Abstract
In embryonic stem cells (ESCs), developmental gene promoters are characterized by their bivalent chromatin state, with simultaneous modification by MLL2 and Polycomb complexes. Although essential for embryogenesis, bivalency is functionally not well understood. Here, we show that MLL2 plays a central role in ESC genome organization. We generate a catalog of bona fide bivalent genes in ESCs and demonstrate that loss of MLL2 leads to increased Polycomb occupancy. Consequently, promoters lose accessibility, long-range interactions are redistributed, and ESCs fail to differentiate. We pose that bivalency balances accessibility and long-range connectivity of promoters, allowing developmental gene expression to be properly modulated.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Raw data and processed information for the ChIP-seq, in situ Hi-C, ATAC-seq, PRO-seq, and RNA-seq sequencing experiments generated in this study have been deposited in the Gene Expression Omnibus (GEO) under accession GSE99530.
References
Tee, W. W. & Reinberg, D. Chromatin features and the epigenetic regulation of pluripotency states in ESCs. Development 141, 2376–2390 (2014).
Bernstein, B. E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006).
Azuara, V. et al. Chromatin signatures of pluripotent cell lines. Nat. Cell Biol. 8, 532–538 (2006).
Voigt, P. et al. Asymmetrically modified nucleosomes. Cell 151, 181–193 (2012).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Piunti, A. & Shilatifard, A. Epigenetic balance of gene expression by Polycomb and COMPASS families. Science 352, aad9780 (2016).
Hu, D. et al. The Mll2 branch of the COMPASS family regulates bivalent promoters in mouse embryonic stem cells. Nat. Struct. Mol. Biol. 20, 1093–1097 (2013).
Denissov, S. et al. Mll2 is required for H3K4 trimethylation on bivalent promoters in embryonic stem cells, whereas Mll1 is redundant. Development 141, 526–537 (2014).
Voigt, P., Tee, W. W. & Reinberg, D. A double take on bivalent promoters. Genes Dev. 27, 1318–1338 (2013).
Ku, M. et al. Genomewide analysis of PRC1 and PRC2 occupancy identifies two classes of bivalent domains. PLoS Genet. 4, e1000242 (2008).
Aloia, L., Di Stefano, B. & Di Croce, L. Polycomb complexes in stem cells and embryonic development. Development 140, 2525–2534 (2013).
Glaser, S. et al. Multiple epigenetic maintenance factors implicated by the loss of Mll2 in mouse development. Development 133, 1423–1432 (2006).
Laugesen, A. & Helin, K. Chromatin repressive complexes in stem cells, development, and cancer. Cell. Stem. Cell. 14, 735–751 (2014).
Mas, G. & Di Croce, L. The role of Polycomb in stem cell genome architecture. Curr. Opin. Cell Biol. 43, 87–95 (2016).
Sexton, T. & Cavalli, G. The role of chromosome domains in shaping the functional genome. Cell 160, 1049–1059 (2015).
Schoenfelder, S. et al. The pluripotent regulatory circuitry connecting promoters to their long-range interacting elements. Genome Res. 25, 582–597 (2015).
Schoenfelder, S. et al. Polycomb repressive complex PRC1 spatially constrains the mouse embryonic stem cell genome. Nat. Genet. 47, 1179–1186 (2015).
Entrevan, M., Schuettengruber, B. & Cavalli, G. Regulation of genome architecture and function by Polycomb proteins. Trends. Cell Biol. 26, 511–525 (2016).
Joshi, O. et al. Dynamic reorganization of extremely long-range promoter–promoter interactions between two states of pluripotency. Cell. Stem. Cell. 17, 748–757 (2015).
Denholtz, M. et al. Long-range chromatin contacts in embryonic stem cells reveal a role for pluripotency factors and polycomb proteins in genome organization. Cell. Stem. Cell. 13, 602–616 (2013).
Cruz-Molina, S. et al. PRC2 Facilitates the regulatory topology required for poised enhancer function during pluripotent stem cell differentiation. Cell. Stem. Cell. 20, 689–705.e9 (2017).
Kundu, S. et al. Polycomb repressive complex 1 generates discrete compacted domains that change during differentiation. Mol. Cell 65, 432–446.e5 (2017).
Kinkley, S. et al. reChIP-seq reveals widespread bivalency of H3K4me3 and H3K27me3 in CD4+ memory T cells. Nat. Commun. 7, 12514 (2016).
Glaser, S. et al. The histone 3 lysine 4 methyltransferase, Mll2, is only required briefly in development and spermatogenesis. Epigenetics Chromatin 2, 5 (2009).
Lubitz, S., Glaser, S., Schaft, J., Stewart, A. F. & Anastassiadis, K. Increased apoptosis and skewed differentiation in mouse embryonic stem cells lacking the histone methyltransferase Mll2. Mol. Biol. Cell. 18, 2356–2366 (2007).
Andreu-Vieyra, C. V. et al. MLL2 is required in oocytes for bulk histone 3 lysine 4 trimethylation and transcriptional silencing. PLoS Biol. 8, e1000453 (2010).
Orlando, D. A. et al. Quantitative ChIP-Seq normalization reveals global modulation of the epigenome. Cell Rep. 9, 1163–1170 (2014).
Schmitges, F. W. et al. Histone methylation by PRC2 is inhibited by active chromatin marks. Mol. Cell 42, 330–341 (2011).
Hu, D. et al. Not All H3K4 methylations are created equal: Mll2/COMPASS dependency in primordial germ cell specification. Mol. Cell 65, 460–475.e6 (2017).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Dekker, J., Marti-Renom, M. A. & Mirny, L. A. Exploring the three-dimensional organization of genomes: interpreting chromatin interaction data. Nat. Rev. Genet. 14, 390–403 (2013).
de Wit, E. et al. The pluripotent genome in three dimensions is shaped around pluripotency factors. Nature 501, 227–231 (2013).
Smith, E. & Shilatifard, A. Enhancer biology and enhanceropathies. Nat. Struct. Mol. Biol. 21, 210–219 (2014).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Mahat, D. B. et al. Base-pair-resolution genome-wide mapping of active RNA polymerases using precision nuclear run-on (PRO-seq). Nat. Protoc. 11, 1455–1476 (2016).
Chamberlain, S. J., Yee, D. & Magnuson, T. Polycomb repressive complex 2 is dispensable for maintenance of embryonic stem cell pluripotency. Stem Cells 26, 1496–1505 (2008).
Freire-Pritchett, P. et al. Global reorganisation of cis-regulatory units upon lineage commitment of human embryonic stem cells. eLife 6, e21926 (2017).
Dixon, J. R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015).
Morey, L. et al. Polycomb regulates mesoderm cell fate-specification in embryonic stem cells through activation and repression mechanisms. Cell Stem Cel. 17, 300–315 (2015).
Poh, Y. C. et al. Generation of organized germ layers from a single mouse embryonic stem cell. Nat. Commun. 5, 4000 (2014).
Pasini, D., Bracken, A. P., Hansen, J. B., Capillo, M. & Helin, K. The Polycomb group protein Suz12 is required for embryonic stem cell differentiation. Mol. Cell. Biol. 27, 3769–3779 (2007).
Margueron, R. et al. Role of the Polycomb protein EED in the propagation of repressive histone marks. Nature 461, 762–767 (2009).
Rickels, R. et al. An evolutionary conserved epigenetic mark of Polycomb response elements implemented by Trx/MLL/COMPASS. Mol. Cell 63, 318–328 (2016).
Ladopoulos, V. et al. The histone methyltransferase KMT2B is required for RNA polymerase II association and protection from DNA methylation at the MagohB CpG island promoter. Mol. Cell. Biol. 33, 1383–1393 (2013).
Singh, A. M. et al. Cell-cycle control of bivalent epigenetic domains regulates the exit from pluripotency. Stem Cell Reports 5, 323–336 (2015).
Herz, H. M. et al. Enhancer-associated H3K4 monomethylation by Trithorax-related, the Drosophila homolog of mammalian Mll3/Mll4. Genes Dev. 26, 2604–2620 (2012).
Hu, D. et al. The MLL3/MLL4 branches of the COMPASS family function as major histone H3K4 monomethylases at enhancers. Mol. Cell. Biol. 33, 4745–4754 (2013).
Yan, J. et al. Histone H3 lysine 4 monomethylation modulates long-range chromatin interactions at enhancers. Cell Res. 28, 204–220 (2018).
Morey, L., Santanach, A. & Di Croce, L. Pluripotency and epigenetic factors in mouse embryonic stem cell fate regulation. Mol. Cell. Biol. 35, 2716–2728 (2015).
Ballare, C. et al. Phf19 links methylated Lys36 of histone H3 to regulation of Polycomb activity. Nat. Struct. Mol. Biol. 19, 1257–1265 (2012).
Beringer, M. et al. EPOP functionally links elongin and Polycomb in pluripotent stem cells. Mol. Cell 64, 645–658 (2016).
Mendez, J. & Stillman, B. Chromatin association of human origin recognition complex, cdc6, and minichromosome maintenance proteins during the cell cycle: assembly of prereplication complexes in late mitosis. Mol. Cell. Biol. 20, 8602–8612 (2000).
Morey, L. et al. Nonoverlapping functions of the Polycomb group Cbx family of proteins in embryonic stem cells. Cell. Stem. Cell. 10, 47–62 (2012).
Aloia, L. et al. Zrf1 is required to establish and maintain neural progenitor identity. Genes Dev. 28, 182–197 (2014).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome. Biol. 10, R25 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome. Biol. 9, R137 (2008).
O’Leary, N. A. et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 44, D733–D745 (2016).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Tyner, C. et al. The UCSC Genome Browser database: 2017 update. Nucleic Acids Res. 45, D626–D634 (2017).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome. Biol. 15, 550 (2014).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads.EMBnet J. 17, 10–12 (2011).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Serra, F. et al. Automatic analysis and 3D-modelling of Hi-C data using TADbit reveals structural features of the fly chromatin colors. PLoS Comput. Biol. 13, e1005665 (2017).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization.Nat Methods 9, 999–1003 (2012).
Lajoie, B. R., Dekker, J. & Kaplan, N. The Hitchhikeras guide to Hi-C analysis: practical guidelines. Methods 72, 65–75 (2015).
Acknowledgements
This work would not have been possible without F. Stewart and M. Obst (Biotechnology Center, TU Dresden, Germany), who generously provided the C57B1/6 Mll2F/F mouse ESCs. We are indebted to L. Morey and members of the Di Croce laboratory for insightful discussions and critical reading of the manuscript. We thank V. A. Raker for scientific editing, and the CRG Genomics Unit for their help in genomic experiments. This work was partially supported by the European Research Council under the 7th Framework Program FP7/2007–2013 (ERC grant agreement 609989) and the European Union’s Horizon 2020 research and innovation programme (grant agreement 676556) to M.A.M.-R. We also acknowledge support of the Spanish Ministry of Economy and Competitiveness (BFU2016-75008-P), Centro de Excelencia Severo Ochoa 2013–2017 (SEV-2012-0208), AGAUR, and Fundació ‘La Marató de TV3’ to L.D.C. We also acknowledge a JSPS Research Fellowship for Young Scientists to support Y.A. of US National Cancer Institute grant R35CA197569 to A.S.H.
Author information
Authors and Affiliations
Contributions
G.M. and L.D.C. designed the study. G.M., C.B. and M.S. carried out ChIP-seq experiments; G.M. and C.B. conducted RNA-seq experiments. G.M. performed western blot, in situ Hi-C, and ATAC-seq experiments. E.B. performed bioinformatic analyses for ChIP-seq, RNA-seq, ATAC-seq, and PRO-seq data. Y.S. and M.A.M.-R. performed Hi-C data processing and analyses. D.H. and A.S. generated the catalytically inactive MLL2 mouse ESCs. Y.A. performed PRO-seq experiments. F.L.D. assisted with in situ Hi-C experiments and helped to interpret Hi-C analyses. All authors contributed to the discussion and interpretation of the results. L.D.C. supervised the experiments and provided intellectual support in the design and interpretation of the data. G.M. wrote the manuscript with input from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Characterization and validation of the catalog of bona fide bivalent genes.
a, Agarose gel image of n = 3 independent cell culture chromatin samples obtained from sonicating Mll2WT ESC lysates for reChIP-seq. Samples were prepared following the manufacturer’s instructions (Methods). b, ChIP-seq profiles of the indicated histone modifications around the TSS of the 3,868 bona fide bivalent genes (left) and a group of 9,948 non-bivalent genes (right). Non-bivalent genes correspond to genes marked by H3K4me3 alone. c, GO term analysis of the 3,868 bona fide bivalent genes showing enrichment in categories corresponding to morphogenesis and differentiation. d, reChIP–qPCR validations at six bivalent gene promoters in wild-type mESCs. ReChIP-seq experiments using H3K4me3–IgG or H3K27me3–IgG were used as background controls. Values represent the mean and error bars correspond to the s.e.m. of n = 3 independent cell cultures. e, Venn diagram showing that about 76% of the bona fide bivalent genes are also MLL2 targets by ChIP-seq. f, Distribution of 524 non-TSS peaks (peaks outside the regions ±2.5 kb around the TSS), which are mostly located at intergenic and intragenic regions of the genome. Peaks were identified by overlapping individual H3K4me3, H3K27me3, and MLL2 ChIP-seq with H3K4me3–H3K27me3 and H3K27me3–H3K4me3 reChIP-seq experiments.
Supplementary Figure 2 MLL2 knockout leads to increased PRC2 and PRC1 at bivalent promoters.
a, Heat map of MLL2, H3K4me3 and H3K27me3 ChIP-seq centered at the TSS (±5 kb) of 1,669 HD bivalent genes, in Mll2WT and Mll2KO ESCs. Genes are ranked by the degree of loss of H3K4me3 upon Mll2 depletion. Numbers show the correspondence between values and the color scale. b, ChIP-seq of H3K4me3 (left) or H3K27me3 (right) around the TSS of the indicated groups of genes (Methods and Supplementary Table 1). c, TSS (±5 kb) plots of histone modifications, SUZ12 or RING1B in Mll2WT and Mll2KO cells, at 1,669 HD bivalent genes (top) and 3,868 bona fide bivalent genes (bottom). ChIP-seq signals were normalized to Drosophila spike-in. d, Western blots of whole-cell extracts in Mll2WT and Mll2KO mESCs. e, Western blots of acid-extracted histones, with total histone H3 as loading control. Numbers correspond to the ratio of band intensities in Mll2KO relative to Mll2WT. f, Western blot of cellular fractionations of Mll2WT and Mll2KO mESCs into total, cytoplasm (cyto), total nuclear (Nuc), nucleoplasm (Np), or chromatin (Chr) fractions. Equal volumes of each fraction were loaded for comparison of band intensities. The Western blot experiments in d–f were repeated for n = 3 independent cell cultures with similar results. g, Scatterplot of significantly differentially expressed genes after Mll2 knockout (n = 2; adjusted P < 0.1). Genes significantly upregulated (orange) and downregulated (blue) are shown. h, Bar plot showing the proportion of genes deregulated after MLL2 depletion that are MLL2 targets by ChIP-seq. i, GO analysis of 744 genes downregulated upon Mll2 knockout. j, Boxplots of expression levels of the indicated groups of genes in Mll2WT and Mll2KO. A pseudocount of 0.1 was added to the total RNA RPKM values. The bottom and top of the boxes correspond to the first and third quartiles, and the line within represents the median (n = 2 independent cell cultures). Whiskers denote the interval within 1.5 times the interquartile range (IQR) from the median. P values were computed using the Wilcoxon test (two-sided). k, Metagene plot of RPB1 ChIP-seq at 1,669 HD bivalent genes in Mll2WT and Mll2KO. l, As in k for RNAPII phosphorylated at Ser5. m, Boxplot of the RNAPII-Ser5P level ±500 bp around the TSS of 1,669 HD bivalent genes in Mll2WT and Mll2KO ESCs. P values were computed using the Wilcoxon test (two-sided).
Supplementary Figure 3 MLL2 knockout induces minor changes in TADs and CTCF occupancy, while reshaping local interactions of bivalent promoters.
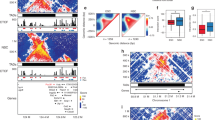
a, Left, percentage of TAD border conservation with two bins between Mll2WT and Mll2KO mESCs (n = 23 TAD conservation values per chromosome; the box extends from lower to upper quartiles, with a line at the median; outliers are shown as individual points). Middle, TAD border strength distributions (n = 2,885 aligned TAD borders; the bar plot shows the mean and 90% confidence interval). Right, relative TAD size distributions (n = 2,803 TAD borders for Mll2WT and n = 3,004 TAD borders for Mll2KO; the bar plot shows the mean and 90% confidence interval). b, CTCF peak enrichment in Mll2WT and Mll2KO mESCs. c, Boxplot of expression levels of genes within A (yellow) or B (blue) compartments in Mll2WT. A 0.1 pseudocount was added to total RNA RPKM values. The bottom and top of the boxes are the first and third quartiles, and the line is the median (n = 2 independent cell cultures). Whiskers correspond to 1.5 times the IQR from the median. P values were computed using the Wilcoxon test (two-sided). d, As in c for 148 bivalent genes within regions switching from A to B compartments in Mll2KO ESCs. P values were computed using the Wilcoxon test (two-sided). e, Hi-C Z-score interaction maps (2-kb resolution) with 500 bins upstream and downstream of the TSS of the indicated genes in wild-type or Mll2KO ESCs. Z scores were calculated by correcting interaction frequencies with genomic distance70. Gene locations are indicated with a circle; red arrows highlight interaction scores that were stronger in wild-type than Mll2KO. f, 3D interaction meta-plots (10-kb resolution) depicting observed versus expected interaction frequencies between bivalent TSS. Hubs visualize inter-TAD (2–10 Mb) crosstalk between TSSs. Areas are centered on the respective TSS (±150 kb).
Supplementary Figure 4 MLL2 catalytic activity contributes to the promoter-focused organization of interactions at bivalent genes.
a, Scatterplot of eigenvectors in Mll2WT versus Mll2CD ESCs. b, Pearson linear correlation between the proportion of bivalent genes with respect to the total number of genes in the genome, with the proportion of A-to-B with respect to B-to-A switchers. Red dots represent chromosomes with a ratio (B to A/A to B) higher than 1.0. c, Matrix of normalized differences in correlation coefficients between Mll2WT and Mll2CD on chromosome 2. The middle panel shows bins that switched from A to B compartments (green dots) and from B to A compartments (orange dots). Lower panels zoom in on the Hoxd cluster and nearby bins, with ChIP-seq of H3K4me3 and H3K27me3, and the behavior of A/B compartments. The genome is binned in 100-kb bins, and the region covers 73–76 Mb of chromosome 2. d, Top, average log10 value of observed versus expected interaction meta-matrices for the region ±250 kb from the TSS of the indicated genes. Interaction submatrices are corrected for direction of transcription and normalized by decay. The distribution of gene size is shown in gray. Bottom, differential interactions in Mll2CD as compared to WT. e, As in d for non-bivalent gene subsets. f,g, 3D interaction meta-plots (10-kb resolution) depicting observed versus expected interaction frequencies between the TSS of bivalent genes (f) and non-bivalent genes (g). Hubs visualize intra-TAD (300 kb–1 Mb) crosstalk between TSS. Areas shown are centered on the respective TSS (±150 kb).
Supplementary Figure 5 MLL2 knockout results in decreased accessibility of bivalent genes encoding proteins involved in neuronal formation and function.
a, Boxplot of the expression levels of 1,258 genes losing accessibility by ATAC-seq in Mll2KO as compared to Mll2WT ESCs. A 0.1 pseudocount was added to total RNA RPKM values. The bottom and top of the boxes correspond to the first and third quartiles, and the line within represents the median. Whiskers denote the interval within 1.5 times the IQR from the median (n = 2 independent cell cultures). P values were computed using the Wilcoxon test (two-sided). b, GO term analysis of 1,258 genes showing at least twofold decrease in ATAC-seq signals in Mll2KO as compared to Mll2WT. c, UCSC Genome Browser tracks of ATAC-seq and H3K4me3 and H3K27me3 ChIP-seq at the indicated bivalent genes in Mll2WT and Mll2KO mESCs. Experiments were repeated two times with similar results. d, Boxplots of ATAC-seq levels (n = 3 independent cell cultures) for genes transitioning from A to B compartments (left) and genes that transition from B to A compartments (right), after MLL2 depletion. P values were calculated using the Wilcoxon test (two-sided). e,f, Overlap between the 626 genes significantly downregulated in Mll2KO ESCs using PRO-seq and the 3,868 bona fide bivalent genes (e) or 4,213 non-bivalent genes (f). g, GO biological process analyses of the 626 genes significantly downregulated in Mll2KO ESCs using PRO-seq. The top five categories are shown. h, UCSC Genome Browser PRO-seq tracks (n = 2 independent cell cultures) in the indicated bivalent genes in Mll2WT and Mll2KO mESCs. Chromosomal coordinates are indicated.
Supplementary Figure 6 MLL2 depletion results in impaired developmental gene expression.
a, Representative images of embryoid bodies (EBs) derived from Mll2WT and Mll2KO ESCs at day 9 of the differentiation protocol. Images were taken at 100× magnification (scale bar, 400 uM). b, GO term analysis of 2,697 genes significantly upregulated in WT-derived 9-d-old EBs compared to ESCs (day 0). c, GO term analysis of 3,111 genes significantly upregulated in Mll2KO-derived 9-d-old EBs compared to Mll2KO ESCs (day 0). d, Venn diagram showing the overlap between the 2,697 genes upregulated in wild-type EBs from day 0 to day 9 and the 3,111 genes upregulated in Mll2KO EBs from day 0 to day 9. e, GO cellular component analysis of 1,159 genes upregulated only in Mll2KO EBs from day 0 to day 9. f, MGI mammalian phenotype term analysis of 1,159 genes upregulated only in Mll2KO EBs from day 0 to day 9.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Note
Supplementary Table 1
Lists of bivalent and non-bivalent genes identified by ChIP-seq and reChIP-seq experiments
Supplementary Table 2
List of primers utilized in ChIP–qPCR and re-ChIP–qPCR experiments
Supplementary Table 3
Differential gene expression analysis using DESeq2 in mESCs and EBs
Supplementary Table 4
Sequencing statistics for ChIP-seq, RNA-seq, ATAC-seq and in situ Hi-C experiments
Rights and permissions
About this article
Cite this article
Mas, G., Blanco, E., Ballaré, C. et al. Promoter bivalency favors an open chromatin architecture in embryonic stem cells. Nat Genet 50, 1452–1462 (2018). https://doi.org/10.1038/s41588-018-0218-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0218-5
This article is cited by
-
A low-input high resolution sequential chromatin immunoprecipitation method captures genome-wide dynamics of bivalent chromatin
Epigenetics & Chromatin (2024)
-
Loss of MLL3/4 decouples enhancer H3K4 monomethylation, H3K27 acetylation, and gene activation during embryonic stem cell differentiation
Genome Biology (2023)
-
Regulation, functions and transmission of bivalent chromatin during mammalian development
Nature Reviews Molecular Cell Biology (2023)
-
A CpG island-encoded mechanism protects genes from premature transcription termination
Nature Communications (2023)
-
Histone exchange sensors reveal variant specific dynamics in mouse embryonic stem cells
Nature Communications (2023)