Abstract
Although much focus is placed on cholera epidemics, the greatest burden occurs in settings in which cholera is endemic, including areas of South Asia, Africa and now Haiti1,2. Dhaka, Bangladesh is a megacity that is hyper-endemic for cholera, and experiences two regular seasonal outbreaks of cholera each year3. Despite this, a detailed understanding of the diversity of Vibrio cholerae strains circulating in this setting, and their relationships to annual outbreaks, has not yet been obtained. Here we performed whole-genome sequencing of V. cholerae across several levels of focus and scale, at the maximum possible resolution. We analyzed bacterial isolates to define cholera dynamics at multiple levels, ranging from infection within individuals, to disease dynamics at the household level, to regional and intercontinental cholera transmission. Our analyses provide a genomic framework for understanding cholera diversity and transmission in an endemic setting.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ali, M., Nelson, A. R., Lopez, A. L. & Sack, D. A. Updated global burden of cholera in endemic countries. PLoS Negl. Trop. Dis. 9, e0003832 (2015).
Clemens, J. D., Nair, G. B., Ahmed, T., Qadri, F. & Holmgren, J. Cholera. Lancet 390, 1539–1549 (2017).
Longini, I. M. et al. Epidemic and endemic cholera trends over a 33-year period in Bangladesh. J. Infect. Dis. 186, 246–251 (2002).
Kendall, E. A. et al. Relatedness of Vibrio cholerae O1/O139 isolates from patients and their household contacts, determined by multilocus variable-number tandem-repeat analysis. J. Bacteriol. 192, 4367–4376 (2010).
Faruque, S. M. et al. Reemergence of epidemic Vibrio cholerae O139, Bangladesh. Emerg. Infect. Dis. 9, 1116–1122 (2003).
Schwartz, B. S. et al. Diarrheal epidemics in Dhaka, Bangladesh, during three consecutive floods: 1988, 1998, and 2004. Am. J. Trop. Med. Hyg. 74, 1067–1073 (2006).
Chowdhury, F. et al. Vibrio cholerae serogroup O139: isolation from cholera patients and asymptomatic household family members in Bangladesh between 2013 and 2014. PLoS Negl. Trop. Dis. 9, e0004183 (2015).
Harris, J. B. et al. Susceptibility to Vibrio cholerae infection in a cohort of household contacts of patients with cholera in Bangladesh. PLoS Negl. Trop. Dis. 2, e221 (2008).
Sugimoto, J. D. et al. Household transmission of Vibrio cholerae in Bangladesh. PLoS Negl. Trop. Dis. 8, e3314 (2014).
George, C. M. et al. Genetic relatedness of Vibrio cholerae isolates within and between households during outbreaks in Dhaka, Bangladesh. BMC Genomics 18, 903 (2017).
Weil, A. A. et al. Clinical outcomes in household contacts of patients with cholera in Bangladesh. Clin. Infect. Dis. 49, 1473–1479 (2009).
Levade, I. et al. Vibrio cholerae genomic diversity within and between patients. Microb. Genom. 3, e000142 (2017).
Faruque, S. M. et al. An improved technique for isolation of environmental Vibrio cholerae with epidemic potential: monitoring the emergence of a multiple-antibiotic-resistant epidemic strain in Bangladesh. J. Infect. Dis. 193, 1029–1036 (2006).
Faruque, A. S. G. et al. Emergence of multidrug-resistant strain of Vibrio cholerae O1 in Bangladesh and reversal of their susceptibility to tetracycline after two years. J. Health Popul. Nutr. 25, 241–243 (2007).
Hendriksen, R. S. et al. Population genetics of Vibrio cholerae from Nepal in 2010: evidence on the origin of the Haitian outbreak. mBio 2, e00157-11 (2011).
Shah, M. A. et al. Genomic epidemiology of Vibrio cholerae O1 associated with floods, Pakistan, 2010. Emerg. Infect. Dis. 20, 13–20 (2014).
Eppinger, M. et al. Genomic epidemiology of the Haitian cholera outbreak: a single introduction followed by rapid, extensive, and continued spread characterized the onset of the epidemic. mBio 5, e01721-14 (2014).
Siddique, A. K. et al. Cholera epidemics in Bangladesh: 1985–1991. J. Diarrhoeal Dis. Res. 10, 79–86 (1992).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Page, A. J. et al. Robust high-throughput prokaryote de novo assembly and improvement pipeline for Illumina data. Microb. Genom. 2, e000083 (2016).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Wong, V. K. et al. Phylogeographical analysis of the dominant multidrug-resistant H58 clade of Salmonella Typhi identifies inter- and intracontinental transmission events. Nat. Genet. 47, 632–639 (2015).
Croucher, N. J. et al. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 43, e15 (2015).
Didelot, X. & Wilson, D. J. ClonalFrameML: efficient inference of recombination in whole bacterial genomes. PLoS Comput. Biol. 11, e1004041 (2015).
Lartillot, N., Lepage, T. & Blanquart, S. PhyloBayes 3: a Bayesian software package for phylogenetic reconstruction and molecular dating. Bioinformatics 25, 2286–2288 (2009).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Yu, G., Smith, D. K., Zhu, H., Guan, Y. & Lam, T. T.-Y. GGTREE: an R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 8, 28–36 (2017).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 44, W242–W245 (2016).
Page, A. J. et al. Roary: rapid large-scale prokaryote pan genome analysis. Bioinformatics 31, 3691–3693 (2015).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Page, A. J. et al. SNP-sites: rapid efficient extraction of SNPs from multi-FASTA alignments. Microb. Genom. 2, e000056 (2016).
Rambaut, A., Lam, T. T., Max Carvalho, L. & Pybus, O. G. Exploring the temporal structure of heterochronous sequences using TempEst (formerly Path-O-Gen). Virus Evol. 2, vew007 (2016).
To, T.-H., Jung, M., Lycett, S. & Gascuel, O. Fast dating using least-squares criteria and algorithms. Syst. Biol. 65, 82–97 (2016).
Mutreja, A. et al. Evidence for several waves of global transmission in the seventh cholera pandemic. Nature 477, 462–465 (2011).
Hunt, M. et al. ARIBA: rapid antimicrobial resistance genotyping directly from sequencing reads. Microb. Genom. e000131 (2017).
Didelot, X. et al. The role of China in the global spread of the current cholera pandemic. PLoS Genet. 11, e1005072 (2015).
Acknowledgements
This research was supported in part by NIAID grants R01 AI106878 to E.T.R., F.Q., S.B.C., F.C., A.I.K., Y.A.B. and R.C.C., R01 AI103055 to J.B.H., F.Q. and R.C.L., U01 AI058935 to S.B.C., F.Q., E.T.R., R.C.L. and J.B.H., U01 AI077883 to E.T.R. and F.Q., the Fogarty International Center-NIH D43 grant TW005572 to M.I.U. and T.R.B., as well as K43 TW010362 to T.R.B. This work was supported by the Wellcome Trust (grant 098051) to N.R.T. M.J.D. is supported by a Wellcome Trust Sanger Institute PhD Studentship. R.C.C. was supported by the Robert Wood Johnson Foundation Harold Amos Medical Faculty Development Program (grant 72424). We thank A. J. Page, J. Keane and the sequencing teams at the Wellcome Trust Sanger Institute. This work was supported by the International Centre for Diarrhoeal Disease Research, Bangladesh (icddr,b) which is grateful to the Governments of Bangladesh, Canada, Sweden and the UK for providing core/unrestricted support.
Author information
Authors and Affiliations
Contributions
F.Q., E.T.R., N.R.T., R.C.C., S.B.C., J.B.H. and R.C.L. designed the study. F.C. and A.I.K. provided patient care and management. F.C., A.I.K., M.I.U., A.P., Y.A.B., R.C.C., T.R.B., J.B.H. and R.C.L. performed the experiments. D.D., M.J.D. and A.M. analyzed the data. D.D. wrote the manuscript, with major contributions from N.R.T., M.J.D., E.T.R. and F.Q. All authors contributed to the editing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
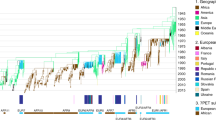
Supplementary Figure 1 Maximum likelihood phylogeny showing the relationships between Vibrio cholerae isolates.
Three isolates sampled in Dhaka, shown in green, do not belong to the 7th Pandemic El Tor (7PET) lineage. The location and date of isolation for each isolate are listed. V. metoecus and Vibrio sp. RC586 were used as outgroups for the phylogeny.
Supplementary Figure 2 Cholera incidence from icddr,b hospital in Dhaka, Bangladesh.
The diarrheal disease surveillance system at icddr,b enrolls every fiftieth individual for full analysis. The different panels discriminate between O1 serotypes and the O139 serogroup.
Supplementary Figure 3 Distribution of SNVs across households and individuals.
a, Pairwise comparison of SNVs shared across households ordered from least to greatest variation within a single household. b, Pairwise variation across individuals sampled more than once. c, Pairwise variability within technical replicates.
Supplementary Figure 4 Phylogenies of isolates sampled from individuals over the course of an infection.
Each panel depicts the relatedness of samples from the same individual. The scale is the number of SNVs per site.
Supplementary Figure 5 Loss of CTX bacteriophage within an individual.
The phylogenetic relatedness of the isolates from this individual is shown in the top diagram. The bottom panel shows the coverage of reads mapped to the CTXϕ region of the reference genome N16961 for samples from day 2 and day 4.
Supplementary Figure 7 Time-scaled phylogeny for the 7PET V. cholerae lineage.
The tips are colored according to the geographic origin of the isolates. The nodes are in the same order as in Fig. 5.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7
Supplementary Table 1
Metadata associated with the 303 Vibrio cholerae isolates from this study
Supplementary Table 2
Accessions and metadata for the 813 Vibrio cholerae genomes used for the global phylogeny
Rights and permissions
About this article
Cite this article
Domman, D., Chowdhury, F., Khan, A.I. et al. Defining endemic cholera at three levels of spatiotemporal resolution within Bangladesh. Nat Genet 50, 951–955 (2018). https://doi.org/10.1038/s41588-018-0150-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0150-8
This article is cited by
-
Genomic epidemiology of Vibrio cholerae during a mass vaccination campaign of displaced communities in Bangladesh
Nature Communications (2023)
-
The potential of genomics for infectious disease forecasting
Nature Microbiology (2022)
-
Vibrio cholerae O139 genomes provide a clue to why it may have failed to usher in the eighth cholera pandemic
Nature Communications (2022)
-
Population genomics of Escherichia coli in livestock-keeping households across a rapidly developing urban landscape
Nature Microbiology (2022)
-
Genomics of the Argentinian cholera epidemic elucidate the contrasting dynamics of epidemic and endemic Vibrio cholerae
Nature Communications (2020)