Abstract
DNA methylation loss occurs frequently in cancer genomes, primarily within lamina-associated, late-replicating regions termed partially methylated domains (PMDs). We profiled 39 diverse primary tumors and 8 matched adjacent tissues using whole-genome bisulfite sequencing (WGBS) and analyzed them alongside 343 additional human and 206 mouse WGBS datasets. We identified a local CpG sequence context associated with preferential hypomethylation in PMDs. Analysis of CpGs in this context (‘solo-WCGWs’) identified previously undetected PMD hypomethylation in almost all healthy tissue types. PMD hypomethylation increased with age, beginning during fetal development, and appeared to track the accumulation of cell divisions. In cancer, PMD hypomethylation depth correlated with somatic mutation density and cell cycle gene expression, consistent with its reflection of mitotic history and suggesting its application as a mitotic clock. We propose that late replication leads to lifelong progressive methylation loss, which acts as a biomarker for cellular aging and which may contribute to oncogenesis.
Similar content being viewed by others
Main
Loss of 5-methylcytosine in both benign and malignant neoplasms was discovered more than 30 years ago1,2,3,4, yet the mechanisms that lead to this hypomethylation and its role in disease remain poorly understood. Genomic studies5,6,7,8,9 established that hypomethylation occurs in only about half the genome, coinciding with megabase-scale domains of repressive chromatin characterized by low gene density, low GC density, late replication timing, localization at the nuclear lamina, and Hi-C ‘B’ domains10,11. These regions were termed PMDs and were contrasted with ‘highly methylated domains’ (HMDs) that make up the remainder of the genome12. PMDs have been confirmed as a common feature of most epithelial cancers13 and other cancer types such as pediatric medulloblastoma14.
Conflicting evidence suggests that PMD hypomethylation could provide tumors with a growth advantage or alternatively may represent only a side effect of cancer15,16. An understanding of the earliest origins of this process could help elucidate a potential role of PMD hypomethylation in cancer initiation, yet results in precancer cell types have been conflicting. Since the 1980s, long-term cell culture has been known to result in significant DNA hypomethylation17, which was later discovered to occur primarily in PMDs8,12,18,19 and to accumulate stochastically in culture20,21. In primary uncultured tissues, one study showed the existence of PMDs in a few highly proliferative tissues such as peripheral white blood cells and placenta, but not in slowly dividing tissues like kidney, lung, or brain9. Other studies have shown the presence of global hypomethylation in placenta22 and more differentiated B cells23 and T cells24, but not in early-stage B cells or T cells nor in myelocytes23,24. The largest WGBS study of normal tissues concluded that PMDs were undetectable in 17 of 19 human tissue types studied (34 of 37 total samples), with the only exceptions being placenta and pancreas25. This reinforced the prevailing view that PMD hypomethylation may be restricted to a very limited set of normal cell types or only initiated upon exposure to environmental factors such as carcinogens26. Our group and one other group detected a small degree of PMD hypomethylation in normal mucosa adjacent to colon tumors5,6 but could not rule out the possibility of a precancer ‘field effect’ in these adjacent tissues.
Here we have analyzed a large and diverse set of WGBS experiments, including new tumor and adjacent normal data from eight common cancer types. By identifying a local sequence signature that defined the most strongly hypomethylated CpGs within PMDs, we were able to determine that most PMDs are shared by cancers and nearly all healthy human and mouse tissue types starting from fetal development. This allowed us to investigate the dynamics of hypomethylation across a large number of normal and malignant tissues and to define the relationship between PMDs, other chromatin features, and genomic mutational processes.
Results
Solo-WCGW CpGs are prone to hypomethylation
We sequenced The Cancer Genome Atlas (TCGA) tumors and adjacent normal samples using paired-end WGBS at ~15× sequence depth, to compile a set of 40 core tumor samples and 9 core normal samples (Methods and Supplementary Table 1). We first defined a set of shared PMDs and HMDs across the majority of our 49-core-sample set using an existing HMM (hidden Markov model)-based method, MethPipe27 (Methods and Supplementary Fig. 1a). Previous studies have suggested that DNA methylation is associated with local sequence context, including local CpG density28,29,30 and the nucleotides directly flanking the CpG29. We used the shared MethPipe PMD set (excluding CpG islands) to determine the local CpG density and tetranucleotide sequence contexts most predictive of DNA hypomethylation.
Low CpG density within windows of ±35 bp was optimal for predicting PMD-specific hypomethylation (Supplementary Fig. 1b). Additionally, CpGs flanked by an A or T (‘W’) on both sides (WCGW tetranucleotides) were consistently more prone to DNA hypomethylation than those flanked by a C or G (‘S’) on either (SCGW) or both (SCGS) sides (Fig. 1a and Supplementary Fig. 1c). In colon tumors and adjacent normal tissues, low CpG density and the WCGW context contributed additively to hypomethylation (Fig. 1b, top). The most hypomethylation-prone sequence context was at CpGs with the combination of zero neighboring CpGs (‘solo’) and the WCGW motif. In two adjacent normal colon samples, only these solo-WCGW CpGs showed significant hypomethylation (Fig. 1b, top). These same sequence dependencies were apparent in a colorectal tumor and normal colon tissue from mice (Fig. 1b, bottom). They were consistent within all other tumor and adjacent normal samples in the core set, using either the WGBS data (Supplementary Fig. 2a) or matched Illumina Infinium HumanMethylation450 (HM450) microarray data (Supplementary Fig. 2b). An additional 390 human and 206 mouse WGBS samples examined later exhibited the same pattern (Supplementary Fig. 3a,b), with the exception of three germ cell samples (Supplementary Fig. 3c).
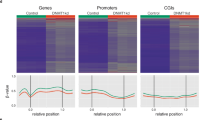
a, Each genomic CpG dinucleotide was placed into one of four CpG density categories (0, 1, 2, or 3+, depending on the number of additional CpGs within a ±35-bp window) and one of the three flanking nucleotide categories (SCGS, SCGW, or WCGW, with S denoting C or G and W denoting A or T). Because CpGs are palindromic, WCGS and SCGW were combined. Each of the 4 × 3 = 12 possible contexts is shown as columns for CpGs within common HMDs (left) or common PMDs (right). In the illustrations, a star indicates the target CpGs and solid circles indicate all neighboring CpGs within the window. The number of CpGs in each context is shown as a percentage of all genomic CpGs; for instance, the first column shows that 6% of all CpGs in the human genome are within HMDs, have 3+ flanking CpGs, and have the SCGS tetranucleotide context. b, The violin plots show the β-value distributions for CpGs in each context for five human tissues (two normal colon tissues and three colon tumors) and two mouse tissues (one normal colon tissue and one colon tumor (CRC)). Violin color indicates mean β value. Columns shaded orange and green indicate the most hypomethylation-resistant and most hypomethylation-prone categories, respectively. c, Average methylation values (non-overlapping 100-kb bins) across a 12-Mb section of chromosome 16p for the human colon samples. Values were calculated using all CpGs (left), only hypomethylation-resistant CpGs (orange; middle), or only solo-WCGW CpGs (green; right). CpG islands were removed in all analyses.
We focused all subsequent analyses on solo-WCGWs, representing 13% of all CpGs in the human genome. While they represent only the extreme of a hypomethylation process that affects other CpGs, focusing on solo-WCGWs alone enhanced the signal of PMD–HMD structure, especially in normal adjacent tissues and weakly hypomethylated tumors such as COAD-3518 (Fig. 1c). The relatively shallow hypomethylation in COAD-3518 could not be attributed to a greater fraction of non-cancer cells in this sample, as the cancer cell fraction in this sample was estimated by molecular estimates30 to be 80%, as compared to 51% for the more strongly hypomethylated COAD-A00R; this suggested that PMD depth was quantitative and driven by an independent property of the cancer cells.
In addition to enhancing the PMD–HMD signal in high-coverage WGBS data, solo-WCGW CpGs allowed accurate PMD structure to be determined with average genomic read coverage as low as 0.05× in downsampled bulk WGBS data (Supplementary Fig. 4a) and in low-coverage single-cell WGBS data31 (Supplementary Fig. 4b), suggesting a possible application for low-coverage or single-cell WGBS studies.
Most PMDs are shared across cancer and normal tissues
Genomic plots of solo-WCGW CpG mean methylation showed strong concordance between PMD locations in all samples in the core set (Fig. 2a). Comparing the average solo-WCGW methylation of the core tumors versus the core normal samples in multiscale plots (Fig. 2b) confirmed that PMDs ranging from 100 kb to 5 Mb in length32 were mostly overlapping between tumors and normal samples, but were less hypomethylated in the normal samples.
a, Average methylation values (non-overlapping 100-kb bins) for chromosome 16p, shown for the core tumor–normal dataset. The “tumor” field indicates tumors (black) versus adjacent normal samples, and the “this study” field indicates samples that were newly sequenced as part of this study (black). Within both the normal and tumor classes, tissue types are grouped and ordered by the average methylation level of samples from the group. For instance, “endometrium” is the first normal group because it has the highest methylation among the normal groups, and likewise for “GBM” among the tumor groups. b, Average methylation across all normal (top) and tumor (bottom) samples, calculated for multiple window sizes from 10 kb to 10 Mb (multiscale plots). c, The s.d. across the 9 core normal samples (top) and 40 core tumor samples (bottom) as multiscale plots. d, 100-kb s.d. values for all non-overlapping genomic bins, plotted for tumors (red histogram, x axis) versus normal samples (blue histogram, y axis). Bimodal peaks for each were identified via a Gaussian mixture model, and cutoffs dividing low and high s.d. values are shown by a dashed line for each axis. A scatter cloud shows the correlation of s.d. values between the tumors and normal samples, indicating the percentage of 100-kb bins falling into each of the four quadrants as well as Spearman’s ρ. e, Illustration of the method used to rescale each sample’s methylation values on the basis of genome-wide levels within a common set of PMDs (Methods). f, As in a, but using rescaled methylation values.
Given the high variability of solo-WCGW PMD hypomethylation across samples (Fig. 2a), we compared the s.d. of 100-kb bins across our core normal tissues and across core tumors, finding that PMDs had higher s.d. than HMDs within each group (Fig. 2c). Across the genome, s.d. was bimodally distributed within 100-kb bins in both the normal and tumor core groups (Fig. 2d), unlike mean methylation (Supplementary Fig. 5) and all other features examined (data not shown). While the highly variable nature of hypomethylation in PMDs has been noted previously5,7, it has not been used as a method for identifying PMDs. Using the bimodal s.d. peaks as a classifier resulted in a segmentation of the genome into HMDs and PMDs, with PMDs covering 63% of the genome in the core tumors (s.d. > 0.125) and 66% of the genome in the core normal samples (s.d. > 0.07). This simple method resulted in classifications for 100-kb bins that were 83% concordant between the normal and tumor groups (Fig. 2d). These PMDs covered 95% of the base pairs in PMDs previously reported in colorectal cancer6 and 93% of the base pairs in PMDs in the IMR90 fibroblast cell line12 (Supplementary Fig. 6). This s.d.-based classification of PMDs allowed us to rescale methylation values for individual samples on the basis of their sample-specific degree of PMD hypomethylation (Fig. 2e,f), further illustrating the high degree of concordance in PMD–HMD structure across tumor and normal samples.
Most PMDs are shared across developmental lineages
We investigated solo-WCGW PMD structure by combining our TCGA dataset with 343 previously published human and 206 mouse WGBS samples (Supplementary Table 1), examining solo-WCGW methylation averages with human samples arranged into six groups (Fig. 3) and mouse samples arranged into four groups (Fig. 4). As in the core set, the overall degree of hypomethylation varied widely, but PMD structure was largely shared for five of the six categories of human tissues. Common PMDs overlapped lamina-associated domains (LADs)33 and late-replicating domains, as expected (Figs. 3a and 4, bottom). The germline and embryo (GE) category was the only exception, with only some samples sharing PMDs (Fig. 3a, group GE; Fig. 4, group GE). Immortalized cell lines (cancer and non-cancer), with the exception of pluripotent embryonic cells, generally showed strongly hypomethylated PMDs that were shared with other groups (Fig. 3a, group of cell lines (CL); Fig. 4, group of embryonic stem cells (ESC)). More discussion on methylation maintenance in embryonic and induced pluripotent stem cells is given in the Supplementary Note and Supplementary Fig. 7a.
a, Average solo-WCGW methylation levels are plotted along chromosome 16p for 390 WGBS samples, organized into six groups: GE, germline and preimplantation embryo; FT, postimplantation embryonic/fetal samples, grouped first by embryonic versus extra-embryonic and then by average methylation; CL, cell lines; PN, postnatal non-blood normal tissue samples; PB, postnatal blood-derived samples; TM, primary tumors. Within each of the six groups, samples are organized by cell type (labeled by color). Lamin B1 signal and the replication timing in IMR90 lung fibroblasts are shown below the methylation heat maps. b, Mean methylation levels within each of the five major groups (excluding group GE), plotted as in Fig. 2b. c, The s.d. within each of the five major groups, plotted as in Fig. 2c. d, The s.d. for the 100-kb scale alone. e, The distribution of s.d. for all non-overlapping 100-kb genomic bins across all samples of the core tumor group (from d) is plotted on the y axis, as compared to each of four major groups (FT, CL, PN, and PB), shown on the x axis. Group GE was omitted owing to lack of PMD structure.
Average solo-WCGW methylation levels are plotted along a representative 30-Mb region of chromosome 17 in mouse. 206 WGBS samples are organized into four groups: ESC, embryonic stem cells; GE, germ line and embryos; FT, fetal tissues; PN, postnatal tissues. Grouping and ordering of samples were performed as described in Fig. 3. Lamin and replication timing are shown on the bottom of the heat map. Lamin A DamID values from wild-type mouse ESCs were downloaded from the Gene Expression Omnibus (GEO) with accession GSE6268369. Replication timing of day 9 differentiated ESCs was downloaded from GEO with accession GSE1798370.
In agreement with the TCGA tumor-adjacent normal samples, most disease-free postnatal tissues showed PMD structure shared with tumors and other groups (Fig. 3a, group of postnatal normal tissues (PN); Fig. 4, group of postnatal normal tissues (PN)). The normal human samples from Schultz et al.25 made up the majority of non-brain samples in our PN group and clearly had shared PMDs in our solo-WCGW analysis, while the original analysis of Schultz et al. identified PMDs in only 3 of these 37 samples. Most brain samples in the PN group were from a different study34, and these stood out as the one postnatal tissue type without clearly detectable PMDs in our analysis, which is possibly attributable to de novo DNA methylation in postmitotic brain cells34. Tissue types with high stem cell turnover35, including liver, colon, skin, and pancreas, displayed the strongest PMD hypomethylation.
All nucleated blood cell types showed shared PMD structure, in contrast to an earlier analysis of many of the same WGBS datasets36 that found PMD hypomethylation to be limited to the lymphoid lineage (Fig. 3a, group of postnatal blood samples (PB)). Both B cells and T cells could generally be divided into subgroups of strong versus weak hypomethylation. Subtypes having undergone antigen presentation and activation (for example, memory B and T cells, regulatory T cells, germinal center B cells, and plasma cells) fell into the strongly hypomethylated class, whereas naive B and T cells fell into the weakly hypomethylated class, consistent with earlier reports showing that B and T cell hypomethylation increased during maturation23,24. However, unlike these earlier reports, our solo-WCGW analysis showed that PMD hypomethylation was already clearly evident by the naive stage (Fig. 3a and Supplementary Fig. 7b). Lymphocyte activation involves clonal expansion (proliferation of individual B or T cells to produce large numbers of daughter cells with the same antigen specificity)37, and the dramatic hypomethylation that occurs after activation strengthens the notion that methylation loss accumulates during successive rounds of cell division (as shown explicitly in long-term cultures21). Our solo-WCGW analysis demonstrates that PMDs occur across all cell types of the myeloid lineage and are largely shared with other cell types (Fig. 3a and Supplementary Fig. 7c).
The tumor group (TM) consisted of 50 solid tumors (largely made up of the 40 core tumors shown previously) plus 50 hematopoietic malignancies (Fig. 3a, group TM). While hematopoietic tumors had more strongly hypomethylated PMDs than normal hematopoietic samples, they generally followed the trend established by their developmental origin: those derived from myeloid cells (AML) had shallower PMDs than those derived from lymphoid cells (CLL, MCL, TPLL, and MM) (one-way Wilcoxon test, P = 9.69 × 10−7). The notable exception among lymphoid-derived tumors was ALL, which had hypomethylation levels similar to those of normal lymphoid cells. The lower degree of hypomethylation in ALL (derived from childhood cases) may reflect the generally lower degree of hypomethylation in cells from younger individuals.
For five of the six cell type groups (excluding group GE), mean methylation across samples in the group (Fig. 3b), as well as s.d. (Fig. 3c,d), showed largely shared PMD structure. The s.d. was bimodally distributed across the genome in all five groups (Fig. 3e) and could thus be used to define PMD regions. For all of these five sample groups, the majority of PMDs defined by high-s.d. bins substantially overlapped PMDs defined earlier from the core tumor group (Fig. 3e and Supplementary Fig. 8). For example, 82% of high-s.d. bins were overlapping between the PN group and the core tumor group, and 84% were overlapping between the PB group and the core tumor group. Our findings reinforce the idea that a large set of cell-type-invariant PMDs dominate the hypomethylation landscape in most tissues.
PMD hypomethylation emerges during embryonic development
The presence of PMD hypomethylation in multiple fetal tissue types led us to further investigate solo-WCGW methylation in gametes and early developmental stages (Fig. 5a–c). Human sperm was highly methylated, with little discernable PMD structure aside from the pericentromeric region (Fig. 5a, group I), while mouse methylomes displayed consistent PMD structures throughout spermatogenesis (Supplementary Fig. 9). Human germinal vesicle oocytes had deep PMD hypomethylation (Fig. 5a, group II), although a subset of PMD boundaries appeared to differ from those in somatic tissues. The rapid and global demethylation that occurs within the inner cell mass (ICM) is thought to be an active process, attributable to a different mechanism than PMD-associated hypomethylation38. While ICM and blastocyst samples were strongly demethylated, they did retain weak PMDs with boundaries resembling those of oocytes rather than those of later somatic cell types (Fig. 5a, group III). Primordial germ cells (PGCs), which are set aside from the soma soon after implantation, showed an even more extreme erasure of DNA methylation than blastocysts, precluding any discernable PMD structure (Fig. 5a, group IV).
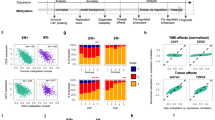
a,b, Multiscale solo-WCGW average plots are shown for samples divided into seven developmental stages, as diagrammed in b: paternal (I) and maternal (II) germ cells, implantation-related tissues (III), primordial germ cells (IV), embryonic soma (V), fetal soma (VI), and postnatal soma (VII). W, week. c, Rank-based analysis of the 792 genomic 100-kb bins from chromosome 16, comparing methylation ranks of the core tumors (y axis) to that of each developmental sample (x axis), with each axis going from a rank of 1 (lowest methylation) to the rank of the highest methylation (excluding bins with missing values in either of the samples). Greater correlation (shown by Spearman’s correlation coefficient, ρ) indicates stronger HMD–PMD structure.
Embryonic somatic tissues (Fig. 5a, group V) were rapidly remethylated across the genome and PMD structure could not be readily resolved, in contrast to what was observed in more mature fetal samples (Fig. 5a, group VI). Tissues sampled at different developmental stages showed a progressive emergence of PMD–HMD structure along organismal development (Fig. 5c). This analysis showed a substantial degree of similarity between PMD structure in brain tissues and PMD structure in other lineages, something that was not apparent from genomic plots. The substantial similarity of PMD structure detected between ICMs, ESCs, embryonic (<8 weeks) stages, and postnatal samples suggests that PMD hypomethylation may begin at the earliest stages of development. This interpretation is strengthened by the observation that the degree of hypomethylation observed at the fetal and postnatal stages for each cell type largely mirrors the lineage-specific hypomethylation rate within the same embryonic cell type.
PMD hypomethylation is associated with chronological age
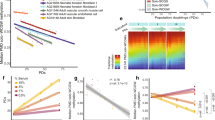
To investigate the link between PMD-associated hypomethylation and cumulative numbers of cell divisions, we tested whether the solo-WCGW methylation level within common PMDs was associated with donor age in different primary cell types. A strong age association was evident from the WGBS profile of sorted CD4+ T cells from a newborn versus those from a 103-year-old individual, with the latter being closer to a T cell–derived leukemia than to the newborn sample (Fig. 6a). To investigate age-related properties within larger studies only performed using the HM450 platform, we used the common PMDs derived from all WGBS samples to define a standard set of solo-WCGW PMD probes represented on HM450 (Methods). In these larger studies, peripheral blood mononuclear cell (PBMC) samples from newborns had significantly less PMD hypomethylation than those from elderly donors (Fig. 6b, left) and fetal liver samples had significantly less PMD hypomethylation than adult liver samples (Fig. 6b, right). Fetal tissues from four different developmental lineages showed nearly linear accumulation of hypomethylation from 9 weeks post-gestation to 22 weeks post-gestation (Fig. 6c). Despite small sample sizes, this was statistically significant for three of the four fetal tissue types. A similar association was observed between PMD hypomethylation and gestational age in multiple mouse fetal tissue types (Supplementary Fig. 10).
a, Multiscale solo-WCGW average plots are shown for CD4+ T cells from a newborn, CD4+ T cells from a 103-year-old (GSE31438), and T cell prolymphocytic leukemia (BLUEPRINT accession S016KWU1). b–f, Summarization of average PMD hypomethylation in HM450-based samples, obtained by averaging β values for 6,214 solo-WCGW probes mapped to common PMDs (Methods). b, Peripheral blood mononuclear cells (PBMCs) in newborns and nonagenarians (left; from GSE30870, P = 8.8 × 10−5, one-way Wilcoxon rank-sum test) and disease-free fetal and adult liver tissue (right; from GSE61278). The center lines of the box plots indicate the median, and the lower and upper bounds indicate the lower and upper quartiles. The lower and upper whiskers correspond to the smallest and largest methylation values. **P ≤ 0.001, Wilcoxon rank-sum test. c–f, HM450-based solo-WCGW averages versus age for individual donors for several tissue types. n is the number of donors or samples, r is Pearson’s product-moment correlation, b1 is the estimated rate of methylation loss, and P is the P value based on Pearson correlation tests. c, Four fetal tissue types at three prenatal time points (from GSE56515). d, Sun-exposed and sun-protected dermis and epidermis (from GSE51954). e, Sorted blood cells of the myeloid lineage (D1, GSE35069; D2, GSE56046). f, Sorted blood cells of the lymphoid lineage (D1, GSE35069; D3, GSE71955; D4, GSE59065).
An earlier study used the HM450 platform to investigate the effects of environmental (UV) exposure on PMD hypomethylation in human skin samples26. While the earlier study described PMD hypomethylation as only occurring within sun-exposed samples of the epidermal layer, our reanalysis of solo-WCGWs showed that both dermal and epidermal cells exhibited age-associated PMD hypomethylation without sun exposure but that this process was dramatically accelerated specifically in epidermal cells upon sun exposure (Fig. 6d). This suggests that, while PMD hypomethylation is a nearly universal process in aging, the degree of hypomethylation is a reflection of the complete mitotic history of the cell, including proliferation associated with normal development and tissue maintenance, plus additional cell turnover occurring as a consequence of environmental insults.
HM450 datasets showed that diverse hematopoietic cell types had a significant association between donor age and degree of hypomethylation, with the myeloid lineage (Fig. 6e) having a much slower rate of age-associated loss than the lymphoid lineage (Fig. 6f). This finding is consistent with the overall lower degree of methylation observed in myeloid cell types from WGBS data. While the rate of loss within the myeloid lineage was extremely low, the association with donor age was significant within the large human monocyte dataset (Fig. 6e). This finding contradicts an earlier analysis based on many of the same samples, which found that monocytes lacked PMD hypomethylation and age-associated hypomethylation24.
PMD hypomethylation and mitotic cell division in cancer
We studied the landscape of cancer hypomethylation in 9,072 tumors from 33 cancer types included in TCGA, using the HM450 solo-WCGWs located within common PMDs (Fig. 7a). PMD hypomethylation was nearly universal but showed extensive variation both within and across cancer types. Comparison to 749 adjacent normal samples from TCGA showed that the relative degree of hypomethylation across cancer types was correlated with that of the disease-free tissue of origin (Supplementary Figs. 11–13). This association was reduced in cancer types for which the normal adjacent specimens contained low fractions of relevant cell types representing putative cells of origin for the tumor.
a, PMD – HMD solo-WCGW methylation difference for 9,072 tumors from TCGA HM450 data. Each sample is ordered within a cancer type by the PMD – HMD difference, and cancer types are ordered by average PMD – HMD difference. b, PMD methylation (x axis) versus somatic mutation density (y axis) for all 3,959 high-purity TCGA cases (purity ≥0.7), with Spearman’s ρ indicated. The blue line represents the regression line for all samples, while the red regression line excludes ‘hypermutator’ samples (Methods). c, Density of somatic LINE-1 insertions (violin plot elements) in non-overlapping 1-Mb genomic bins (n = 3,053), stratified by the percentage of the bin overlapping common PMDs (only cases with whole-genome sequencing are included). d, PMD methylation (x axis) versus LINE-1 insertion counts (y axis) for nine TCGA cancer types having substantial LINE-1 insertion counts. *P < 0.05, **P ≤ 0.01, Spearman’s test significance. e, The ten most significantly enriched Gene Ontology (GO) terms for the 60 genes whose expression was most strongly correlated with PMD hypomethylation in TCGA tumors, showing fold enrichment (gray) and false discovery rate (olive). f, Gene set enrichment analysis (GSEA) for 350 cell cycle–dependent genes from Cyclebase44, ranking all genes according to degree of correlation between expression and PMD hypomethylation. The y axis shows the enrichment score for gene set S (ES (S)). g, Normalized expression (z score) of cell cycle–dependent genes from Cyclebase (categorized by cell cycle phase) in 3,414 high-purity TCGA tumor samples (purity ≥0.7), ordered by PMD – HMD methylation difference.
Somatic mutation events are known to display mitotic clock-like properties39. Within TCGA tumors, we found that higher genome-wide somatic mutation densities were significantly associated with deeper PMD hypomethylation, suggesting that mitotic turnover may underlie both somatic mutation and PMD hypomethylation (Fig. 7b). This association was consistent using different purity thresholds (Supplementary Fig. 13c), indicating that it was not the result of confounding due to differential detection sensitivity related to purity.
PMD hypomethylation was also associated with somatic copy number aberration density (Supplementary Fig. 13d). Activation and insertion of LINE-1 endogenous retrotransposable elements represents a common event in human cancer and can result in structural alterations, copy number alterations, and induction of oncogenes36,40,41. Using somatic LINE-1 insertions identified from whole-genome sequencing of TCGA tumors36, we found that LINE-1 insertion breakpoints were preferentially enriched in PMD regions (Fig. 7c), in agreement with an earlier study40. Tumors with deeper PMD hypomethylation had more LINE-1 insertions in eight of nine cancer types, with the only exception being endometrial cancer (Fig. 7d and Supplementary Fig. 14). While the mechanisms controlling LINE-1 insertion density in cancer are not well understood, they may be stochastically linked to the number of cell divisions (like single-nucleotide variants (SNVs)) and/or require derepression of ‘hot’ LINE-1 elements, a process that may be linked to DNA hypomethylation42,43.
We reasoned that tumors highly proliferative at the time of specimen collection may also reflect an extensive history of past cell division. Using TCGA samples with matched gene expression data, we identified the 60 genes whose expression was most strongly associated with PMD hypomethylation, finding that these genes were most enriched in Gene Ontology functional terms associated with proliferation and mitotic cell division (Fig. 7e). In further support of this link between ongoing cell proliferation and PMD hypomethylation, the genes with the greatest association with PMD hypomethylation were strongly enriched within a list of 350 cell cycle–dependent genes from Cyclebase44 (Fig. 7f). Ranking tumor samples by their degree of PMD hypomethylation showed that this association involved most cell cycle–dependent genes across different mitotic stages (Fig. 7g). Proliferative tumors had deep PMD hypomethylation despite having higher levels of DNMT1, DNMT3A, and DNMT3B, which are expressed as part of a general DNA replication program (Supplementary Note). The most hypomethylated tumors also had high expression of UHRF1 (a contributor to DNMT1 methylation maintenance activity), underscoring the idea that PMD hypomethylation accumulates despite strong expression of the DNA methylation maintenance machinery. We also investigated whether overexpression of TET genes, which participate in active DNA demethylation, might contribute to PMD hypomethylation. None of the three TET genes had the highest expression in the tumors with strongly hypomethylated PMDs, indicating that TET enzymes are not responsible for DNA methylation loss in PMD regions (in contrast to promoters and CpG islands, where extensive evidence exists for TET-mediated demethylation). All of our tumor mutation and expression results suggest cumulative mitotic cell divisions as the major driving force behind accumulation of PMD hypomethylation.
Replication timing and H3K36me3 both affect methylation
We used the one cell type with publicly available data for all relevant histone and topological marks, IMR90, to systematically analyze our solo-WCGW-based PMD definition. This analysis confirmed previous findings6,7 that HMD–PMD structure coincides with nuclear architecture, as characterized by Hi-C A and B compartments, lamin B1 distribution, and replication timing (Fig. 8a). At the single-CpG scale, solo-WCGW CpG methylation was most strongly correlated with replication timing, followed by the histone mark H3K36me3 (Supplementary Fig. 15a). The de novo methyltransferase DNMT3B has recently been shown to be guided to transcribed gene bodies via a direct interaction with the H3K36 methylation mark45. Active genes marked by H3K36me3 are overwhelmingly located in early-replicating regions, and it has been suggested that both active transcription of gene bodies and early replication timing contribute to differential methylation throughout the genome9. To disentangle the contributions of H3K36me3 and replication timing to genome-wide DNA methylation levels and PMDs, we performed a stratified analysis of all solo-WCGW CpGs in the genome (Fig. 8b,c). We found that the 14% of solo-WCGWs overlapping H3K36me3 were highly methylated, irrespective of position relative to gene annotations or replication timing (Fig. 8b, left). The remaining 86% of solo-WCGWs (those not overlapping an H3K36me3 peak) had lower methylation across all contexts, but were strongly dependent on replication timing (Fig. 8b, right). In IMR90 cells, the degree of methylation maintenance associated with early replication timing was even greater than the degree associated with H3K36me3 (Fig. 8b, right). The relative contribution of replication timing versus H3K36me3 was reversed in the H1 (hESC) cell line (Fig. 8c), a cell type with exceptionally high DNMT3A and DNMT3B activity that makes it one of the few cell types able to survive loss of DNMT1 function46,47. Because most somatic cell types had detectably hypomethylated PMDs like IMR90 (and unlike H1), our observations support a model in which highly effective methylation maintenance at H3K36me3-marked regions is achieved through a process mediated by the direct recruitment of DNMT3B through its PWWP domain45. Consistent with earlier observations9, this H3K36me3-linked maintenance appears to act independently of the effect of replication timing on PMD methylation loss (Fig. 8d).
a, Multiscale plot of chromosome 16p showing similarity between solo-WCGW methylation and other chromatin marks in the IMR90 fibroblast cell line. b, Average methylation level of all genomic solo-WCGWs in IMR90 fibroblasts, stratified by (i) overlap with H3K36me3 peaks (left versus right), (ii) context relative to gene annotations (“genic” versus “intergenic”), and (iii) Repli-seq replication timing bin (red, yellow, light blue, dark blue). For solo-WCGWs residing within ±10 kb of an annotated gene (genic), meta-gene plots show methylation averages in relation to the transcription start site (TSS) and the transcription termination site (TTS). For all other solo-WCGWs (intergenic), each replication timing group is shown as a single violin plot. c, Data plotted as in b for the H1 hESC cell line (using Repli-chip rather than Repli-seq data). d, Schematic summary, showing solo-WCGW CpG methylation loss primarily determined by replication timing domain but locally protected by H3K36me3. e, Schematic model illustrating DNMT1 processivity favoring dense CpGs and leading to incomplete remethylation of solo CpGs. f, Schematic illustration of the remethylation timing model where genomic regions synthesized earlier in S phase (HMDs) spend more time exposed to methylation maintenance machinery and thus show more complete methylation maintenance than PMDs. g, Illustration of the relationship between major determinants of hypomethylation and 3D nuclear topology, with LADs occupying a distinct heterochromatic nuclear compartment.
Discussion
In this study, we identified four distinct features influencing DNA methylation levels in large portions of the human and mouse genomes: first, the local sequence context of the CpG dinucleotide; second, the timing of DNA replication; third, the presence of the H3K36me3 histone mark; and fourth, the accumulated number of cell divisions. The sequence context, replication timing, and H3K36me3 marks each confer differential susceptibility to replication-associated DNA methylation loss and thus collectively shape PMD–HMD structure, while the degree of PMD hypomethylation is a function of the cumulative number of cell divisions from the earliest stages of embryonic development.
We showed that two local sequence features (CpG density and the WCGW sequence context) exert a strong influence on the rate of DNA methylation loss at individual CpGs within PMDs and that these influences are consistent across cell types and species. The bulk of DNA methylation maintenance is performed by DNMT1 and augmented by DNMT3A and DNMT3B48. DNMT1 has been shown to act processively, with increased efficiency in the presence of multiple CpG sites in close proximity49, a feature consistent with the poorer methylation maintenance of ‘solo’ CpGs (Fig. 8e). In vitro biochemical studies have yielded conflicting findings regarding the role of the immediate CpG-flanking positions on DNMT1 activity, with one study suggesting higher affinity for GC-rich flanking sequences50 and another suggesting higher affinity for AT-rich sequences51. The in vivo effects of the WCGW motif described here on methylation maintenance efficiency should be followed up with careful mechanistic studies to identify the causative factor or factors. The discovery of the solo-WCGW signature largely allowed for our improved analysis of HMD–PMD structure, which may lead to better characterization of not just the ‘common PMDs’ studied here but also important classes of cell-type-specific PMDs6,7,14,52 (Supplementary Note).
Most solo-WCGW CpGs were not marked by H3K36me3, and we identified replication timing as the major determinant for methylation levels at these H3K36me3-negative CpGs. We propose that replication late in S phase provides the cell with less time for remethylation of newly synthesized daughter strands during DNA replication (Fig. 8f). This remethylation window model is supported by a recent study that reconstructed methylation gains and losses at individual CpGs upon clonal expansions of individual somatic cells in culture21, showing that progressive methylation loss was most pronounced at late-replicating domains. Further strengthening the remethylation window model, biochemical studies have shown that remethylation during mitosis is in fact relatively slow and not fully completed until after the S–G2 checkpoint53,54. Therefore, remethylation efficiency is likely dependent on the time window between daughter strand synthesis and the beginning of M phase. This is consistent with the mitotic clock-like PMD methylation loss we observe specifically within late-replicating regions (Fig. 8f).
The presence of H3K36me3 appeared to override this late-replication-associated methylation loss at solo-WCGW CpGs (Fig. 8d). Genetic evidence suggests that maintenance of DNA methylation at H3K36me3-marked CpGs is mediated by the direct recruitment of DNMT3B to H3K36me3-marked nucleosomes45,55. The independent contributions of replication timing and H3K36me3 are consistent with earlier findings based on actively transcribed gene bodies9 and help to resolve the long-standing paradox concerning positive associations between actively transcribed gene bodies and DNA methylation56. This would also explain why head and neck squamous cell carcinomas with NSD1 mutations, which exhibit significant reductions in H3K36me2 and H3K36me3 levels57, have substantial loss of DNA methylation in the HMD compartment (Supplementary Fig. 15b). It is important to note that the two major genomic contexts we found to contribute to hypomethylation are strongly associated with specific nuclear territories (Fig. 8g). As the heterochromatin likely represents a distinct compartment separated by a physical boundary, we cannot rule out the possibility of other compositional differences of this compartment contributing to the less efficient DNA methylation maintenance observed there.
A number of studies have identified specific CpGs predictive of chronological age58,59,60 as well as gestation age at birth61. These signatures are largely non-overlapping with PMDs, as shown in earlier work26, and with the PMD solo-WCGW CpGs identified here. We believe that this is because PMD hypomethylation captures underlying mitotic dynamics, which are only loosely associated with chronological age per se. Organismal aging and the associated physiological changes affect transcriptional regulation of various genes and pathways, and many or most of the loci identified on the basis of age alone58,59,60 likely represent transcriptionally coupled chromatin changes at these genes (for example, changes to somatostatin, which regulated growth hormone58). As we suggest, PMD hypomethylation is likely a more direct clock-like readout of mitotic age, which is generally correlated with chronological age but can be accelerated by environmental factors or processes that promote cell turnover, such as cellular damage, wounding, inflammation, etc.
DNA hypomethylation has long been proposed to allow for aberrant expression and transposition of retroelements, which can play a role in cancer by inducing chromosomal aberrations at the point of insertion62,63,64,65,66. Genetically engineered Dnmt1 hypomorphism in mouse was shown to cause lymphomas frequently harboring retrotranspon-induced Notch1 activation events43. Whole-genome sequencing has shown that approximately 50% of human tumors contain somatic retrotranspositions of LINE-1 elements and that these often lead to structural alterations40,41,67,68 enriched within PMDs40. In one study, human lung tumors exhibiting mobilization of LINE-1 elements shared a common DNA hypomethylation signature42. Across a large TCGA cohort, we showed that tumors with higher degrees of PMD hypomethylation are more likely to have LINE-1 insertions (Fig. 7c,d). This evidence is correlative in nature, and it is certainly possible that LINE-1 activity is caused by a methylation-independent event. However, our results are consistent with previous genetic models, and it is tempting to hypothesize that LINE-1 activity could be accelerated by PMD hypomethylation.
The methylation loss process described here affects a sizeable fraction of all CpGs in the genome and thus could exert a significant influence on methylation-dependent mutational processes, most importantly CpG-to-TpG substitutions driven by methylation-dependent deamination of CpGs. This mutational signature accounts for a large fraction of the single-nucleotide mutations observed in both evolution and cancer, and thus we might expect systematic DNA methylation changes to influence the rate of these mutations. A simplistic model would predict that hypomethylated solo-WCGW CpGs within late-replicating PMDs would be protected from deamination and thus have a lower CpG-to-TpG mutation rate. Indeed, we observed some evidence in support of this model for both somatic mutations (from tumor sequencing) and de novo mutations in the human germ line (from whole-genome trio sequencing). This analysis, described in detail in Supplementary Fig. 16 and the Supplementary Note, does not take into account other important factors such as the rate of transcription-coupled DNA repair and should be confirmed in future studies.
URLs. Roadmap Epigenomics data, ftp://ftp.ncbi.nlm.nih.gov/pub/geo/DATA/roadmapepigenomics/; BLUEPRINT epigenome project data, ftp://ftp.ebi.ac.uk/pub/databases/blueprint/; ENCODE data project, http://www.encodeproject.org/; BisSNP easy run procedure, http://people.csail.mit.edu/dnaase/bissnp2011/stepByStep.html; Picard tools,
http://broadinstitute.github.io/picard. Our entire customized workflow ECWorkflows is hosted and freely available at https://github.com/uec/ECWorkflows. Our annotations including the genomic coordinates of solo-WCGWs and PMD definitions are available at https://zwdzwd.github.io/pmd.
Methods
Whole-genome bisulfite sequencing
Cases for the WGBS assay were selected from eight of the most common cancer types (lung squamous cell carcinoma, lung adenocarcinoma, breast, colorectal, endometrial, stomach, bladder, glioblastoma). For at least one tumor from each cancer type, we also sequenced its adjacent histologically normal tissue; for the rest, only the tumor was profiled. We combined these samples with one tumor and matched normal colon cancer pair from our earlier study6, yielding a core set of 40 well-characterized tumors and 9 adjacent normal samples (Supplementary Table 1). These tumors and normal samples are referred to as core tumors and core normal samples in the text. The paired-end WGBS-PE protocol was adapted from earlier protocols developed by our group6. Briefly, sample genomic DNA (2 μg) was sonicated using a Diagenode Bioruptor and size selected to a range of 400–500 bp. Sodium bisulfite conversion of all DNA samples was performed using the EZ DNA Methylation kit (Zymo Research). All libraries were subjected to quality control by Agilent Bioanalyzer examination and quantified using the Kapa Biosystems kit. Cluster generation and paired-end sequencing were performed according to Illumina guidelines for the HiSeq 2000, using the latest version of reagents and software updates.
External data
The external human WGBS data consisted of 19 germ cells and preimplantation embryonic tissues, 13 postimplantation embryonic and fetal tissues, 37 cell lines, 59 non-blood normal primary tissues (including normal adjacent tissues of tumors as well as disease-free samples), 154 blood or blood component samples, 11 solid tumors, and 50 blood malignancies (Supplementary Table 1). The 206 mouse WGBS datasets were constituted by 13 embryonic stem cells, 17 germ cells and embryonic tissues, 123 primary fetal tissues, and 53 primary postnatal normal samples. Human postnatal normal samples were retrieved from Roadmap Epigenomics Project (see URLs). Sorted blood WGBS and blood malignancies were downloaded from the BLUEPRINT epigenome project (see URLs). Mouse fetal WGBS samples were downloaded from the ENCODE project (see URLs). Other postnatal and fetal WGBS samples were downloaded from MethBase27. For MethBase samples, we only included datasets that passed the quality control standard of the database. We list the relevant citations and sources of the WGBS datasets used in this work in Supplementary Table 1. HM450 datasets and the corresponding meta-information used for age association were obtained from Gene Expression Omnibus by downloading the following datasets: GSE30870, GSE35069, GSE56046, GSE59065, GSE51954, GSE61278, and GSE56515. Mutation prevalence for TCGA tumor samples was obtained from the Broad Institute TCGA Genome Data Analysis Center (2016): MutSigCV v0.9 cross-sample somatic mutation rate estimates (28 January 2016 release). Tumors that had POLE or APOBEC family mutations or were classified as having microsatellite instability were annotated to be hypermutator tumors. When hypermutator samples were excluded, samples without annotation were also excluded. Numbers of somatic LINE-1 insertions in 1-Mb bins were downloaded from an earlier report36.
Alignment and extraction of methylcytosine levels
Reads were aligned to the genome (build GRCh37) using BSmap71 under the following parameters: “-p 27 -s 16 -v 10 -q 2 -A AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTAC ACGACGCTCTTCCGATCT -A AGATCGGAA GAGCGTCGTGT AGGGAAAGAGTGTAGAT CTCGGTGGTCGCC GTATCATT”. We marked duplicated reads using Picard tools (version 1.38; see URLs). DNA methylation rates and SNP information were called using Bis-SNP72, with the default easy-run procedure (see URLs). Bis-SNP allows for C>T mutations to be distinguished from bisulfite conversion by investigating the complementary strand. CpGs with fewer than ten reads of coverage were excluded from analysis.
Genomic binning
To show megabase-scale HMD/PMD structures, we chose a 100-kb window size so that the segments would contain a sufficient number of solo-WCGWs to give reliable methylation averages (Supplementary Fig. 17 and Supplementary Note), without losing resolution to detect the majority of PMD positions, which fall within PMDs of 500 kb or greater6.
Definition of preliminary PMD/HMD domains based on all CpGs
We used WGBS at ~15× coverage to profile the methylation patterns of 40 tumors (39 new TCGA samples and one from our prior study6) from eight of the most common cancer types, and tumors were selected on the basis of high cancer cell content (Supplementary Table 1). For one case from each of the eight cancer types, we profiled both the tumor and adjacent normal tissue; for the rest, only the tumor was profiled. Most of our tumor samples had a high degree of hypomethylation, so we first used an existing HMM-based tool, MethPipe27, using a window size setting of 10 kb, to identify PMDs in each sample individually (Supplementary Fig. 1a). While the fraction of the genome covered by PMDs in different samples differed by two to threefold (Supplementary Fig. 1b), there was sufficient overlap to define a shared MethPipe PMD set of 417 PMDs (covering 13% of the genome) that was shared among at least 21 of the 30 tumors. As a comparison group, we defined a shared MethPipe HMD (highly methylated domain) set that was not covered by PMDs in any tumor sample and included 830 regions (covering 32% of the genome).
Final definition of PMDs/HMDs based on s.d. of solo-WCGW methylation
Every 100-kb bin was dichotomized as PMD or HMD using a Gaussian mixture model (implemented in the R package mixtools) based on cross-sample s.d. of β values from our core tumor samples (n = 40). The Gaussian mixture model assumes two subpopulations of 100-kb bins—those located in PMDs with higher cross-sample s.d. and those located in HMDs with lower cross-sample s.d. The final threshold of cross-sample s.d. for classifying PMDs from HMDs was determined to be 0.125. The more conservative sets of ‘common PMDs’ and ‘common HMDs’ were defined by the criteria s.d. > 0.15 and s.d. < 0.10, respectively. Overlap of the PMD boundaries of two samples was measured in the percentage of 100-kb bins identified as both in PMDs and in HMDs in the two samples, respectively. The mouse PMDs/HMDs were defined in the same way using 32 postnatal non-brain WGBS samples (Supplementary Table 1). The s.d. threshold for classifying PMDs from HMDs in mouse was determined to be 0.09. Genomic coordinates of solo-WCGW CpGs and our defined PMDs and HMDs are available (see URLs).
HM450 analysis
For TCGA HM450 datasets, raw IDATs were preprocessed by first applying background subtraction73 and then linear dye-bias correction matching the signal intensities of the two detection channels. Probe signals with detection P < 0.05, as well as probes overlapping common SNPs and putative repetitive elements that cause potential cross-hybridization, were then masked74. For external datasets where raw IDATs were unavailable, we used processed β values downloaded from GEO. On the basis of our WGBS analysis, we classified HM450 probes according to the number of neighboring CpGs and the tetranucleotide sequence context. Only probes targeting solo-WCGW CpGs were retained. We also removed probes falling into annotated CpG islands or that were unmethylated (β < 0.2) in at least 20 of the 749 matched normal tissue samples included in TCGA. This resulted in 6,214 probes in common PMDs and 9,040 probes in common HMDs. Four-letter acronyms for cancer types were taken following the official TCGA nomenclature. We used the difference in methylation between the mean methylation of solo-WCGW probes located in common PMDs and those in common HMDs to measure the degree of PMD-associated DNA hypomethylation in each sample. This method avoids confounding in the case of cancer types derived from globally demethylated cell types such as primordial germ cells (Supplementary Figs. 12 and 13).
Analysis of the IMR90 epigenome
Features were clustered using 1 – |ρ| as the distance, where ρ is the Spearman’s correlation coefficient. Centromeres were excluded from IMR90 analysis. IMR90 epigenome data were downloaded from the ENCODE project data center (accessions listed in Supplementary Table 1). Wavelet-transformed signals for replication timing were downloaded from GEO (GSM923447)75. Histone mark signal was quantified using the percentage of base overlaps of each window with gapped peaks downloaded from the Roadmap Epigenomics Consortium. Gene bodies were extracted from GENCODE transcript annotation version 26. Base overlap was used as the gene body signal. RNA-seq signal was the log2-transformed number of reads overlapping with each window using Bedtools76. Only the protein-coding gene annotation from the HAVANA team was used for the genic analysis in Fig. 8d. Intergenic regions excluded all transcript annotation from all sources. Solo-WCGW CpG lamin B1 ChIP and Hi-C data were downloaded from GEO under accessions GSE53331 and GSE35156, respectively.
Rescaling based on PMD methylation
We calculated the distribution of methylation values within common PMD 100-kb bins. We then trimmed the top and bottom 20% of this distribution for each sample, setting low values to 0 and high values to 1, and linearly rescaled all values between 20% and 80% to the range [0,1] (Fig. 2e). The same genomic region of chromosome 16p is visualized in Fig. 2f.
Stratified analysis of solo-WCGW CpGs in the genome
The solo-WCGW CpGs were first classified (Fig. 8b,c) by their overlap with H3K36me3 into H3K36me3-positive (left) and H3K36me3-negative (right) categories, then by relative position to gene structures and placement in one of the four replication timing bin quartiles (colors, with threshold ≤40, (40,60], (60,75], >75 for IMR90 Repli-Seq and ≤–0.5, (–0.5,0.4], (0.4,1.15], >1.15 for H1 Repli-ChIP). For solo-WCGWs residing within ±10 kb of an annotated gene, metagene plots (Fig. 8b,c) were used to show average methylation levels across all genes in relation to the transcription start site (TSS) and the transcription termination site (TTS). For all other solo-WCGWs (intergenic), we showed the distribution of methylation values together for each replication timing group as a single violin plot.
Statistics
Except for when described explicitly in the text, P values for two-group comparison were calculated using one-tailed Wilcoxon’s rank-sum test. Correlation coefficients were computed with Spearman’s method, with the exact P values calculated in R using algorithm AS 89, otherwise via asymptotic t approximation when exact computation was not feasible.
Life Sciences Reporting Summary
Further information on experimental design is available in the Life Sciences Reporting Summary.
Code availability
Our customized workflow for preprocessing WGBS sequencing data is freely accessible (see URLs).
Data availability
The WGBS data are available in the Genome Data Commons (GDC) under the TCGA project with IDs and file names listed in Supplementary Table 1.
References
Ehrlich, M. & Wang, R. Y. 5-Methylcytosine in eukaryotic DNA. Science 212, 1350–1357 (1981).
Feinberg, A. P. & Vogelstein, B. Hypomethylation distinguishes genes of some human cancers from their normal counterparts. Nature 301, 89–92 (1983).
Gama-Sosa, M. A. et al. The 5-methylcytosine content of DNA from human tumors. Nucleic Acids Res. 11, 6883–6894 (1983).
Goelz, S., Vogelstein, B. & Feinberg, A. Hypomethylation of DNA from benign and malignant human colon neoplasms. Science 228, 187–190 (1985).
Hansen, K. D. et al. Increased methylation variation in epigenetic domains across cancer types. Nat. Genet. 43, 768–775 (2011).
Berman, B. P. et al. Regions of focal DNA hypermethylation and long-range hypomethylation in colorectal cancer coincide with nuclear lamina–associated domains. Nat. Genet. 44, 40–46 (2011).
Fortin, J.-P. & Hansen, K. D. Reconstructing A/B compartments as revealed by Hi-C using long-range correlations in epigenetic data. Genome Biol. 16, 180 (2015).
Weber, M. et al. Chromosome-wide and promoter-specific analyses identify sites of differential DNA methylation in normal and transformed human cells. Nat. Genet. 37, 853–862 (2005).
Aran, D., Toperoff, G., Rosenberg, M. & Hellman, A. Replication timing–related and gene body–specific methylation of active human genes. Hum. Mol. Genet. 20, 670–680 (2011).
Bergman, Y. & Cedar, H. DNA methylation dynamics in health and disease. Nat. Struct. Mol. Biol. 20, 274–281 (2013).
Quante, T. & Bird, A. Do short, frequent DNA sequence motifs mould the epigenome? Nat. Rev. Mol. Cell Biol. 17, 257–262 (2016).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Timp, W. et al. Large hypomethylated blocks as a universal defining epigenetic alteration in human solid tumors. Genome Med. 6, 61 (2014).
Hovestadt, V. et al. Decoding the regulatory landscape of medulloblastoma using DNA methylation sequencing. Nature 510, 537–541 (2014).
Baylin, S. & Bestor, T. H. Altered methylation patterns in cancer cell genomes: cause or consequence? Cancer Cell 1, 299–305 (2002).
Brennan, K. & Flanagan, J. M. Is there a link between genome-wide hypomethylation in blood and cancer risk? Cancer Prev. Res. 5, 1345–1357 (2012).
Ehrlich, M. et al. Amount and distribution of 5-methylcytosine in human DNA from different types of tissues of cells. Nucleic Acids Res. 10, 2709–2721 (1982).
Lister, R. et al. Hotspots of aberrant epigenomic reprogramming in human induced pluripotent stem cells. Nature 471, 68–73 (2011).
Hansen, K. D. et al. Large-scale hypomethylated blocks associated with Epstein–Barr virus–induced B-cell immortalization. Genome Res. 24, 177–184 (2014).
Landan, G. et al. Epigenetic polymorphism and the stochastic formation of differentially methylated regions in normal and cancerous tissues. Nat. Genet. 44, 1207–1214 (2012).
Shipony, Z. et al. Dynamic and static maintenance of epigenetic memory in pluripotent and somatic cells. Nature 513, 115–119 (2014).
Schroeder, D. I. et al. The human placenta methylome. Proc. Natl Acad. Sci. USA 110, 6037–6042 (2013).
Kulis, M. et al. Whole-genome fingerprint of the DNA methylome during human B cell differentiation. Nat. Genet. 47, 746–756 (2015).
Durek, P. et al. Epigenomic profiling of human CD4+ T cells supports a linear differentiation model and highlights molecular regulators of memory development. Immunity 45, 1148–1161 (2016).
Schultz, M. D. et al. Human body epigenome maps reveal noncanonical DNA methylation variation. Nature 523, 212–216 (2015).
Vandiver, A. R. et al. Age and sun exposure-related widespread genomic blocks of hypomethylation in nonmalignant skin. Genome Biol. 16, 80 (2015).
Song, Q. et al. A reference methylome database and analysis pipeline to facilitate integrative and comparative epigenomics. PLoS One 8, e81148 (2013).
Edwards, J. R. et al. Chromatin and sequence features that define the fine and gross structure of genomic methylation patterns. Genome Res. 20, 972–980 (2010).
Gaidatzis, D. et al. DNA sequence explains seemingly disordered methylation levels in partially methylated domains of mammalian genomes. PLoS Genet. 10, e1004143 (2014).
Carter, S. L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
Farlik, M. et al. DNA methylation dynamics of human hematopoietic stem cell differentiation. Cell Stem Cell 19, 808–822 (2016).
Knijnenburg, T. A. et al. Multiscale representation of genomic signals. Nat. Methods 11, 689–694 (2014).
Guelen, L. et al. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453, 948–951 (2008).
Lister, R. et al. Global epigenomic reconfiguration during mammalian brain development. Science 341, 1237905 (2013).
Tomasetti, C. & Vogelstein, B. Variation in cancer risk among tissues can be explained by the number of stem cell divisions. Science 347, 78–81 (2015).
Rodriguez-Martin, B. et al. Pan-cancer analysis of whole genomes reveals driver rearrangements promoted by LINE-1 retrotransposition in human tumours. Preprint at bioRxiv https://doi.org/10.1101/179705 (2017).
Burnet, F. M. A modification of Jerne’s theory of antibody production using the concept of clonal selection. CA Cancer J. Clin. 26, 119–121 (1976).
Wu, H. & Zhang, Y. Reversing DNA methylation: mechanisms, genomics, and biological functions. Cell 156, 45–68 (2014).
Alexandrov, L. B. et al. Clock-like mutational processes in human somatic cells. Nat. Genet. 47, 1402–1407 (2015).
Lee, E. et al. Landscape of somatic retrotransposition in human cancers. Science 337, 967–971 (2012).
Tubio, J. M. C. et al. Extensive transduction of nonrepetitive DNA mediated by L1 retrotransposition in cancer genomes. Science 345, 1251343–1251343 (2014).
Iskow, R. C. et al. Natural mutagenesis of human genomes by endogenous retrotransposons. Cell 141, 1253–1261 (2010).
Howard, G., Eiges, R., Gaudet, F., Jaenisch, R. & Eden, A. Activation and transposition of endogenous retroviral elements in hypomethylation induced tumors in mice. Oncogene 27, 404–408 (2008).
Santos, A., Wernersson, R. & Jensen, L. J. Cyclebase 3.0: a multi-organism database on cell-cycle regulation and phenotypes. Nucleic Acids Res. 43, D1140–D1144 (2015).
Baubec, T. et al. Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation. Nature 520, 243–247 (2015).
Li, E., Bestor, T. H. & Jaenisch, R. Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell 69, 915–926 (1992).
Li, Z. et al. Distinct roles of DNMT1-dependent and DNMT1-independent methylation patterns in the genome of mouse embryonic stem cells. Genome Biol. 16, 115 (2015).
Jones, P. A. & Liang, G. Rethinking how DNA methylation patterns are maintained. Nat. Rev. Genet. 10, 805–811 (2009).
Hermann, A., Goyal, R. & Jeltsch, A. The Dnmt1 DNA-(cytosine-C5)-methyltransferase methylates DNA processively with high preference for hemimethylated target sites. J. Biol. Chem. 279, 48350–48359 (2004).
Flynn, J., Azzam, R. & Reich, N. DNA binding discrimination of the murine DNA cytosine-C5 methyltransferase. J. Mol. Biol. 279, 101–116 (1998).
Bashtrykov, P., Ragozin, S. & Jeltsch, A. Mechanistic details of the DNA recognition by the Dnmt1 DNA methyltransferase. FEBS Lett. 586, 1821–1823 (2012).
Johann, P. D. et al. Atypical teratoid/rhabdoid tumors are comprised of three epigenetic subgroups with distinct enhancer landscapes. Cancer Cell 29, 379–393 (2016).
Liang, G. et al. Cooperativity between DNA methyltransferases in the maintenance methylation of repetitive elements. Mol. Cell. Biol. 22, 480–491 (2002).
Schermelleh, L. et al. Dynamics of Dnmt1 interaction with the replication machinery and its role in postreplicative maintenance of DNA methylation. Nucleic Acids Res. 35, 4301–4312 (2007).
Neri, F. et al. Intragenic DNA methylation prevents spurious transcription initiation. Nature 543, 72–77 (2017).
Jones, P. A. The DNA methylation paradox. Trends Genet. 15, 34–37 (1999).
Papillon-Cavanagh, S. et al. Impaired H3K36 methylation defines a subset of head and neck squamous cell carcinomas. Nat. Genet. 49, 180–185 (2017).
Hannum, G. et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol. Cell 49, 359–367 (2013).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, R115 (2013).
Slieker, R. C. et al. Age-related accrual of methylomic variability is linked to fundamental ageing mechanisms. Genome Biol. 17, 191 (2016).
Knight, A. K. et al. An epigenetic clock for gestational age at birth based on blood methylation data. Genome Biol. 17, 206 (2016).
Walsh, C. P., Chaillet, J. R. & Bestor, T. H. Transcription of IAP endogenous retroviruses is constrained by cytosine methylation. Nat. Genet. 20, 116–117 (1998).
Bourc’his, D. & Bestor, T. H. Meiotic catastrophe and retrotransposon reactivation in male germ cells lacking Dnmt3L. Nature 431, 96–99 (2004).
Trinh, B. N., Long, T. I., Nickel, A. E., Shibata, D. & Laird, P. W. DNA methyltransferase deficiency modifies cancer susceptibility in mice lacking DNA mismatch repair. Mol. Cell. Biol. 22, 2906–2917 (2002).
Eden, A. Chromosomal instability and tumors promoted by DNA hypomethylation. Science 300, 455 (2003).
Ehrlich, M. DNA hypomethylation in cancer cells. Epigenomics 1, 239–259 (2009).
Solyom, S. et al. Pathogenic orphan transduction created by a nonreference LINE-1 retrotransposon. Hum. Mutat. 33, 369–371 (2012).
Helman, E. et al. Somatic retrotransposition in human cancer revealed by whole-genome and exome sequencing. Genome Res. 24, 1053–1063 (2014).
Amendola, M. & van Steensel, B. Nuclear lamins are not required for lamina-associated domain organization in mouse embryonic stem cells. EMBO Rep. 16, 610–617 (2015).
Hiratani, I. et al. Genome-wide dynamics of replication timing revealed by in vitro models of mouse embryogenesis. Genome Res. 20, 155–169 (2010).
Xi, Y. & Li, W. BSMAP: whole genome bisulfite sequence MAPping program. BMC Bioinformatics 10, 232 (2009).
Liu, Y., Siegmund, K. D., Laird, P. W. & Berman, B. P. Bis-SNP: combined DNA methylation and SNP calling for Bisulfite-seq data. Genome Biol. 13, R61 (2012).
Triche, T. J. Jr., Weisenberger, D. J., Van Den Berg, D., Laird, P. W. & Siegmund, K. D. Low-level processing of Illumina Infinium DNA Methylation BeadArrays. Nucleic Acids Res. 41, e90 (2013).
Zhou, W., Laird, P. W. & Shen, H. Comprehensive characterization, annotation and innovative use of Infinium DNA methylation BeadChip probes. Nucleic Acids Res. 45, e22 (2017).
Hansen, R. S. et al. Sequencing newly replicated DNA reveals widespread plasticity in human replication timing. Proc. Natl Acad. Sci. USA 107, 139–144 (2010).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Acknowledgements
We thank T. Hinoue and H. Noushmehr for help selecting TCGA WGBS samples based on analysis of other TCGA data types and H. Goodridge and D.-C. Lin for useful discussions and comments. We thank The Cancer Genome Atlas Research program office, especially K. Shaw, for helping to get the WGBS project up and running. We also thank the members of the TCGA Research Network along with the dozens of other research groups that generated the published datasets that were used here to gain new insights. This project was supported by the Van Andel Research Institute, the Cedars-Sinai Center for Bioinformatics and Functional Genomics and the Samuel Oschin Comprehensive Cancer Institute, and the University of Southern California USC Epigenome Center. The work was funded by the following grants: National Institutes of Health/National Cancer Institute grants U24 CA143882 (P.W.L., B.P.B., H.Q.D., and H.S.); R01 CA170550 (P.W.L.); U01 CA184826 (B.P.B.); U24 CA210969 (P.W.L., B.P.B., and H.S.), Ovarian Cancer Research Fund Grant 373933 (H.S.), and National Institutes of Health/National Human Genome Research Institute grant R01 HG006705 (B.P.B.).
Author information
Authors and Affiliations
Contributions
H.S., P.W.L., and B.P.B. conceived the study. C.M.N., P.W.L., and B.P.B. oversaw the data generation and data quality control, with assistance from D.J.W. Z.R. automated the next-generation sequencing analysis and quality control steps, and submission of data to NCI repositories. W.Z., H.Q.D., H.S., and B.P.B. performed computational analysis and produced figures. W.Z., H.S., P.W.L., and B.P.B. wrote the manuscript, with significant contributions from H.Q.D. H.S., P.W.L., and B.P.B. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–19 and Supplementary Note
Supplementary Table 1
Meta-information and sources of WGBS samples reported and analyzed
Rights and permissions
About this article
Cite this article
Zhou, W., Dinh, H.Q., Ramjan, Z. et al. DNA methylation loss in late-replicating domains is linked to mitotic cell division. Nat Genet 50, 591–602 (2018). https://doi.org/10.1038/s41588-018-0073-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0073-4
This article is cited by
-
The impact of DNA methylation on CTCF-mediated 3D genome organization
Nature Structural & Molecular Biology (2024)
-
DNMT3B PWWP mutations cause hypermethylation of heterochromatin
EMBO Reports (2024)
-
Biologically informed deep learning for explainable epigenetic clocks
Scientific Reports (2024)
-
Cell cycle gene alterations associate with a redistribution of mutation risk across chromosomal domains in human cancers
Nature Cancer (2024)
-
Metabologenomic characterization uncovers a clinically aggressive IDH mutant glioma subtype
Acta Neuropathologica (2024)