and placed through the same consensus-finding process described above. AugustusPB transcripts that could not be confidently assigned to parent transcripts were discarded and not evaluated for novel contribution.
The consensus gene sets were subsetted into a basic gene set following the methodology used by GENCODE60. Briefly, coding transcripts were retained if they were marked as having complete end information. If no complete transcripts are present, one longest CDS is picked for the gene. For non-coding transcripts, the fewest number of transcripts to keep at least 80% of present non-coding splice junctions were retained.
Sliding window analysis
Only coordinates in which at least one strain had a hSNP call were retained. These coordinates were then used to estimate the combined density of hSNPs using a 10 kb sliding window (step of 2 kb) across the mouse reference genome. Windows were grouped according to the number of hSNPs they contained. The windows were then ordered by density of SNP (lowest, 1 hSNP per 10 kb window, to highest). The top 5% of hSNP dense windows was identified and a shared density cut-off per 10 kb window calculated (equivalent to 71 hSNPs per 10 kb window). This represented the density at which the interval content and total unique overlapping base pairs was observed to be clustered around distinct loci (Supplementary Fig. 6a).
Strain-specific analyses
For each strain separately, the density of hSNPs in 10 kb sliding windows (step of 2 kb) was estimated. Only windows with greater than or equal to the shared density cut-off per window were retained. These windows were then intersected with GENCODE M8 gene annotations; the total number of unique genes and base pair positions overlapping pass windows for each strain was calculated (Fig. 1c). For each strain separately, coding genes from GENCODE M815 overlapping pass heterozygote dense windows were identified. Gene sets for each strain were then combined and, using PantherDB76, were classified based on protein class annotations (Fig. 1d, left). To establish an expected rate for each protein class, the same analysis was carried out using the entire protein-coding CDS annotated gene set from GENCODE M8. Strain-specific gene sets (Supplementary Data 3) and PantherDB classifications are contained in Supplementary Table 10. Genes involved in defense and immunity (the largest protein class represented by the combined gene set) were then retrieved and the strains that contributed genes to this protein class identified. Strain-specific defense genes are listed in Supplementary Data 4. To identify defense genes from the analysis shared among classical inbred strains and each of the wild-derived strains, each of the strain-specific gene sets were merged into five categories, namely classical inbred (BALB/cJ, CBA/J, DBA/2J, C3H/HeJ, 129S1/SvImJ, A/J, C57BL/6NJ, NOD/ShiLtJ, LP/J, NZO/HlLtJ, FVB/NJ and AKR/J), PWK/PhJ, CAST/EiJ, WSB/EiJ, and SPRET/EiJ (Fig. 1d, right).
Generation of Efcab3-like knockout mice
All mice were maintained in a specific pathogen-free facility with sentinel monitoring at standard temperature (19–23 °C) and humidity (55% ± 10%), on a 12 h dark, 12 h light cycle (lights on 7:30–19:00) and fed a standard rodent chow (LabDiet 5021–3, 9% crude fat content, 21% kcal as fat, 0.276 ppm cholesterol). Both food and water were available ad libitum. The mice were housed for phenotyping in groups of 3 or 4 mice per cage in either blue line (Tecniplast Seal Safe 1285L: overall dimensions of caging 365×207×140 mm3, floor area 530 cm2) or green line (Tecniplast GM500: overall dimensions of caging 391×199×160 mm3, floor area 501 cm2) individually ventilated caging receiving 60 air changes per hour. In addition to Aspen bedding substrate, standard environmental enrichment of a nestlet and a cardboard tunnel were provided. All animals were regularly monitored for health and welfare and were additionally checked before and after procedures. The Efcab3-like gene has previously been represented by two loci MGI:3651790 and MGI:1918144, corresponding to the 5′ and 3′ regions, respectively. Both loci have been targeted using a conditional approach as part of the International Knockout Mouse Consortium (IKMC) resource. The Efcab3-like gene was targeted using CRISPR/Cas9 methodology77. Briefly, the constitutive coding exon 5 (chromosome 11: 104700610-104700692, GRCm38), which is well-supported by RNA-Seq data in multiple tissues (ENSMUST00000212287; ENSMUSE00000376310 (ENSEMBL v90)) was deleted using the SpCas9 endonuclease to induce a frameshift mutation. Pairs of flanking guide RNAs (gRNAs) were designed using the WTSI Genome Editing (WGE) tool78 creating four gRNAs (two gRNAs 5′ and two gRNAs 3′ to the CE region, Supplementary Table 21). Cas9 mRNA (Trilink) together with the four gRNAs was injected into the cytoplasm of single-cell C57BL/6NTac zygotes. Injected embryos were briefly cultured and oviductal embryo transfer performed in 0.5 days postcoital pseudopregnant female recipients (CBA/C57BL/6J). F0 mice were screened for the exon deletion by a combination of end-point PCR and loss of wild-type allele quantitative PCR. Positive F0 mice were further bred with C57BL/6NTac mice. F1 mice were rescreened by PCR and breakpoints confirmed by Sanger sequencing (Supplementary Data 11). A single genotype-confirmed F1 mouse (Efcab3em1(IMPC)Wtsi) was used to generate mice for phenotyping. The care and use of mice in the study was carried out in accordance with UK Home Office regulations, UK Animals (Scientific Procedures) Act of 1986 under a UK Home Office license that approved this work, which was reviewed regularly by the WTSI Animal Welfare and Ethical Review Body.
Neuroanatomical studies of Efcab3-like knockout
Neuroanatomical studies were performed blind with experimenters not knowing the genotype of the mouse, on three 16-week-old matched control male mice in C57BL/6N background and three 16-week-old homozygous knockout of Efcab3. Standard operating procedures are described in more details elsewhere79. Mouse brain samples were immersion-fixed in 10% buffered formalin for 48 h, before paraffin embedding and sectioning at 5 μm thickness using a sliding microtome (Leica RM 2145). One precise sagittal section was stereostatically defined as the plane Lateral +0.72 mm of the Mouse Brain Atlas. Brain sections were double-stained using luxol fast blue for myelin and cresyl violet for neurons and scanned at cell-level resolution using the Nanozoomer whole-slide scanner (Hamamatsu Photonics). Using in-house ImageJ (see URLs) plugins, covariates, for example sample processing dates and usernames, were collected at every step of the procedure, as well as 40 brain morphological parameters of 25 area and 14 length measurements, and the number of cerebellar folia (Supplementary Table 15). This resulted in the quantification of 22 unique brain structures, including: (1) total brain area; (2) primary and secondary motor cortices; (3) pons; (4) cerebellar area, internal granular layer of the cerebellum and medial cerebellar nucleus; (5) lateral ventricle; (6) corpus callosum; (7) thalamus; (8) caudate putamen; (9) hippocampus and its associated features; (10) fimbria of the hippocampus; (11) anterior commissure; (12) stria medullaris; (13) fornix; (14) optic chiasm; (15) hypothalamus; (16) pontine nuclei; (17) substantia nigra; (18) fibers of the pons; (19) cingulate cortex; (20) dorsal subiculum; (21) inferior colliculus; and (22) superior colliculus. All samples were also systematically assessed for cellular ectopia (misplaced neurons). Neuroanatomical data (Supplementary Table 16) were analyzed using Student’s two-tailed equal variance test.
Further details of methods are given in the Supplementary Note.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The genome sequencing reads are available from the European Nucleotide Archive and the assemblies are part of BioProject PRJNA310854 (Supplementary Table 22). The genome assemblies and annotation are available via the Ensembl genome browser and the UCSC Genome Browser. Sequence accessions for the three immune-related loci on chromosome 11 are available from the European Nucleotide Archive (Supplementary Table 23).
References
Beck, J. A. et al. Genealogies of mouse inbred strains. Nat. Genet. 24, 23–25 (2000).
Church, D. M. et al. Lineage-specific biology revealed by a finished genome assembly of the mouse. PLoS Biol. 7, e1000112 (2009).
Svenson, K. L. et al. Multiple trait measurements in 43 inbred mouse strains capture the phenotypic diversity characteristic of human populations. J. Appl. Physiol. 102, 2369–2378 (2007).
Americo, J. L., Moss, B. & Earl, P. L. Identification of wild-derived inbred mouse strains highly susceptible to monkeypox virus infection for use as small animal models. J. Virol. 84, 8172–8180 (2010).
Ideraabdullah, F. Y. et al. Genetic and haplotype diversity among wild-derived mouse inbred strains. Genome Res. 14, 1880–1887 (2004).
Churchill, G. A. et al. The Collaborative Cross, a community resource for the genetic analysis of complex traits. Nat. Genet. 36, 1133–1137 (2004).
French, J. E. et al. Diversity Outbred mice identify population-based exposure thresholds and genetic factors that influence benzene-induced genotoxicity. Environ. Health Perspect. 123, 237–245 (2015).
Ferris, M. T. et al. Modeling host genetic regulation of influenza pathogenesis in the collaborative cross. PLoS Pathog. 9, e1003196 (2013).
Rasmussen, A. L. et al. Host genetic diversity enables Ebola hemorrhagic fever pathogenesis and resistance. Science 346, 987–991 (2014).
Kelada, S. N. P. et al. Integrative genetic analysis of allergic inflammation in the murine lung. Am. J. Respir. Cell Mol. Biol. 51, 436–445 (2014).
Mouse Genome Sequencing Consortium et al. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
Keane, T. M. et al. Mouse genomic variation and its effect on phenotypes and gene regulation. Nature 477, 289–294 (2011).
Yalcin, B. et al. Sequence-based characterization of structural variation in the mouse genome. Nature 477, 326–329 (2011).
Simpson, E. M. et al. Genetic variation among 129 substrains and its importance for targeted mutagenesis in mice. Nat. Genet. 16, 19–27 (1997).
Shultz, L. D., Ishikawa, F. & Greiner, D. L. Humanized mice in translational biomedical research. Nat. Rev. Immunol. 7, 118–130 (2007).
Skarnes, W. C. et al. A conditional knockout resource for the genome-wide study of mouse gene function. Nature 474, 337–342 (2011).
Flint, J. & Mott, R. Applying mouse complex-trait resources to behavioural genetics. Nature 456, 724–727 (2008).
Churchill, G. A., Gatti, D. M., Munger, S. C. & Svenson, K. L. The Diversity Outbred mouse population. Mamm. Genome 23, 713–718 (2012).
Putnam, N. H. et al. Chromosome-scale shotgun assembly using an in vitro method for long-range linkage. Genome Res. 26, 342–350 (2016).
Goios, A., Pereira, L., Bogue, M., Macaulay, V. & Amorim, A. mtDNA phylogeny and evolution of laboratory mouse strains. Genome Res. 17, 293–298 (2007).
Doran, A. G. et al. Deep genome sequencing and variation analysis of 13 inbred mouse strains defines candidate phenotypic alleles, private variation and homozygous truncating mutations. Genome Biol. 17, 167 (2016).
Yalcin, B. et al. The fine-scale architecture of structural variants in 17 mouse genomes. Genome Biol. 13, R18 (2012).
Fiddes, I. T. et al. Comparative Annotation Toolkit (CAT) – simultaneous clade and personal genome annotation. (2017). https://doi.org/10.1101/231118
König, S., Romoth, L. W., Gerischer, L. & Stanke, M. Simultaneous gene finding in multiple genomes. Bioinformatics 32, 3388–3395 (2016).
Zhang, Z. et al. PseudoPipe: an automated pseudogene identification pipeline. Bioinformatics 22, 1437–1439 (2006).
Liu, Q. et al. Sensory neuron-specific GPCR Mrgprs are itch receptors mediating chloroquine-induced pruritus. Cell 139, 1353–1365 (2009).
Weiner, J. A., Wang, X., Tapia, J. C. & Sanes, J. R. Gamma protocadherins are required for synaptic development in the spinal cord. Proc. Natl Acad. Sci. USA 102, 8–14 (2005).
Dummer, P. D. et al. APOL1 Kidney disease risk variants: an evolving landscape. Semin. Nephrol. 35, 222–236 (2015).
Capewell, P., Cooper, A., Clucas, C., Weir, W. & Macleod, A. A co-evolutionary arms race: trypanosomes shaping the human genome, humans shaping the trypanosome genome. Parasitology 142(Suppl, 1), S108–S119 (2015).
Monroe, K. M. et al. IFI16 DNA sensor is required for death of lymphoid CD4 T cells abortively infected with HIV. Science 343, 428–432 (2014).
Boniotto, M. et al. Population variation in NAIP functional copy number confers increased cell death upon Legionella pneumophila infection. Hum. Immunol. 73, 196–200 (2012).
Patierno, S. R. et al. Uteroglobin: a potential novel tumor suppressor and molecular therapeutic for prostate cancer. Clin. Prostate Cancer 1, 118–124 (2002).
Cai, Y. et al. Preclinical evaluation of human secretoglobin 3A2 in mouse models of lung development and fibrosis. Am. J. Physiol. Lung Cell. Mol. Physiol. 306, L10–L22 (2014).
Braunewell, K. H. & Gundelfinger, E. D. Intracellular neuronal calcium sensor proteins: a family of EF-hand calcium-binding proteins in search of a function. Cell Tissue Res. 295, 1–12 (1999).
Dickinson, M. E. et al. High-throughput discovery of novel developmental phenotypes. Nature 537, 508–514 (2016).
Ewing, A. D. et al. Retrotransposition of gene transcripts leads to structural variation in mammalian genomes. Genome Biol. 14, R22 (2013).
Schrider, D. R. et al. Gene copy-number polymorphism caused by retrotransposition in humans. PLoS Genet. 9, e1003242 (2013).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Giordano, J. et al. Evolutionary history of mammalian transposons determined by genome-wide defragmentation. PLoS Comput. Biol. 3, e137 (2007).
Liebenauer, L. L. & Slotnick, B. M. Social organization and aggression in a group of olfactory bulbectomized male mice. Physiol. Behav. 60, 403–409 (1996).
Saraiva, L. R. et al. Combinatorial effects of odorants on mouse behavior. Proc. Natl Acad. Sci. USA 113, E3300–E3306 (2016).
Ibarra-Soria, X. et al. Variation in olfactory neuron repertoires is genetically controlled and environmentally modulated. eLife 6, (2017).
Zhang, H., Hardamon, C., Sagoe, B., Ngolab, J. & Bui, J. D. Studies of the H60a locus in C57BL/6 and 129/Sv mouse strains identify the H60a 3’UTR as a regulator of H60a expression. Mol. Immunol. 48, 539–545 (2011).
Diefenbach, A., Jamieson, A. M., Liu, S. D., Shastri, N. & Raulet, D. H. Ligands for the murine NKG2D receptor: expression by tumor cells and activation of NK cells and macrophages. Nat. Immunol. 1, 119–126 (2000).
O’Sullivan, T., Dunn, G. P., Lacoursiere, D. Y., Schreiber, R. D. & Bui, J. D. Cancer immunoediting of the NK group 2D ligand H60a. J. Immunol. 1950 187, 3538–3545 (2011).
Ye, Z. et al. Expression of H60 on mice heart graft and influence of cyclosporine. Transplant. Proc. 38, 2168–2171 (2006).
Durrant, C. et al. Collaborative Cross mice and their power to map host susceptibility to Aspergillus fumigatus infection. Genome Res. 21, 1239–1248 (2011).
Lilue, J., Müller, U. B., Steinfeldt, T. & Howard, J. C. Reciprocal virulence and resistance polymorphism in the relationship between Toxoplasma gondii and the house mouse. eLife 2, e01298 (2013).
Levinsohn, J. L. et al. Anthrax lethal factor cleavage of Nlrp1 is required for activation of the inflammasome. PLoS Pathog. 8, e1002638 (2012).
Bustos, O. et al. Evolution of the Schlafen genes, a gene family associated with embryonic lethality, meiotic drive, immune processes and orthopoxvirus virulence. Gene 447, 1–11 (2009).
Bauernfeind, F. & Hornung, V. Of inflammasomes and pathogens – sensing of microbes by the inflammasome. EMBO Mol. Med. 5, 814–826 (2013).
Boyden, E. D. & Dietrich, W. F. Nalp1b controls mouse macrophage susceptibility to anthrax lethal toxin. Nat. Genet. 38, 240–244 (2006).
Broz, P. & Dixit, V. M. Inflammasomes: mechanism of assembly, regulation and signalling. Nat. Rev. Immunol. 16, 407–420 (2016).
Sastalla, I. et al. Transcriptional analysis of the three Nlrp1 paralogs in mice. BMC Genomics 14, 188 (2013).
Hunn, J. P., Feng, C. G., Sher, A. & Howard, J. C. The immunity-related GTPases in mammals: a fast-evolving cell-autonomous resistance system against intracellular pathogens. Mamm. Genome 22, 43–54 (2011).
Taylor, G. A. IRG proteins: key mediators of interferon-regulated host resistance to intracellular pathogens. Cell Microbiol. 9, 1099–1107 (2007).
Li, M. et al. Codon-usage-based inhibition of HIV protein synthesis by human schlafen 11. Nature 491, 125–128 (2012).
Stremlau, M. et al. The cytoplasmic body component TRIM5alpha restricts HIV-1 infection in Old World monkeys. Nature 427, 848–853 (2004).
Bell, T. A. et al. The paternal gene of the DDK syndrome maps to the Schlafen gene cluster on mouse chromosome 11. Genetics 172, 411–423 (2006).
Mudge, J. M. & Harrow, J. Creating reference gene annotation for the mouse C57BL6/J genome assembly. Mamm. Genome 26, 366–378 (2015).
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 22, 1760–1774 (2012).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Arbogast, T. et al. Mouse models of 17q21.31 microdeletion and microduplication syndromes highlight the importance of Kansl1 for cognition. PLoS Genet. https://doi.org/10.1371/journal.pgen.1006886 (2017).
Lanier, L. L. Evolutionary struggles between NK cells and viruses. Nat. Rev. Immunol. 8, 259–268 (2008).
Paten, B. et al. Cactus: Algorithms for genome multiple sequence alignment. Genome Res. 21, 1512–1528 (2011).
Kolmogorov, M. et al. Chromosome assembly of large and complex genomes using multiple references. Prepritn available at https://www.biorxiv.org/content/early/2018/02/11/088435 (2018).
Arbogast, T. et al. Mouse models of 17q21.31 microdeletion and microduplication syndromes highlight the importance of Kansl1 for cognition. PLoS Genet. 13, (2017).
Loviglio, M. N. et al. Chromosomal contacts connect loci associated with autism, BMI and head circumference phenotypes. Mol. Psychiatry 22, 836–849 (2017).
Srivastava, A. et al. Genomes of the Mouse Collaborative Cross. Genetics 206, 537–556 (2017).
Park, N. et al. An improved approach to mate-paired library preparation for Illumina sequencing. Methods Gener. Seq. 1, https://doi.org/10.2478/mngs-2013-0001 (2013).
Kirby, A. et al. Fine mapping in 94 inbred mouse strains using a high-density haplotype resource. Genetics 185, 1081–1095 (2010).
Simpson, J. T. & Durbin, R. Efficient de novo assembly of large genomes using compressed data structures. Genome Res. 22, 549–556 (2012).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1, 18 (2012).
Keller, O., Kollmar, M., Stanke, M. & Waack, S. A novel hybrid gene prediction method employing protein multiple sequence alignments. Bioinformatics 27, 757–763 (2011).
Stanke, M., Diekhans, M., Baertsch, R. & Haussler, D. Using native and syntenically mapped cDNA alignments to improve de novo gene finding. Bioinformatics 24, 637–644 (2008).
Mi, H., Muruganujan, A. & Thomas, P. D. PANTHER in 2013: modeling the evolution of gene function, and other gene attributes, in the context of phylogenetic trees. Nucleic Acids Res. 41, D377–D386 (2013).
Boroviak, K., Doe, B., Banerjee, R., Yang, F. & Bradley, A. Chromosome engineering in zygotes with CRISPR/Cas9. Genesis 54, 78–85 (2016).
Hodgkins, A. et al. WGE: a CRISPR database for genome engineering. Bioinformatics 31, 3078–3080 (2015).
Mikhaleva, A., Kannan, M., Wagner, C. & Yalcin, B. Histomorphological phenotyping of the adult mouse brain. Curr. Protoc. Mouse Biol. 6, 307–332 (2016).
Acknowledgements
This work was supported by the Medical Research Council (MR/L007428/1), BBSRC (BB/M000281/1), and the Wellcome Trust. D.J.A. is supported by Cancer Research-UK and the Wellcome Trust. M.K.S. was supported by a research grant from CONICYT/FONDECYT/REGULAR No.1171004 and the European Commission (EUFP7 BLUEPRINT grant HEALTH-F5-2011-282510). D.T.O. work was supported by Cancer Research UK (20412), the Wellcome Trust (202878/A/16/Z), and the European Research Council (615584). P.F. was supported by the Wellcome Trust (grant numbers WT108749/Z/15/Z, WT098051, WT202878/B/16/Z), the National Human Genome Research Institute (U41HG007234), and the European Molecular Biology Laboratory. The research leading to these results has received funding from the European Union’s Seventh Framework Programme (FP7/2007–2013) under grant agreement HEALTH-F4-2010-241504 (EURATRANS). C.E.M. is supported by R01 DK074656. We thank members of the Sanger Institute Mouse Pipelines teams (Mouse Informatics, Molecular Technologies, Genome Engineering Technologies, Mouse Production Team, Mouse Phenotyping) and the Research Support Facility for the provision and management of the mice. We thank V. Vancollie for assistance with phenotyping data.
Author information
Authors and Affiliations
Contributions
T.M.K., D.J.A., B.P., and J.F. designed and supervised the study. M.Q., L.S., N.P., L.Re., A.C., M.Du., and A.F.-S. prepared the samples and carried out the sequencing. J.Li., T.M.K., L.G., K.H., S.K.P., S.P., J.T., J.W., M.K., W.C., F.Y., K.W., B.F., and J.C. carried out the genome assembly. B.P., I.T.F., M.A., J.A., M.Di., D.D.D., D.E., A.F., M.G., J.G., J.H., S.N., P.M., F.C.P.N., L.Ro., M.S., C.Si., C.St., G.T., R.B., J.Lo., M.S.H., D.T., and J.Li. carried out the genome annotation. J.Li., A.G.D., T.M.K., B.P., I.T.F., M.A., P.D., D.W.L., X.I.S., R.D., P.F., C.E.M., R.M., and D.T.O. carried out the genome analysis. S.C., C.L., M.T., and B.Y. carried out the knockout generation and phenotyping.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figs. 1–16, Supplementary Tables 1–26 and Supplementary Note
Supplementary Data 1
Novel genes predicted in classical and wild-derived mouse strains
Supplementary Data 2
hSNP-dense regions in C57BL/6J and C57BL/6NJ and issues of GRCm38
Supplementary Data 3
Gene list in hSNP-dense regions
Supplementary Data 4
Defense and immunity genes found in hSNP-dense regions
Supplementary Data 5
Olfr CDS in CAST/EiJ de novo assembly
Supplementary Data 6
Assembly quality control of three immunity-related loci in WSB/EiJ, PWK/PhJ and CAST/EiJ
Supplementary Data 7
Reference corrections through strain assembly information
Supplementary Data 8
GENCODE corrections
Supplementary Data 9
Phenotype of Efcab3-like–/– mouse compared to control group
Supplementary Data 10
Assemblathon assembly stats
Supplementary Data 11
PCR confirmation of Efcab3-like knockout
Supplementary Data 12
PCR primers for assembly quality control
Supplementary Data 13
Definition of repeat classes and families
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lilue, J., Doran, A.G., Fiddes, I.T. et al. Sixteen diverse laboratory mouse reference genomes define strain-specific haplotypes and novel functional loci. Nat Genet 50, 1574–1583 (2018). https://doi.org/10.1038/s41588-018-0223-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0223-8
This article is cited by
-
Single-mitochondrion sequencing uncovers distinct mutational patterns and heteroplasmy landscape in mouse astrocytes and neurons
BMC Biology (2024)
-
Machine learning on alignment features for parent-of-origin classification of simulated hybrid RNA-seq
BMC Bioinformatics (2024)
-
NLRP inflammasomes in health and disease
Molecular Biomedicine (2024)
-
Sex-specific DNA-replication in the early mammalian embryo
Nature Communications (2024)
-
Retention of ES cell-derived 129S genome drives NLRP1 hypersensitivity and transcriptional deregulation in Nlrp3tm1Flv mice
Cell Death & Differentiation (2024)