Abstract
CRISPR–Cas genome editing creates targeted DNA double-strand breaks (DSBs) that are processed by cellular repair pathways, including the incorporation of exogenous DNA via single-strand template repair (SSTR). To determine the genetic basis of SSTR in human cells, we developed a coupled inhibition-cutting system capable of interrogating multiple editing outcomes in the context of thousands of individual gene knockdowns. We found that human Cas9-induced SSTR requires the Fanconi anemia (FA) pathway, which is normally implicated in interstrand cross-link repair. The FA pathway does not directly impact error-prone, non-homologous end joining, but instead diverts repair toward SSTR. Furthermore, FANCD2 protein localizes to Cas9-induced DSBs, indicating a direct role in regulating genome editing. Since FA is itself a genetic disease, these data imply that patient genotype and/or transcriptome may impact the effectiveness of gene editing treatments and that treatments biased toward FA repair pathways could have therapeutic value.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Sternberg, S. H. & Doudna, J. A. Expanding the biologist’s toolkit with CRISPR-Cas9. Mol. Cell 58, 568–574 (2015).
DeWitt, M. A. et al. Selection-free genome editing of the sickle mutation in human adult hematopoietic stem/progenitor cells. Sci. Transl. Med. 8, 360ra134 (2016).
Cox, D. B., Platt, R. J. & Zhang, F. Therapeutic genome editing: prospects and challenges. Nat. Med. 21, 121–131 (2015).
Maggio, I. & Gonçalves, M. A. Genome editing at the crossroads of delivery, specificity, and fidelity. Trends Biotechnol. 33, 280–291 (2015).
Chen, F. et al. High-frequency genome editing using ssDNA oligonucleotides with zinc-finger nucleases. Nat. Methods 8, 753–755 (2011).
Jasin, M. Genetic manipulation of genomes with rare-cutting endonucleases. Trends Genet. 12, 224–228 (1996).
Richardson, C. D., Ray, G. J., DeWitt, M. A., Curie, G. L. & Corn, J. E. Enhancing homology-directed genome editing by catalytically active and inactive CRISPR-Cas9 using asymmetric donor DNA. Nat. Biotechnol. 34, 339–344 (2016).
Sander, J. D. & Joung, J. K. CRISPR-Cas systems for editing, regulating and targeting genomes. Nat. Biotechnol. 32, 347–355 (2014).
Wright, A. V. et al. Rational design of a split-Cas9 enzyme complex. Proc. Natl Acad. Sci. USA 112, 2984–2989 (2015).
Kim, S., Kim, D., Cho, S. W., Kim, J. & Kim, J.-S. Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res. 24, 1012–1019 (2014).
Horlbeck, M. A. et al. Compact and highly active next-generation libraries for CRISPR-mediated gene repression and activation. eLife 5, e19760 (2016).
Huang, daW., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
George, B. et al. Regulation and function of Myb-binding protein 1A (MYBBP1A) in cellular senescence and pathogenesis of head and neck cancer. Cancer Lett. 358, 191–199 (2015).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Lin, S., Staahl, B. T., Alla, R. K. & Doudna, J. A. Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. eLife 3, e04766 (2014).
Bothmer, A. et al. Characterization of the interplay between DNA repair and CRISPR/Cas9-induced DNA lesions at an endogenous locus. Nat. Commun. 8, 13905 (2017).
Nakanishi, K. et al. Human Fanconi anemia monoubiquitination pathway promotes homologous DNA repair. Proc. Natl Acad. Sci. USA 102, 1110–1115 (2005).
Davis, L. & Maizels, N. Homology-directed repair of DNA nicks via pathways distinct from canonical double-strand break repair. Proc. Natl Acad. Sci. USA 111, E924–E932 (2014).
Hustedt, N. & Durocher, D. The control of DNA repair by the cell cycle. Nat. Cell Biol. 19, 1–9 (2016).
Walden, H. & Deans, A. J. The Fanconi anemia DNA repair pathway: structural and functional insights into a complex disorder. Annu. Rev. Biophys. 43, 257–278 (2014).
Moldovan, G.-L. & D’Andrea, A. D. How the Fanconi anemia pathway guards the genome. Annu. Rev. Genet. 43, 223–249 (2009).
Palovcak, A., Liu, W., Yuan, F. & Zhang, Y. Maintenance of genome stability by Fanconi anemia proteins. Cell Biosci. 7, 8 (2017).
Kim, J. M. et al. Inactivation of murine Usp1 results in genomic instability and a Fanconi anemia phenotype. Dev. Cell 16, 314–320 (2009).
Kee, Y. & D’Andrea, A. D. Molecular pathogenesis and clinical management of Fanconi anemia. J. Clin. Invest. 122, 3799–3806 (2012).
Adamo, A. et al. Preventing nonhomologous end joining suppresses DNA repair defects of Fanconi anemia. Mol. Cell 39, 25–35 (2010).
Renaud, E., Barascu, A. & Rosselli, F. Impaired TIP60-mediated H4K16 acetylation accounts for the aberrant chromatin accumulation of 53BP1 and RAP80 in Fanconi anemia pathway-deficient cells. Nucleic Acids Res. 44, 648–656 (2016).
Howard, S. M., Yanez, D. A. & Stark, J. M. DNA damage response factors from diverse pathways, including DNA crosslink repair, mediate alternative end joining. PLoS Genet. 11, e1004943 (2015).
van Overbeek, M. et al. DNA repair profiling reveals nonrandom outcomes at Cas9-mediated breaks. Mol. Cell 63, 633–646 (2016).
Ellis, N. A. et al. The Bloom’s syndrome gene product is homologous to RecQ helicases. Cell 83, 655–666 (1995).
Adelman, C. A. et al. HELQ promotes RAD51 paralogue-dependent repair to avert germ cell loss and tumorigenesis. Nature 502, 381–384 (2013).
Takata, K., Reh, S., Tomida, J., Person, M. D. & Wood, R. D. Human DNA helicase HELQ participates in DNA interstrand crosslink tolerance with ATR and RAD51 paralogs. Nat. Commun. 4, 2338 (2013).
Chun, J., Buechelmaier, E. S. & Powell, S. N. Rad51 paralog complexes BCDX2 and CX3 act at different stages in the BRCA1-BRCA2-dependent homologous recombination pathway. Mol. Cell. Biol. 33, 387–395 (2013).
Knipscheer, P. et al. The Fanconi anemia pathway promotes replication-dependent DNA interstrand cross-link repair. Science 326, 1698–1701 (2009).
Liang, C.-C. et al. The FANCD2–FANCI complex is recruited to DNA interstrand crosslinks before monoubiquitination of FANCD2. Nat. Commun. 7, 12124 (2016).
Federico, M. B. et al. Chromosomal integrity after UV irradiation requires FANCD2-mediated repair of double strand breaks. PLoS Genet. 12, e1005792 (2016).
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Nakanishi, K. et al. Interaction of FANCD2 and NBS1 in the DNA damage response. Nat. Cell Biol. 4, 913–920 (2002).
Roques, C. et al. MRE11–RAD50–NBS1 is a critical regulator of FANCD2 stability and function during DNA double-strand break repair. EMBO J. 28, 2400–2413 (2009).
Song, J. et al. RS-1 enhances CRISPR/Cas9- and TALEN-mediated knock-in efficiency. Nat. Commun. 7, 10548 (2016).
Orthwein, A. et al. A mechanism for the suppression of homologous recombination in G1 cells. Nature 528, 422–426 (2015).
Osborn, M. J. et al. Fanconi anemia gene editing by the CRISPR/Cas9 system. Hum. Gene Ther. 26, 114–126 (2015).
Skvarova Kramarzova, K. et al. CRISPR/Cas9-mediated correction of the FANCD1 gene in primary patient cells. Int. J. Mol. Sci. 18, E1269 (2017).
Thul, P. J. et al. A subcellular map of the human proteome. Science 356, eaal3321 (2017).
Knight, S. C. et al. Dynamics of CRISPR-Cas9 genome interrogation in living cells. Science 350, 823–826 (2015).
Petermann, E., Orta, M. L., Issaeva, N., Schultz, N. & Helleday, T. Hydroxyurea-stalled replication forks become progressively inactivated and require two different RAD51-mediated pathways for restart and repair. Mol. Cell 37, 492–502 (2010).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Gilbert, L. A. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661 (2014).
Kampmann, M., Bassik, M. C. & Weissman, J. S. Nat. Protoc. 9, 1825–1847 (2014).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25, 402–408 (2001).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Acknowledgements
This work used the Vincent J. Coates Genomics Sequencing Laboratory at UC Berkeley, supported by NIH S10 OD018174 Instrumentation Grant. We thank the Berkeley Macrolab for support with protein expression and purification. This work was supported by grants from the Li Ka Shing Foundation, the Heritage Medical Research Institute and the Fanconi Anemia Research Foundation. S.N.F. is an HHMI fellow of the Helen Hay Whitney Foundation. S.J.F. is a recipient of the SURF Rose Hills Fellowship and the Regents’ and Chancellor’s Fellowship. C.D.R. and J.E.C. are listed as inventors on a patent application based on this work.
Author information
Authors and Affiliations
Contributions
C.D.R. and J.E.C. designed the experiments. C.D.R., A.J.S. and S.N.F. performed the pooled screens. C.D.R., K.R.K. and S.J.F. performed the follow-up experiments. C.D.R., N.L.B. and J.E.C. analyzed the data. E.Z. and K.R.K. generated the knockout clones. C.D.R. and J.E.C. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
C.D.R. and J.E.C are inventors on a patent application related to this work.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 SSTR efficiency varies in different human cell lines.
Nine cell lines were edited using RNP targeting the EMX1 locus either without or with ssODN containing the PciI sequence. The edited locus was amplified, reannealed, and digested using T7 endonuclease I (t), which quantifies gene disruption, or the restriction enzyme PciI (R), which quantifies SSTR. Data presented are representative of independent experiments.
Supplementary Figure 2 BFP, GFP, and non-fluorescent populations are effectively resolved during a pooled CRISPRi screen.
a, Replicate 1 (of n = 2 independent experiments) of dCas9-KRAB cells infected with gRNA library and edited at the BFP locus. Live single cells were plotted to compare BFP and GFP intensities. Three populations were sorted: BFP+GFP– (BFP), BFP−GFP+ (GFP), and BFP−GFP− (UN). Percentages in each gate are presented for 100,000 cells. b, Total cells sorted for each edited population.
Supplementary Figure 3 Pooled CRISPRi screens identify DNA repair genes that contribute to multiple phenotypes.
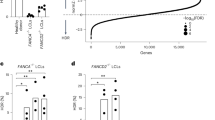
a, Essential genes in the K562 cell background. A volcano plot identifies genes from the DNA repair library enriched or depleted in CRISPRi cells relative to gRNA-only controls. Representative essential genes are highlighted in blue, negative control untargeted gRNAs are shown in black, and targeted gRNAs are shown in orange. Data were generated from n = 2 independent experiments. log2 fold change and Mann–Whitney P values were calculated as described11. b, Genes synthetic lethal with a DSB. A volcano plot identifies genes from the DNA repair library enriched or depleted in CRISPRi cells treated with Cas9 relative to untreated cell populations. Representative synthetic lethal genes are highlighted in blue, negative control untargeted gRNAs are shown in black, and targeted gRNAs are shown in orange. Data were generated from n = 2 independent experiments. log2 fold change and Mann–Whitney P values were calculated as described11.
Supplementary Figure 4 Characterizing non-SSTR phenotypes in pooled screen data.
a, Essential genes identified in this study substantially overlap with previous studies. The distribution of gRNAs in n = 2 unedited dCas9-KRAB cell libraries was separately compared to the gRNA distribution in an unedited K562 cell library and gene scores for each target gene were calculated. Essential genes were defined as those showing significant (P < 0.05, Mann–Whitney U test) depletion (log2 (fold change) < –0.5) in the dCas9-KRAB population and compiled into a single list. These essential genes were compared to the Horlbeck K562 dataset11, filtered for significance (P < 0.05) and magnitude of effect (phenotype < –0.2). Overlap between gene lists is presented as a Venn diagram. b, Essential genes are progressively lost from the library. Data presented are the mean ± s.d. of n = 2 independent experiments. c, Genes that are synthetically lethal with a single DSB are abruptly lost from the library following Cas9 treatment. Data presented are the mean ± s.d. of n = 2 independent experiments.
Supplementary Figure 5 Effective knockdown of DNA repair factors using siRNA.
a, Knockdown and editing of DSB repair factors reveals genetic differences between HDR and SSTR. K562 cells expressing a BFP reporter were treated with the indicated siRNAs and edited by co-administration of RNP targeting the BFP reporter with ssODN or dsDonor containing a BFP-to-GFP mutation. Data presented are the mean ± s.d. of n = 2 independent experiments. b, siRNA of RAD51, PARP1, and LIG4 in the K562 cells from a. Cells were siRNA treated for 48 h prior to editing. Fold depletion of the siRNA-target transcript over controls (ACTB, GAPDH) was measured by qPCR. Data presented represent the transcriptional state of the cells at the time of editing. The mean ± s.d. was calculated from n = 2 independent experiments. c, siRNA knockdown of FANCA and FANCF in human dermal fibroblasts. Cells were siRNA treated prior to editing (Fig. 2c) as described in b.
Supplementary Figure 6 Transcriptional manipulation of DNA repair factors using CRISPRi.
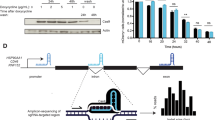
a, Stable CRISPRi cells targeting the indicated gene with or without re-expression of a cDNA of the indicated gene were harvested (Fig. 2a,b) and fold depletion of the indicated transcripts over control transcripts (ACTB, GAPDH) was measured by qPCR. The mean ± s.d. was calculated from n = 2 independent experiments. b, Epitope tags reveal robust re-expression of FA proteins. HA blots of parental and cDNA re-expression cell lines are presented along with predicted sizes for re-expressed genes. Data are representative of n = 2 independent experiments. c, Blotting for FANCA confirms knockdown/re-expression data. Top row, FANCA blot of parental (K562) cells, FANCA knockdown CRISPRi cells, and FANCA knockdown CRISPRi cells with re-expressed FANCA. Bottom row, loading controls for each sample. Data are representative of n = 2 independent experiments. d, Blotting for FANCD2 confirms knockdown/re-expression data. Top row, FANCD2 blot of parental (K562) cells, two FANCD2 knockdown cell lines, and FANCD2 knockdown g2 with re-expressed FANCD2. Bottom row, loading controls for each sample. Data are representative of n = 2 independent experiments. e, CD90 abundance validates transcription of cDNA constructs. Re-expression constructs express Thy1.1 co-transcriptionally with the cDNA, which was monitored by live-cell flow cytometry. Data are representative of n = 2 independent experiments.
Supplementary Figure 7 Flow cytometry and amplicon sequencing produce similar editing rates.
Editing at the BFP locus was quantified by flow cytometry (light blue), amplicon sequencing followed by in-house bioinformatic analysis (medium blue), or amplicon sequencing followed by CRISPRessoPOOL (Pinello, L. et al. Analyzing CRISPR genome-editing experiments with CRISPResso. Nat. Biotechnol. 34, 695–697 (2016)) analysis (dark blue). The mean ± s.d. was generated from n = 2 independent experiments.
Supplementary Figure 8 Knockout of FANCC and FANCG disrupts SSTR.
a, Editing strategy used to make clonal knockouts of FANCC and FANCG. Parental cells were edited with paired nucleases targeting the indicated sequences. Knockout was verified using amplification primers outside the edited locus. b, Paired cutting with CRISPR–Cas9 effectively removes the first exon of FANCC and FANCG. Diagnostic PCR (top) and western blotting (bottom) was performed to verify exon removal and elimination of gene product for FANCC or FANCG in clonal cell lines. Predicted sizes for genomic PCR amplifying the first exon of FANCC or FANCG (blue triangle; FANCC, 600 bp; FANCG, 950 bp) or for predicted excision outcomes (FANCC, 190 bp, maroon arrow; FANCG, 470 bp, green arrow) are presented alongside gels. Data are representative of n = 2 independent experiments. c, Cells lacking FANCC or FANCG show diminished levels of SSTR. SSTR levels were measured in multiple, individually cloned FANCC (n = 4) or FANCG (n = 3) knockout cell lines and plotted alongside values for clonally derived wild-type cell lines (n = 5). The editing rate for each sample (circle) is presented along with the center value (mean, black bar) and s.d. (error bars) for each group. *P < 0.05 (Welch’s t-test).
Supplementary Figure 9 Schematic of amplicon sequencing used at the BFP or HBB loci.
Protospacer (gray) and PAM (black) are presented within amplified sequence. ssODN size and orientation are also displayed. Vertical hash marks indicate locations where ssODN and genomic sequences differ.
Supplementary Figure 10 Total editing remains consistent when SSTR is disrupted.
Indel (NHEJ, light blue), SSTR (medium blue), and total (dark blue) editing events were quantified for the indicated CRISPRi cell lines and normalized to untargeted controls. a, RNP targeting the HBB locus with donor ssODN. b, RNP targeting the HBB locus without ssODN. c, RNP targeting the BFP locus with donor ssODN. d, RNP targeting the BFP locus without ssODN. Data presented are the mean ± s.d. of n = 2 independent experiments.
Supplementary Figure 11 HELQ interaction partners have a role in SSTR.
a, An interaction map of HELQ reproduced from an earlier study (Adelman, 2013) illustrates direct physical interactions between HELQ and other complexes and reported interactions from the BIOGRID, STRING and MINT databases. b, HELQ interaction partners have a role in SSTR. K562 cells were siRNA or CRISPRi treated against the indicated target genes prior to editing at the BFP locus. Data presented are the mean ± s.d. of n = 2 independent experiments. c, HELQ interaction partners can be effectively depleted by siRNA or CRISPRi. Fold depletion of the target transcript over controls (ACTB, GAPDH) was measured by qPCR. The mean ± s.d. was calculated from n = 2 independent experiments.
Supplementary Figure 12 FANCD2 ChIP-seq identifies on- and off-target Cas9 cut sites.
a, FANCD2 ChIP-seq overlaps with Guide-seq36 sites. The table describes the top 20 cut sites for the VEGFA_site2 sgRNA or the 10 annotated cut sites for the HEK293_site1 sgRNA. Peaks identified by MACS2 for n = 2 FANCD2 ChIP-seq datasets (green highlighting) are presented along with calculated q values. b, FANCD2 is enriched at the HBB cut site in HEK293T cells. Presented data are ChIP-seq pileups generated from comparing RNP + donor (treatment; n = 2 independent experiments) and Apo (control; n = 2 independent experiments) samples. c, FANCD2 is enriched at VEGFA_site2 on- and off-target loci in HEK293T cells. Presented data are ChIP-qPCR fold enrichments generated from non-specific IgG ChIP and FANCD2 ChIP at the indicated loci. The mean ± s.d. was calculated from n = 2 independent experiments.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12
Supplementary Table 1
Molecular biology reagents
Supplementary Table 2
Processed screen data: collapsed gene scores
Rights and permissions
About this article
Cite this article
Richardson, C.D., Kazane, K.R., Feng, S.J. et al. CRISPR–Cas9 genome editing in human cells occurs via the Fanconi anemia pathway. Nat Genet 50, 1132–1139 (2018). https://doi.org/10.1038/s41588-018-0174-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0174-0
This article is cited by
-
The Fanconi anemia core complex promotes CtIP-dependent end resection to drive homologous recombination at DNA double-strand breaks
Nature Communications (2024)
-
Improving prime editing with an endogenous small RNA-binding protein
Nature (2024)
-
Enhancing homology-directed repair efficiency with HDR-boosting modular ssDNA donor
Nature Communications (2024)
-
Removal of TREX1 activity enhances CRISPR–Cas9-mediated homologous recombination
Nature Biotechnology (2024)
-
FANCD2–FANCI surveys DNA and recognizes double- to single-stranded junctions
Nature (2024)