Abstract
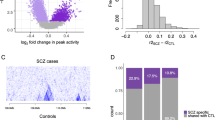
Genome-wide association studies (GWAS) have identified over 100 risk loci for schizophrenia, but the causal mechanisms remain largely unknown. We performed a transcriptome-wide association study (TWAS) integrating a schizophrenia GWAS of 79,845 individuals from the Psychiatric Genomics Consortium with expression data from brain, blood, and adipose tissues across 3,693 primarily control individuals. We identified 157 TWAS-significant genes, of which 35 did not overlap a known GWAS locus. Of these 157 genes, 42 were associated with specific chromatin features measured in independent samples, thus highlighting potential regulatory targets for follow-up. Suppression of one identified susceptibility gene, mapk3, in zebrafish showed a significant effect on neurodevelopmental phenotypes. Expression and splicing from the brain captured most of the TWAS effect across all genes. This large-scale connection of associations to target genes, tissues, and regulatory features is an essential step in moving toward a mechanistic understanding of GWAS.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Price, A. L., Spencer, C. C. & Donnelly, P. Progress and promise in understanding the genetic basis of common diseases. Proc. R. Soc. B 282, 20151684 (2015).
Soldner, F. et al. Parkinson-associated risk variant in distal enhancer of α-synuclein modulates target gene expression. Nature 533, 95–99 (2016).
Sekar, A. et al. Schizophrenia risk from complex variation of complement component 4. Nature 530, 177–183 (2016).
Claussnitzer, M. et al. FTO obesity variant circuitry and adipocyte browning in humans. N. Engl. J. Med. 373, 895–907 (2015).
Grubert, F. et al. Genetic control of chromatin states in humans involves local and distal chromosomal interactions. Cell 162, 1051–1065 (2015).
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Trynka, G. et al. Chromatin marks identify critical cell types for fine mapping complex trait variants. Nat. Genet. 45, 124–130 (2013).
Pickrell, J. K. Joint analysis of functional genomic data and genome-wide association studies of 18 human traits. Am. J. Hum. Genet. 94, 559–573 (2014).
Gusev, A. et al. Partitioning heritability of regulatory and cell-type-specific variants across 11 common diseases. Am. J. Hum. Genet. 95, 535–552 (2014).
Kichaev, G. et al. Integrating functional data to prioritize causal variants in statistical fine-mapping studies. PLoS Genet. 10, e1004722 (2014).
Won, H.-H. et al. Disproportionate contributions of select genomic compartments and cell types to genetic risk for coronary artery disease. PLoS Genet. 11, e1005622 (2015).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Degner, J. F. et al. DNase I sensitivity QTLs are a major determinant of human expression variation. Nature 482, 390–394 (2012).
McVicker, G. et al. Identification of genetic variants that affect histone modifications in human cells. Science 342, 747–749 (2013).
Kasowski, M. et al. Extensive variation in chromatin states across humans. Science 342, 750–752 (2013).
Kilpinen, H. et al. Coordinated effects of sequence variation on DNA binding, chromatin structure, and transcription. Science 342, 744–747 (2013).
Waszak, S. M. et al. Population variation and genetic control of modular chromatin architecture in humans. Cell 162, 1039–1050 (2015).
Taudt, A., Colomé-Tatché, M. & Johannes, F. Genetic sources of population epigenomic variation. Nat. Rev. Genet. 17, 319–332 (2016).
Farh, K. K.-H. et al. Genetic and epigenetic fine mapping of causal autoimmune disease variants. Nature 518, 337–343 (2015).
Moyerbrailean, G. A. et al. Which genetics variants in DNase-seq footprints are more likely to alter binding? PLoS Genet. 12, e1005875 (2016).
Gamazon, E. R. et al. A gene-based association method for mapping traits using reference transcriptome data. Nat. Genet. 47, 1091–1098 (2015).
Gusev, A. et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat. Genet. 48, 245–252 (2016).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 48, 481–487 (2016).
Fromer, M. et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. 19, 1442–1453 (2016).
Wright, F. A. et al. Heritability and genomics of gene expression in peripheral blood. Nat. Genet. 46, 430–437 (2014).
Li, Y. I. et al. RNA splicing is a primary link between genetic variation and disease. Science 352, 600–604 (2016).
Zhou, X., Carbonetto, P. & Stephens, M. Polygenic modeling with Bayesian sparse linear mixed models. PLoS Genet. 9, e1003264 (2013).
Nicolae, D. L. et al. Trait-associated SNPs are more likely to be eQTLs: annotation to enhance discovery from GWAS. PLoS Genet. 6, e1000888 (2010).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012). S1–S3.
Nica, A. C. et al. Candidate causal regulatory effects by integration of expression QTLs with complex trait genetic associations. PLoS Genet. 6, e1000895 (2010).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Won, H. et al. Chromosome conformation elucidates regulatory relationships in developing human brain. Nature 538, 523–527 (2016).
Purcell, S. M. et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Vilhjálmsson, B. J. et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am. J. Hum. Genet. 97, 576–592 (2015).
Palla, L. & Dudbridge, F. A fast method that uses polygenic scores to estimate the variance explained by genome-wide marker panels and the proportion of variants affecting a trait. Am. J. Hum. Genet. 97, 250–259 (2015).
Daetwyler, H. D., Villanueva, B. & Woolliams, J. A. Accuracy of predicting the genetic risk of disease using a genome-wide approach. PLoS One 3, e3395 (2008).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Ongen, H., Buil, A., Brown, A. A., Dermitzakis, E. T. & Delaneau, O. Fast and efficient QTL mapper for thousands of molecular phenotypes. Bioinformatics 32, 1479–1485 (2016).
Ryan, J. & Saffery, R. Crucial timing in schizophrenia: role of DNA methylation in early neurodevelopment. Genome Biol. 15, 495 (2014).
Akbarian, S. et al. The PsychENCODE project. Nat. Neurosci. 18, 1707–1712 (2015).
van de Geijn, B., McVicker, G., Gilad, Y. & Pritchard, J. K. WASP: allele-specific software for robust molecular quantitative trait locus discovery. Nat. Methods 12, 1061–1063 (2015).
Reilly, S. K. et al. Evolutionary changes in promoter and enhancer activity during human corticogenesis. Science 347, 1155–1159 (2015).
Golzio, C. et al. KCTD13 is a major driver of mirrored neuroanatomical phenotypes of the 16p11.2 copy number variant. Nature 485, 363–367 (2012).
Maillard, A. M. et al. The 16p11.2 locus modulates brain structures common to autism, schizophrenia and obesity. Mol. Psychiatry 20, 140–147 (2015).
McCarthy, S. E. et al. Microduplications of 16p11.2 are associated with schizophrenia. Nat. Genet. 41, 1223–1227 (2009).
Migliavacca, E. et al. A potential contributory role for ciliary dysfunction in the 16p11.2 600kb BP4–BP5 pathology. Am. J. Hum. Genet. 96, 784–796 (2015).
Föcking, M. et al. Proteomic and genomic evidence implicates the postsynaptic density in schizophrenia. Mol. Psychiatry 20, 424–432 (2015).
Sibley, C. R., Blazquez, L. & Ule, J. Lessons from non-canonical splicing. Nat. Rev. Genet. 17, 407–421 (2016).
Nelson, C. E. et al. In vivo genome editing improves muscle function in a mouse model of Duchenne muscular dystrophy. Science 351, 403–407 (2016).
Li, Y. I. et al. Annotation-free quantification of RNA splicing using LeafCutter. Nat. Genet. 50, 151–158 (2018).
Chen, C.-Y. et al. Improved ancestry inference using weights from external reference panels. Bioinformatics 29, 1399–1406 (2013).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Yang, J. et al. Genetic variance estimation with imputed variants finds negligible missing heritability for human height and body mass index. Nat. Genet. 47, 1114–1120 (2015).
Hormozdiari, F., Kostem, E., Kang, E. Y., Pasaniuc, B. & Eskin, E. Identifying causal variants at loci with multiple signals of association. Genetics 198, 497–508 (2014).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Jordan, D. M. et al. Identification of cis-suppression of human disease mutations by comparative genomics. Nature 524, 225–229 (2015).
Acknowledgements
We acknowledge M. Gandal, B. van de Geijn, A. Ko, P.-R. Loh, L. O’Connor, P. Pajukanta, and N. Zaitlen for helpful discussions. This research was funded by NIH grants F32GM106584 (A.G.), R01GM105857 (A.L.P.), R01MH109978 (A.L.P.), R01MH107649 (B.M.N.), R01MH105472 (G.E.C. and P.F.S.), R01HG009120 (B.P.), U01 MH103339-03S2 (D.H.G.), and R01 MH110927-02 (D.H.G.). H.K.F. was supported by the Fannie and John Hertz Foundation. The project described was also supported by award no. T32GM007753 from the National Institute of General Medical Sciences.This study was supported by a P50MH094268 grant (to N.K.). N.K. is suported as a distinguished Jean and George Brumley Professor. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institute of General Medical Sciences or the National Institutes of Health. We are grateful to the CommonMind Consortium and the PsychENCODE Consortium for making data publicly and readily available. Data were generated as part of the CommonMind Consortium supported by funding from Takeda Pharmaceuticals Company Limited; F. Hoffman-La Roche Ltd.; and NIH grants R01MH085542, R01MH093725, P50MH066392, P50MH080405, R01MH097276, RO1-MH-075916, P50M096891, P50MH084053S1, R37MH057881, R37MH057881S1, HHSN271201300031C, AG02219, AG05138, and MH06692. Brain tissue for the study was obtained from the following brain bank collections: the Mount Sinai NIH Brain and Tissue Repository, the University of Pennsylvania Alzheimers Disease Core Center, the University of Pittsburgh NeuroBioBank and Brain and Tissue Repositories, and the NIMH Human Brain Collection Core. CMC Leadership: P. Sklar, J. Buxbaum (Icahn School of Medicine at Mount Sinai), B. Devlin, D. Lewis (University of Pittsburgh), R. Gur, C.-G. Hahn (University of Pennsylvania), K. Hirai, H. Toyoshiba (Takeda Pharmaceuticals Company Limited), E. Domenici, L. Essioux (F. Hoffman-La Roche Ltd.), L. Mangravite, M. Peters (Sage Bionetworks), T. Lehner, and B. Lipska (NIMH). Data were generated as part of the PsychENCODE Consortium, supported by U01MH103339, U01MH103365, U01MH103392, U01MH103340, U01MH103346, R01MH105472, R01MH094714, R01MH105898, R21MH102791, R21MH105881, R21MH103877, and P50MH106934 awarded to S. Akbarian (Icahn School of Medicine at Mount Sinai), G. Crawford (Duke), S. Dracheva (Icahn School of Medicine at Mount Sinai), P. Farnham (USC), M. Gerstein (Yale), D. Geschwind (UCLA), T. M. Hyde (LIBD), A. Jaffe (LIBD), J. A. Knowles (USC), C. Liu (UIC), D. Pinto (Icahn School of Medicine at Mount Sinai), N. Sestan (Yale), P. Sklar (Icahn School of Medicine at Mount Sinai), M. State (UCSF), P. Sullivan (UNC), F. Vaccarino (Yale), S. Weissman (Yale), K. White (University of Chicago), and P. Zandi (JHU).
Author information
Authors and Affiliations
Consortia
Contributions
A.G., B.P., and A.L.P. designed the study. A.G., N.M., H.W., H.K.F., and Y.R. conducted analyses. M.K., L.S., A.S., G.E.C., D.H.G., N.K., and P.F.S. conducted and supervised experiments. The Psychiatric Genomics Consortium, S.M., B.M.N., R.A.O., M.C.O., and P.F.S. collected the data. A.G., B.P., and A.L.P. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–39, Supplementary Tables 1, 2, 4–18 and 20–25, and Supplementary Note
Supplementary Tables 3 and 19
Supplementary Tables 3 and 19
Rights and permissions
About this article
Cite this article
Gusev, A., Mancuso, N., Won, H. et al. Transcriptome-wide association study of schizophrenia and chromatin activity yields mechanistic disease insights. Nat Genet 50, 538–548 (2018). https://doi.org/10.1038/s41588-018-0092-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0092-1
This article is cited by
-
Decreased CNNM2 expression in prefrontal cortex affects sensorimotor gating function, cognition, dendritic spine morphogenesis and risk of schizophrenia
Neuropsychopharmacology (2024)
-
Investigating the role of common cis-regulatory variants in modifying penetrance of putatively damaging, inherited variants in severe neurodevelopmental disorders
Scientific Reports (2024)
-
Examination of a novel expression-based gene-SNP annotation strategy to identify tissue-specific contributions to heritability in multiple traits
European Journal of Human Genetics (2024)
-
Cell subtype-specific effects of genetic variation in the Alzheimer’s disease brain
Nature Genetics (2024)
-
Polygenic profiles define aspects of clinical heterogeneity in attention deficit hyperactivity disorder
Nature Genetics (2024)