Abstract
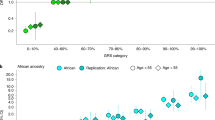
Genome-wide association studies (GWAS) and fine-mapping efforts to date have identified more than 100 prostate cancer (PrCa)-susceptibility loci. We meta-analyzed genotype data from a custom high-density array of 46,939 PrCa cases and 27,910 controls of European ancestry with previously genotyped data of 32,255 PrCa cases and 33,202 controls of European ancestry. Our analysis identified 62 novel loci associated (P < 5.0 × 10−8) with PrCa and one locus significantly associated with early-onset PrCa (≤55 years). Our findings include missense variants rs1800057 (odds ratio (OR) = 1.16; P = 8.2 × 10−9; G>C, p.Pro1054Arg) in ATM and rs2066827 (OR = 1.06; P = 2.3 × 10−9; T>G, p.Val109Gly) in CDKN1B. The combination of all loci captured 28.4% of the PrCa familial relative risk, and a polygenic risk score conferred an elevated PrCa risk for men in the ninetieth to ninety-ninth percentiles (relative risk = 2.69; 95% confidence interval (CI): 2.55–2.82) and first percentile (relative risk = 5.71; 95% CI: 5.04–6.48) risk stratum compared with the population average. These findings improve risk prediction, enhance fine-mapping, and provide insight into the underlying biology of PrCa1.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
08 January 2019
In the version of this article initially published, the name of author Manuela Gago-Dominguez was misspelled as Manuela Gago Dominguez. The error has been corrected in the HTML and PDF version of the article.
References
Goh, C. L. et al. Genetic variants associated with predisposition to prostate cancer and potential clinical implications. J. Intern. Med. 271, 353–365 (2012).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2016. CA Cancer J. Clin. 66, 7–30 (2016).
Cuzick, J. et al. Prevention and early detection of prostate cancer. Lancet Oncol. 15, e484–e492 (2014).
Altekruse, S. F. et al. Spatial patterns of localized-stage prostate cancer incidence among white and black men in the southeastern United States, 1999–2001. Cancer Epidemiol. Biomarkers Prev. 19, 1460–1467 (2010).
Stanford, J. L. & Ostrander, E. A. Familial prostate cancer. Epidemiol. Rev. 23, 19–23 (2001).
Bunker, C. H. et al. High prevalence of screening-detected prostate cancer among Afro-Caribbeans: the Tobago Prostate Cancer Survey. Cancer Epidemiol. Biomarkers Prev. 11, 726–729 (2002).
Ghadirian, P., Howe, G. R., Hislop, T. G. & Maisonneuve, P. Family history of prostate cancer: a multi-center case-control study in Canada. Int. J. Cancer 70, 679–681 (1997).
Grönberg, H., Damber, L. & Damber, J. E. Familial prostate cancer in Sweden: a nationwide register cohort study. Cancer 77, 138–143 (1996).
Matikaine, M. P. et al. Relatives of prostate cancer patients have an increased risk of prostate and stomach cancers: a population-based, cancer registry study in Finland. Cancer Causes Control 12, 223–230 (2001).
Eeles, R. et al. The genetic epidemiology of prostate cancer and its clinical implications. Nat. Rev. Urol. 11, 18–31 (2014).
Eeles, R. A. et al. Identification of 23 new prostate cancer susceptibility loci using the iCOGS custom genotyping array. Nat. Genet. 45, 385–391 (2013). e1–e2.
Al Olama, A. A. et al. A meta-analysis of 87,040 individuals identifies 23 new susceptibility loci for prostate cancer. Nat. Genet. 46, 1103–1109 (2014).
Al Olama, A. A. et al. Multiple loci on 8q24 associated with prostate cancer susceptibility. Nat. Genet. 41, 1058–1060 (2009).
Amundadottir, L. T. et al. A common variant associated with prostate cancer in European and African populations. Nat. Genet. 38, 652–658 (2006).
Eeles, R. A. et al. Identification of seven new prostate cancer susceptibility loci through a genome-wide association study. Nat. Genet. 41, 1116–1121 (2009).
Eeles, R. A. et al. Multiple newly identified loci associated with prostate cancer susceptibility. Nat. Genet. 40, 316–321 (2008).
Gudmundsson, J. et al. Genome-wide association and replication studies identify four variants associated with prostate cancer susceptibility. Nat. Genet. 41, 1122–1126 (2009).
Gudmundsson, J. et al. Genome-wide association study identifies a second prostate cancer susceptibility variant at 8q24. Nat. Genet. 39, 631–637 (2007).
Gudmundsson, J. et al. Common sequence variants on 2p15 and Xp11.22 confer susceptibility to prostate cancer. Nat. Genet. 40, 281–283 (2008).
Gudmundsson, J. et al. Two variants on chromosome 17 confer prostate cancer risk, and the one in TCF2 protects against type 2 diabetes. Nat. Genet. 39, 977–983 (2007).
Haiman, C. A. et al. Genome-wide association study of prostate cancer in men of African ancestry identifies a susceptibility locus at 17q21. Nat. Genet. 43, 570–573 (2011).
Kote-Jarai, Z. et al. Seven prostate cancer susceptibility loci identified by a multi-stage genome-wide association study. Nat. Genet. 43, 785–791 (2011).
Schumacher, F. R. et al. Genome-wide association study identifies new prostate cancer susceptibility loci. Hum. Mol. Genet. 20, 3867–3875 (2011).
Sun, J. et al. Evidence for two independent prostate cancer risk-associated loci in the HNF1B gene at 17q12. Nat. Genet. 40, 1153–1155 (2008).
Takata, R. et al. Genome-wide association study identifies five new susceptibility loci for prostate cancer in the Japanese population. Nat. Genet. 42, 751–754 (2010).
Thomas, G. et al. Multiple loci identified in a genome-wide association study of prostate cancer. Nat. Genet. 40, 310–315 (2008).
Yeager, M. et al. Genome-wide association study of prostate cancer identifies a second risk locus at 8q24. Nat. Genet. 39, 645–649 (2007).
Duggan, D. et al. Two genome-wide association studies of aggressive prostate cancer implicate putative prostate tumor suppressor gene DAB2IP. J. Natl. Cancer Inst. 99, 1836–1844 (2007).
Amin Al Olama, A. et al. A meta-analysis of genome-wide association studies to identify prostate cancer susceptibility loci associated with aggressive and non-aggressive disease. Hum. Mol. Genet. 22, 408–415 (2013).
Amos, C. I. et al. The OncoArray Consortium: a network for understanding the genetic architecture of common cancers. Cancer Epidemiol. Biomarkers Prev. 26, 126–135 (2017).
Delaneau, O., Marchini, J. & Zagury, J. F. A linear complexity phasing method for thousands of genomes. Nat. Methods 9, 179–181 (2011).
Howie, B. N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
de Bakker, P. I. et al. Practical aspects of imputation-driven meta-analysis of genome-wide association studies. Hum. Mol. Genet. 17, R122–R128 (2008).
Leongamornlert, D. et al. Frequent germline deleterious mutations in DNA repair genes in familial prostate cancer cases are associated with advanced disease. Br. J. Cancer 110, 1663–1672 (2014).
Mateo, J. et al. DNA-repair defects and Olaparib in metastatic prostate cancer. N. Engl. J. Med. 373, 1697–1708 (2015).
Meyer, A. et al. ATM missense variant P1054R predisposes to prostate cancer. Radiother. Oncol. 83, 283–288 (2007).
Sánchez-Huertas, C. & Lüders, J. The augmin connection in the geometry of microtubule networks. Curr. Biol. 25, R294–R299 (2015).
Volodko, N., Gordon, M., Salla, M., Ghazaleh, H. A. & Baksh, S. RASSF tumor suppressor gene family: biological functions and regulation. FEBS Lett. 588, 2671–2684 (2014).
Kudo, T. et al. The RASSF3 candidate tumor suppressor induces apoptosis and G1-S cell-cycle arrest via p53. Cancer Res. 72, 2901–2911 (2012).
Wissmann, C. et al. WIF1, a component of the Wnt pathway, is down-regulated in prostate, breast, lung, and bladder cancer. J. Pathol. 201, 204–212 (2003).
Yee, D. S. et al. The Wnt inhibitory factor 1 restoration in prostate cancer cells was associated with reduced tumor growth, decreased capacity of cell migration and invasion and a reversal of epithelial to mesenchymal transition. Mol. Cancer 9, 162 (2010).
Allen, H. F., Wade, P. A. & Kutateladze, T. G. The NuRD architecture. Cell. Mol. Life Sci. 70, 3513–3524 (2013).
Lai, A. Y. & Wade, P. A. Cancer biology and NuRD: a multifaceted chromatin remodelling complex. Nat. Rev. Cancer 11, 588–596 (2011).
Basta, J. & Rauchman, M. The nucleosome remodeling and deacetylase complex in development and disease. Transl. Res. 165, 36–47 (2015).
McDermott, D. F. & Atkins, M. B. PD-1 as a potential target in cancer therapy. Cancer Med. 2, 662–673 (2013).
Zhong, H. & Prentice, R. L. Bias-reduced estimators and confidence intervals for odds ratios in genome-wide association studies. Biostatistics 9, 621–634 (2008).
Kiciński, M., Vangronsveld, J. & Nawrot, T. S. An epidemiological reappraisal of the familial aggregation of prostate cancer: a meta-analysis. PLoS One 6, e27130 (2011).
Albright, F. et al. Prostate cancer risk prediction based on complete prostate cancer family history. Prostate 75, 390–398 (2015).
Wang, M. et al. Large-scale association analysis in Asians identifies new susceptibility loci for prostate cancer. Nat. Commun. 6, 8469 (2015).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics. 26, 2190–2191 (2010).
Amin Al Olama, A. et al. Risk analysis of prostate cancer in PRACTICAL, a multinational consortium, using 25 known prostate cancer susceptibility loci. Cancer Epidemiol. Biomarkers Prev. 24, 1121–1129 (2015).
Merico, D., Isserlin, R., Stueker, O., Emili, A. & Bader, G. D. Enrichment map: a network-based method for gene-set enrichment visualization and interpretation. PLoS One 5, e13984 (2010).
Joshi-Tope, G. et al. Reactome: a knowledgebase of biological pathways. Nucleic Acids Res. 33, D428–D432 (2005).
Schaefer, C. F. et al. PID: the Pathway Interaction Database. Nucleic Acids Res. 37, D674–D679 (2009).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Romero, P. et al. Computational prediction of human metabolic pathways from the complete human genome. Genome Biol. 6, R2 (2005).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Kandasamy, K. et al. NetPath: a public resource of curated signal transduction pathways. Genome Biol. 11, R3 (2010).
Thomas, P. D. et al. PANTHER: a library of protein families and subfamilies indexed by function. Genome Res. 13, 2129–2141 (2003).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Wang, L., Jia, P., Wolfinger, R. D., Chen, X. & Zhao, Z. Gene set analysis of genome-wide association studies: methodological issues and perspectives. Genomics 98, 1–8 (2011).
Wang, K., Li, M. & Hakonarson, H. Analysing biological pathways in genome-wide association studies. Nat. Rev. Genet. 11, 843–854 (2010).
Wang, K., Li, M. & Bucan, M. Pathway-based approaches for analysis of genomewide association studies. Am. J. Hum. Genet. 81, 1278–1283 (2007).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Acknowledgements
We pay tribute to Brian Henderson for his vision and leadership; he was a driving force behind the OncoArray project and he unfortunately passed away before seeing its fruition. We also thank the individuals who participated in these studies enabling this work.
Genotyping of the OncoArray was funded by the US National Institutes of Health (NIH) (U19 CA 148537 for the ELLIPSE project and X01HG007492 to the Center for Inherited Disease Research (CIDR) under contract no. HHSN268201200008I. Additional analytical support was provided by NIH NCI U01 CA188392 (to F.R.S.).
Funding for the iCOGS infrastructure came from the European Community's Seventh Framework Programme under grant agreement no. 223175 (HEALTH-F2-2009-223175) (COGS), Cancer Research UK (C1287/A10118, C1287/A 10710, C12292/A11174, C1281/A12014, C5047/A8384, C5047/A15007, C5047/A10692, and C8197/A16565), the NIH (CA128978) and Post-Cancer GWAS Initiative (1U19 CA148537, 1U19 CA148065, and 1U19 CA148112; the GAME-ON initiative), the Department of Defense (W81XWH-10-1-0341), the Canadian Institutes of Health Research (CIHR) for the CIHR Team in Familial Risks of Breast Cancer, Komen Foundation for the Cure, the Breast Cancer Research Foundation, and the Ovarian Cancer Research Fund.
This work was supported by the Canadian Institutes of Health Research; the European Commission's Seventh Framework Programme grant agreement no. 223175 (HEALTH-F2-2009-223175); Cancer Research UK grants C5047/A7357, C1287/A10118, C1287/A16563, C5047/A3354, C5047/A10692, and C16913/A6135; and NIH Cancer Post-Cancer GWAS initiative grant no. 1 U19 CA 148537-01 (the GAME-ON initiative).
We also thank the following for funding support: the Institute of Cancer Research and the Everyman Campaign, the Prostate Cancer Research Foundation, Prostate Research Campaign UK (now Prostate Action), the Orchid Cancer Appeal, the National Cancer Research Network UK, and the National Cancer Research Institute (NCRI) UK. We are grateful for the support of NIHR funding to the NIHR Biomedical Research Centre at the Institute of Cancer Research and the Royal Marsden NHS Foundation Trust.
The Prostate Cancer Program of Cancer Council Victoria also acknowledges grant support from the National Health and Medical Research Council, Australia (126402, 209057, 251533, 396414, 450104, 504700, 504702, 504715, 623204, 940394, and 614296), VicHealth, Cancer Council Victoria, the Prostate Cancer Foundation of Australia, the Whitten Foundation, PricewaterhouseCoopers, and Tattersall’s. E.A.O., D.M.K., and E.M.K. acknowledge the Intramural Program of the National Human Genome Research Institute for support.
The BPC3 was supported by the NIH, National Cancer Institute (cooperative agreements U01-CA98233 to D.J.H., U01-CA98710 to S.M.G., U01-CA98216 to E.R., and U01-CA98758 to B.E.H., and the Intramural Research Program of the NIH/National Cancer Institute, Division of Cancer Epidemiology and Genetics).
The CAPS GWAS study was supported by the Swedish Cancer Foundation (grant nos. 09-0677, 11-484, and 12-823), the Cancer Risk Prediction Center (CRisP; http://ki.se/en/meb/crisp/), a Linneus Centre grant (contract ID 70867902) financed by the Swedish Research Council, and the Swedish Research Council (grant nos. K2010-70X-20430-04-3, and 2014-2269).
PEGASUS was supported by the Intramural Research Program, Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH.
A full description of funding and acknowledgements can be found in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
Writing group: F.R.S., C.A.H., D.V.C., A.A.A.O., S.I.B., M. Ahmed, Z.S.K.-J., and R.A.E. Conception and coordination of OncoArray synthesis: F.R.S., C.I.A., D.F.E., S.J.C., C.A.H., B.E.H., and F.W. Database management: S.B., M.N.B., X.S., K.M., and A.L. Bioinformatics support: E.J.S., T.D., D. Leongamornlert, E.A., C.C.-B., and C.G. Genotyping calling and QC: L.F., J.D., and J.T. Provision of DNA samples and/or phenotypic data: V.L.S., S.M.G., B.D.C., C.M.T., P.J.G., I.M.T., J.B., S.C., L. Moya, J.C., L.H., W.T., G.P.R., H.G., M. Aly, T.N., P. Pharoah, N.P., J.S., T.L.J.T., C. Slavov, A.A., D.A., S.W., A.W., N.H., C.M.L.W., A.M.D., N.B., L.A.M., E.G., G.L.A., O.C.,G.C.T., S.K., L.E.B.F., K.D.S., T.F.O., M.B., L. Maehle, E.M.G., D.E.N., J.L.D., F.C.H., R.M.M., R.C.T., T.J.K., R.J.H., N.E.F., A.F., S.A.I., M.C. Stern, B.S.R., S.L.K., H.O., Y.-J.L., H.-W.Z., N.F., X.M., X.G., G.W., Z.S., G.G.G., M.C. Southey, R.J.M., L.M.F., A.S.K., B.M.K., J. Lubinski, G.C.-V., K.L.P., M.S., J.Y.P., T.A.S., H.-Y.L., J.L.S., C.C., D.W., J. Lim, E.A.O., M.S.G., B.G.N., S.F.N., M.W., R.B., M.A.R., P.I., H.B., K.C., B.H.,C.M., M.L., T.S., J.K., C.J.L., E.M.J., M.R.T., P. Paulo, M.C., S.L.N., L.S., Y.C.D., K.D.R., G.D.M., P.O., A.R., J. Llorca, S.-H.T., D.W.L., L.F.N., D. Lessel, M.G., T.K., R.K., N.U., S.S., C. Sipeky, V.M., M.P., F. Canzian, S.J., T.V.d.B., S. Larkin, P.A.T., C.A.H., M.G.D., J.E.C., M.E.M., M.J.R., G.J., R.H.N.v.S., F.M., T.T., Y.A.K., J.X., K.-T.K., L.C.-A., H.P., A.M., S.N.T., S.K.M., D.J.S., S. Lindstrom, C.T., J.M., D.J.H., E.R., A.S., F. Claessens, L.N.K., L.L.M., R.N.H., M.J.M., Z.C., P.K., F.W., S.J.C., B.E.H., C.A.-H., R.A.E., A.V., A.G.-C., B.F.D., G.C.-T., APCB investigators, IMPACT Study, Canary PASS investigators, BPC3, PRACTICAL, CAPS, PEGASUS, GAME-ON/ELLIPSE, and Profile Study Steering Committee. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1–19 and Supplementary Note
Rights and permissions
About this article
Cite this article
Schumacher, F.R., Al Olama, A.A., Berndt, S.I. et al. Association analyses of more than 140,000 men identify 63 new prostate cancer susceptibility loci. Nat Genet 50, 928–936 (2018). https://doi.org/10.1038/s41588-018-0142-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0142-8
This article is cited by
-
FORGEdb: a tool for identifying candidate functional variants and uncovering target genes and mechanisms for complex diseases
Genome Biology (2024)
-
An early-onset specific polygenic risk score optimizes age-based risk estimate and stratification of prostate cancer: population-based cohort study
Journal of Translational Medicine (2024)
-
Associations between gut microbiota and three prostate diseases: a bidirectional two-sample Mendelian randomization study
Scientific Reports (2024)
-
A distinct class of pan-cancer susceptibility genes revealed by an alternative polyadenylation transcriptome-wide association study
Nature Communications (2024)
-
Clinical features of prostate cancer by polygenic risk score
Familial Cancer (2024)