Abstract
Methylation at the 5 position of cytosine in DNA (5meC) is a key epigenetic mark in eukaryotes. Once introduced, 5meC can be maintained through DNA replication by the activity of ‘maintenance’ DNA methyltransferases (DNMTs). Despite their ancient origin, DNA methylation pathways differ widely across animals, such that 5meC is either confined to transcribed genes or lost altogether in several lineages. We used comparative epigenomics to investigate the evolution of DNA methylation. Although the model nematode Caenorhabditis elegans lacks DNA methylation, more basal nematodes retain cytosine DNA methylation, which is targeted to repeat loci. We found that DNA methylation coevolved with the DNA alkylation repair enzyme ALKB2 across eukaryotes. In addition, we found that DNMTs introduced the toxic lesion 3-methylcytosine into DNA both in vitro and in vivo. Alkylation damage is therefore intrinsically associated with DNMT activity, and this may promote the loss of DNA methylation in many species.
Similar content being viewed by others
Main
DNA methylation is an important regulatory mechanism in eukaryotes, with important functions such as transposable element (TE) silencing and gene regulation1. 5meC acts as an epigenetic modification, which, once introduced by de novo methyltransferases (DNMT3a and DNMT3b in mammals), can be maintained through cell division by the activity of maintenance methyltransferases (DNMT1 in mammals)2. Both de novo and maintenance methylation are conserved in many species across eukaryotes, including animals, plants and fungi3,4. Nevertheless, DNA methylation pathways evolve rapidly in multiple lineages. Levels of DNA methylation vary widely, with many insects displaying sparse DNA methylation that is confined to a subset of transcribed genes5,6,7,8,9. Moreover, in many species, including the model organisms Drosophila melanogaster, C. elegans and Saccharomyces cerevisiae, cytosine DNA methylation has been lost altogether5,10. The factors driving such rapid evolution of DNA methylation pathways and their targets remain unclear. We investigated the evolution of DNA methylation in the nematode phylum and more widely across eukaryotes. We found that DNA methylation coevolved with DNA repair pathways and with the ALKB2 alkylation repair system in particular. To explain this, we identified a hitherto unknown off-target effect of DNMTs, in which they introduce alkylation damage into DNA. Indeed, we found that DNMTs are the major endogenous source of the alkylation 3-methylcytosine (3meC) lesion in cells. We hypothesize that this toxic activity may act to promote the loss of DNA methylation altogether in multiple lineages.
Results
DNA methylation is conserved in basal nematodes
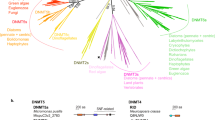
To study the factors driving the evolution of DNA methylation, we searched for cytosine DNMTs in nematode genomes across the phylum (Fig. 1a). We first used the Pfam core domain to identify potential cytosine DNMTs and then grouped these using phylogenetic analysis with known eukaryotic DNMTs. All of the identified nematode DNMTs are homologs of DNMT1, DNMT2 or DNMT3. DNMT2, which predominantly methylates tRNA3,4,11, is the most widespread among nematodes, but has been lost independently in some lineages, including Caenorhabditis, whereas it is conserved in the closely related parasitic nematode Nippostrongylus brasiliensis (clade V; Fig. 1b). Consistent with previous analyses of individual species12,13, we found that the cytosine DNMTs DNMT1 and DNMT3 have been retained in early-branching lineages, confirming that they are ancestral to nematodes. DNMT1 and DNMT3 were most likely lost completely in the common ancestor of the Rhabditida group that contains C. elegans (clades III–V; Fig. 1b). Notably, among the nematodes retaining cytosine DNMTs, some nematodes possess both DNMT1 and DNMT3 (Romanomermis culicivorax), whereas some species possess only DNMT1 (Plectus sambesii) or DNMT3 (Trichuris muris and Trichinella spiralis) (Fig. 1b,c). In species in which DNMT3 is the sole identified DNMT (T. spiralis or T. muris), this protein has not adopted any additional domains from DNMT1 (Fig. 1d,e).
a, Cladogram of nematodes, including the species profiled in this study. Clade nomenclature and phylogenetic positions are taken from ref. 33. b, The presence of DNMTs in nematodes as assessed by reciprocal BLAST. Gray indicates no best reciprocal BLAST hit. c, Multiple-sequence alignment showing key motifs important for DNMT activity in nematode DNMTs along with human DNMT1 and DNMT3 for comparison. d, Domains in DNMT1 and DNMT3 from human. e, Conservation of domains in nematode DNMTs as assessed by comparison to the Pfam seed of each domain. Domains from d that are in the Pfam database are shown, as are those found in at least one nematode DNMT. The N-terminal regions of DNMT1 from R. culicivorax and P. sambesii are missing, precluding definitive assessment of the presence or absence of CXXC; this is probably a result of incomplete genome assembly (Methods).
To investigate the effect of the presence of various combinations of cytosine methyltransferases, we measured the abundance of cytosine methylation (5meC) in genomic DNA using ultrasensitive liquid chromatography/mass spectrometry (LC/MS). 5meC was clearly detectable in all of the species containing DNMT1 or DNMT3 (Fig. 2a). We did not detect any 5meC in C. briggsae, which does not have DNMTs, and only detected very low levels in N. brasiliensis, which only has DNMT2. Notably, R. culicivorax, which has both DNMT1 and DNMT3, contained higher levels of genomic 5meC than the other nematodes.
a, Quantification of 5meC in DNA by LC/MS for different nematode species. Bar lines indicate the mean, and error bars represent s.d. Each overlayed point shows the mean of two technical repeats for n independent DNA extractions (n = 6, P. sambesii; n = 4, R. culicivorax, T. spiralis, T. muris, C. briggsae; n = 2, N. brasiliensis). n.d., not detected. b,c, The overall fraction of sites with >10% methylation for each of the specified CG-containing trinucleotides. The total number of CGs analyzed is presented in Supplementary Table 3. d, The average methylation of each CG (meCG) in different annotated regions, as compared with unmethylated spike-in. The total number of CGs analyzed is presented in Supplementary Note 3. FET, Fisher's exact test. e–h, Individual examples of repeat element consensus sequences with high levels of DNA methylation.
Nematode DNA methylation is enriched at transposable elements
To investigate the targeting of DNMT1 and DNMT3 to different genomic regions, we carried out whole-genome bisulfite sequencing (Supplementary Table 1). Consistent with our LC/MS analysis (Fig. 2a), we detected significant levels of DNA methylation above the bisulfite non-conversion rate, as estimated based on the inclusion of an unmethylated spike-in in the bisulfite reaction (P < 10–100, Fisher’s exact test; Fig. 2d), in all nematodes with DNMT1 or DNMT3; this methylation was significantly enriched at CG sites over non-CG sites (P < 10–100, Fisher’s exact test; Supplementary Fig. 1a). We did not observe significant differences in non-CG methylation between nematodes with just DNMT1 or DNMT3 (Supplementary Fig. 1a). Although we detected trace amounts of 5meC in N. brasiliensis possessing just DNMT2, our bisulfite analysis did not show any significant enrichment of 5meC above the non-conversion rate (Fig. 2d and Supplementary Fig. 1a), suggesting at most a very low and nonspecific activity of DNMT2 on cytosine in DNA, as has been observed in D. melanogaster10. Of note, the DNMT1-only methylome and the DNMT3-only methylomes showed different preferences for the nucleotide following the methylation (CG) site, normalized to the abundance of each trinucleotide in the genome. Comparison with bisulfite sequencing data from mouse embryonic stem cells (ESCs) lacking either DNMT1 or DNMT314 and from the arthropod Bombyx mori, which only has DNMT18, showed that the trinucleotide preferences of DNMT1 were highly similar between nematodes, mammalian cells and B. mori. In contrast, the DNMT3 preferences were different between nematodes and mammalian cells, suggesting differential conservation of DNA interactions for these two types of DNMTs (Fig. 2b,c).
Next, we annotated methylation sites across the entire genome. All of the nematodes with DNMT1 or DNMT3 showed significant enrichment of CG methylation above the genome-wide level (P < 10–5, Fisher’s exact test) normalized to CG content for at least one category of repetitive elements (Fig. 2d). In contrast, we did not observe enrichment in overall methylation at genes. Notably, this observation could not be explained by different trinucleotide composition within repeats, as trinucleotide content was similar across different repeat types (Supplementary Fig. 1b,c). C. briggsae (no DNMTs) and N. brasiliensis (DNMT2 only) showed no such enrichment (Fig. 2d). Notably, P. sambesii (DNMT1 only) showed marked enrichment for repeats over the genome-wide background (Fig. 2d). DNA methylation could be found across the entire body of many repetitive elements in all species (Fig. 2e–h and Supplementary Figs. 2 and 3) and, genome wide, elements with high levels of methylation were enriched for at least one category of repeats (Fig. 3a–d).
a–d, Histograms of methylation levels averaged across the body of different genomic features. e, Violin plots of methylation levels of all of the genes across the different species with the subset of genes carrying high levels of DNA methylation observed in R. culicivorax indicated by an arrow. The dot is at the median, the box shows interquartile range and the whiskers extend to the greatest point that is no more than 1.5 times the interquartile range.
We next examined DNA methylation in protein-coding genes. Analysis of the DNA methylation level across genes revealed that there were no notable populations of genes with higher levels of DNA methylation in P. sambesii (DNMT1 only), T. spiralis (DNMT3 only) or T. muris (DNMT3 only) (Fig. 3e and Supplementary Figs. 2 and 3). The few genes that showed appreciable DNA methylation in these species were likely misannotated repeats, as genes with homology to repeats had higher CG methylation than genes without homology (Supplementary Fig. 4). T. spiralis has been reported to show gene body methylation12; however, that study did not normalize for CG content. Given that CG density is markedly higher in the coding regions of all nematodes examined, this likely accounts for the discrepancy with our findings (Supplementary Fig. 5).
In R. culicivorax (DNMT1 and DNMT3), there was a bimodal distribution of DNA methylation across genes, with a small population of genes showing elevated DNA methylation (Fig. 3e). This finding is potentially reminiscent of gene body methylation in other invertebrates5,6,7,8,9. However, Gene Ontology (GO) analysis of the top 50 methylated genes with GO annotations revealed that ~14% were annotated as nucleic acid integration (enrichment P = 10–55 compared with all genes, chi-squared test with Benjamani and Hochberg (BH) multiple-test correction; Supplementary Tables 2 and 3); thus, even in R. culicivorax, at least some genes with high levels of methylation may be either misannotated TEs or genes with TE insertions.
Altogether, our analysis of DNA methylation across nematodes indicates that methylation of repeats is its most widely conserved function and was likely to have been present in the common ancestor of nematodes. Methylation in the bodies of transcribed protein-coding genes has been lost altogether in the lineage leading to T. spiralis and T. muris and in P. sambesii, and exists only in a minority of genes in R. culicivorax, and it is therefore not a conserved feature of DNA methylation in nematodes.
It has been argued that gene body methylation is a universal feature of DNMT1 and DNMT3 activity but that repeat-targeted cytosine methylation evolved independently in plants and vertebrates5,6. Our data are in accordance with a more nuanced view that the functions of DNA methylation evolve rapidly5,15 and that repeat-targeted DNA methylation is found in invertebrates16,17. Overall, the rapid evolution of both DNA methylation mechanisms and their targets in nematodes adds to the growing picture of the complex evolution of epigenetic mechanisms in animals5,15,18, in which the ancestral animals had a rich set of epigenetic mechanisms that have subsequently been lost independently in many descendent organisms.
DNA methylation coevolves with DNA alkylation damage repair across eukaryotes
What drives the rapid evolution of cytosine DNA methylation pathways in animals? One approach to this question is to identify genes coevolving with DNMTs, which may indicate pathways that are linked to the presence or absence of DNA methylation. We analyzed animal genomes in Ensembl (Release 28) and identified 133 human proteins that coevolved with DNMT1 or DNMT3 (P < 0.01, Fisher’s exact test after multiple-test correction; Supplementary Table 4). To our surprise, we found that the most strongly enriched GO term was for DNA repair (Fig. 4a and Supplementary Table 5). In particular, we noted the presence of alkylation repair enzymes among this set (Supplementary Fig. 6 and Supplementary Table 4), including the enzyme ALKB2 and its paralog ALKB3 (a mammalian-specific duplication; hereafter referred to as ALKB2/3). ALKB2/3 enzymes are members of the Fe2+-dependent oxygenase family of DNA repair enzymes homologous to Escherichia coli ALKB19. Whereas E. coli ALKB repairs a wide range of alkylated adducts, including protein, RNA and DNA, the family has diversified in eukaryotes, with different ALKB family enzymes specializing in the repair of particular substrates. Mammalian ALKB2/3 enzymes are the only members of the ALKB family that repair alkylation damage in DNA19,20 and are the only members that coevolved with the DNMTs DNMT1 and DNMT3 (Fig. 4c and Supplementary Fig. 7). To independently verify the association between DNA methylation and ALKB2/3, we carried out phylogenetic profiling of ALKB2/3 and DNMTs across the eukaryotic genomes in the Ensembl database (fungi, protozoa and animals) and tested for coevolution between ALKB2/3 and DNA methylation. Notably, in this analysis, we corrected for the over-representation of several closely related species in Ensembl (for example, the Drosophila genus, in which there are 12 species represented in Ensembl, all of which have no ALKB2/3 and no DNMT1 or DNMT3, or mammals, all of which have ALKB2/3 and DNMT1 and DNMT3) by ensuring that only one member from each lineage with the same profile of ALKB2/3 and DNMTs was included in the analysis (Fig. 4b and Supplementary Figs. 8–11; see Supplementary Tables 6–8 for the list of all of the species considered for the analysis). All three groups showed statistically significant co-occurrence between ALKB2/3 and the presence of at least one cytosine DNMT (DNMT1 and DNMT3) (P < 0.001 for fungi, P < 0.005 for animals, P < 0.01 for protazoa using Fisher’s exact test; Fig. 4b). In addition, in some fungi in which ALKB2/3 are present but DNMT1 is absent, DNMT5, which acts on CG sequences15, is conserved (Supplementary Figs. 8–11).
a, The top five most statistically significantly enriched GO terms in the set of genes that coevolved with DNMTs across animals. Terms associated with DNA repair are shown in red; others are shown in gray. b, The conservation of ALKB2 along with DNMTs across different taxonomic groups. Losses among n independent branches are shown (n = 20, animals; n = 31, fungi; n = 23, protists). c, The conservation of different members of the ALKB family in nematodes with and without DNA methylation.
We note that there are some potentially interesting exceptions to the general coevolution between ALKB2/3 and DNMTs, particularly in arthropods, where several species have lost ALKB2/3 while retaining DNMTs. To investigate this further, we compared genome-wide methylation levels across arthropods using previously published data from 18 insects9, the crustaceans Parhylale hawaiensis17, Daphnia pulex and Daphnia magna21, and the desert locust Schistocerca gregaria16. We found that species retaining ALKB2/3 had >10-fold higher median levels of DNA methylation than species that have lost ALKB2/3; this was true in both coding sequences and genome wide (P < 0.01, Wilcoxon unpaired test; Supplementary Fig. 12a,b and Supplementary Table 9).
DNMTs introduce 3meC alkylation damage into DNA
Overall, our analysis confirmed robust and widespread coevolution between ALKB2/3 and DNMTs across eukaryotes. On the basis of this observation, we wondered about a possible mechanistic link between DNA methylation and the presence of alkylation DNA damage. The preferred substrates for ALKB2/3 in DNA are 1-methyladenine (1meA) and 3meC22,23. We wondered whether the activity of cytosine DNMTs might be associated with the generation of 3meC in addition to these enzymes producing 5meC. To test this, we used synthetic nucleoside standards to develop an ultrasensitive mass spectroscopy (LC/MS) approach that enabled us to specifically distinguish between and quantify 3meC and 5meC in DNA (Fig. 5a,b, Methods and Supplementary Fig. 13a). To further verify this detection method, we treated a plasmid with the mutagen MMS, which, among other lesions, is known to introduce 3meC into DNA. The LC/MS analysis revealed a robust induction of 3meC, but no induction of 5meC (Supplementary Fig. 13b,c).
a,b, Validation of the method to detect 3meC specifically in the presence of 5meC using LC/MS. c,d, LC/MS measurement of 3meC introduced by the catalytic domain of DNMT3a in vitro compared with 3meC induction by the F646A mutant, which does not bind the cofactor SAM. Each of the three individual points for each sample shows the mean of two technical replicates for an independent in vitro reaction. Measurements below the signal-to-noise (s/n) ratio are shown in cyan. e, Example LC/MS traces for 3meC and 5meC for ESCs with or without DNMTs. Screenshots of the LC/MS analysis are shown. Colors for peaks are automatically assigned by the software on the basis of the peak settings. f, LC/MS analysis of 3meC in mouse ESCs with and without DNMTs and ALKBH2 (the Ensembl gene name of the mouse ALKB2 ortholog). The box plots show the interquartile range of 3meC normalized to dG, with a line at the median and whiskers extending to the furthest point within 95% of the range. Each of the three points for each cell line shows the mean of two technical replicates for independent DNA extractions. g, Model for how DNMTs influence methylation on different positions of cytosine.
To further examine the possible association between DNMTs and 3meC, we tested whether cytosine DNMT activity might be sufficient to produce DNA alkylation damage in vitro. We carried out in vitro methyltransferase reactions using the recombinant catalytic domain of DNMT3a. The subsequent LC/MS analysis identified the robust production of 5meC, as well as clear evidence for 3meC induction (Fig. 5c,d and Supplementary Fig. 13d,e). The induction of 3meC was far less abundant and occurred in the ratio 1:2,850 for 3meC:5meC, that is, 3meC = ~0.035% of 5meC (Fig. 5c,d). To verify that this result required the catalytic activity of DNMT3a, we expressed and purified the F646A point mutant of the catalytic domain of DNMT3a, which has a reduced ability to bind the cofactor SAM (Supplementary Fig. 14). Consistent with previous results24, we found that this enzyme had markedly reduced catalytic activity in introducing 5meC (Fig. 5c). Notably, this mutation also completely eliminated 3meC formation, demonstrating that catalytic activity is essential for DNMT3a to promote 3meC introduction (Fig. 5d). Taken together, these results suggest that DNMTs can use SAM to promote the introduction of 3meC at a low rate in addition to their usual 5meC product. Notably, the bacterial methyltransferase mSSSI also introduced 3meC in vitro (Supplementary Fig. 13c), suggesting that the introduction of 3meC may be a general property of cytosine methyltransferases. It is possible that the generation of 3meC involves a direct catalytic activity of the enzyme; alternatively, DNMTs may promote this indirectly by flipping the base out from the double helix25 and positioning it close to SAM.
To test whether DNMTs can promote introduction of 3meC in vivo, we used our LC/MS method to examine 3meC levels in mouse ESCs carrying DNMT1, DNMT3a and DNMT3b deletions (triple knockout, TKO)26. In wild-type (WT) mouse ESCs, we detected a clear signal for 3meC. Notably, the measured 3meC level was around tenfold lower than the level measured in vitro (Fig. 5c–f), consistent with the existence of endogenous DNA repair mechanisms capable of removing 3meC (Fig. 5g). In contrast, we were not able to detect any 3meC in TKO cells (P = 0.0017, ANOVA; Fig. 5e,f). As an independent validation, dot blots using an antibody specific for 3meC showed similar data (Supplementary Fig. 14a,b). We therefore conclude that the presence of active DNMT1 and DNMT3a/b is clearly associated with increased levels of 3meC in genomic DNA.
Mammalian ALKB2/3 enzymes have been shown to repair 3meC in vitro and in cultured mouse cells20, 22, 23. To test whether 3meC induced by DNMT activity is processed by ALKB2 in mouse ESCs, we used the CRISPR–Cas9 system to target deletions to the first exon of Alkb2 in both WT and TKO cells (Supplementary Fig. 15a). We obtained clones with homozygous deletions in both alleles of Alkb2, which showed a reduction in ALKB2 protein, in both WT and TKO cells (Supplementary Fig. 15b,c). Moreover, these clones showed increased sensitivity to the mutagen MMS relative to their parent line (P = 0.042, ANOVA test for ALKB deletion; Supplementary Fig. 15d), consistent with disruption of ALKB2 function in repairing alkylation DNA damage. We next analyzed 3meC levels and found that the loss of ALKB2 led to a ~15% increase in steady-state 3meC levels (P = 0.02, ANOVA test for ALKB2 deletion; Fig. 5f), implicating ALKB2 in the removal of 3meC. Notably, in TKO cells, even the lack of ALKB2 did not raise the level of 3meC above the detection limit of our LC/MS quantification (Fig. 5f). Overall, these data are consistent with ALKB2 being involved in the removal of 3meC associated with the activity of DNMTs in vivo.
The presence of 5meC in DNA is known to be mutagenic as a result of the deamination of 5meC to thymine, resulting in the depletion of CG dinucleotides over evolutionary time27, 28. 5meC-to-thymine deamination results in a G-T mismatch. However, alkylation damage such as 3meC poses a much more severe threat, as 3meC blocks the DNA polymerases involved in normal DNA replication29,30. Thus, our finding that 3meC is produced by DNMTs indicates that DNMT activity may directly cause replication stress in cells. On the basis of the average GC composition of the mouse genome, we calculated that the level of 3meC that we observe in vivo corresponds to approximately five modified cytosines in every 106 base pairs. The most common form of endogenous DNA damage known is the formation of abasic sites through cytosine deamination and subsequent uracil excision, as well as spontaneous depurination20. This form of DNA damage has a marked effect on shaping nucleotide frequencies through evolutionary processes28. Abasic sites have been measured in cultured cells and tissues, with estimates ranging from 1–20 nucleotides per 106 base pairs31. Our results indicate that 3meC, introduced by the off-target activity of DNMTs, exists at similar levels as abasic sites and is therefore one of the most abundant forms of spontaneous DNA damage in cells.
Discussion
Our results reveal that DNA methylation is a rapidly evolving epigenetic system. We found that, although C. elegans and other nematodes lost their DNA methylation system, other nematode species contain combinations of DNMTs homologous to the mammalian DNMT1 and DNMT3 enzymes that install genomic DNA methylation in these species. Furthermore, we found that, at least in nematodes, DNA methylation is primarily targeted to repetitive elements in the genome.
Notably, our evolutionary analysis of DNA methylation highlights an unexpected coevolution between DNA methylation and DNA repair systems. Our data indicate that DNMT activity is associated with the generation of 3meC both in vitro and in vivo and that ALKB2 demethylase is required to process this type of alkylation damage. We suggest that the relatively high level of endogenous DNA damage introduced by this off-target activity of DNMTs explains why ALKB2/3 enzymes are generally needed in organisms with 5meC (Fig. 5g). Even in the presence of ALKB2/3, 3meC introduction by DNMTs is likely to pose a threat to genome stability by causing DNA polymerases to stall, leading to the appearance of double-strand DNA breaks. Consistent with this possibility, members of the BRCA complex and RAD18, both of which are important in DNA double-strand break repair32, coevolved with DNMTs (Supplementary Fig. 6 and Supplementary Table 5).
Although future investigation into the relationship between DNA methylation and DNA repair may identify additional mechanistic links, our data indicate that the propensity of cytosine DNMTs to induce alkylation damage may be an important factor explaining the frequent independent losses of DNA methylation across different animal groups. Our data provide an important example of how analysis of the evolutionary relationships between proteins can identify previously unknown biochemical mechanisms.
Methods
Nematode collection and DNA isolation
R. culicivorax adults were a gift from C. Kraus (University of Köln) and derived from the culture of E. Platzer (University of California, Riverside). T. spiralis animals were prepared according to standard methodology. T. muris adults were collected using fine forceps from the ceca of SCID mice orally infected 42 d previously with 400 embryonated eggs.
P. sambesii animals were grown on low-salt agar with semiliquid HB101 at 25 °C. Adults were isolated from mixed-stage cultures by sorting on a COPAS large-particle sorter.
Analysis of DNA methylation sequencing
We assembled a draft P. sambesii genome from Illumina short-read sequencing (see below). Other genomes were taken from Wormbase (C. briggsae, WS240), Wormbase Parasite (N. brasiliensis, T. spiralis, T. muris WBPS4) or Nembase (R. culicivorax). Libraries for bisulfite sequencing were prepared using the Pico Methyl-Seq kit (Zymo Research). Bisulfite sequencing reads were mapped using Bismark, using the bowtie2 option. To obtain the methylation levels for different CG contexts and for different categories of genomic annotation (Fig. 2), we used the Bismark methylation extractor module to convert Bismark alignments into genome-wide coverage files reporting the methylation status. As the Bismark methylation extractor can only operate on a small number of contigs, before alignments, we had to artificially condense all genomes except that of C. briggsae (which is already assembled into 6 chromosomes) into ten ‘pseudo-contigs’ without disrupting the sequence of the contigs themselves. Subsequently, we selected cytosines covered by at least ten reads using a custom Perl script for further analysis. We converted genome coordinates from pseudo-contigs back to the original contigs and annotated individual CG sites according to gene predictions, either our own (P. sambesii) or those taken from Wormbase (WBPS4; WS245), and repeats were annotated using RepeatMasker using the parameters --no-low, --no-is, --species animalia, using Bedtools34. We then obtained percentage methylation by summing the methylated reads and the unmethylated reads across all CG sites within different regions.
Statistical enrichment of CG methylation was calculated using the Fisher’s exact test comparing the number of methylated sites and the number of unmethylated sites in both of the genomic regions of interest (that is, genes versus the entire genome).
To analyze the distribution across genes and TEs in more detail (Figs. 2d and 3), we again used Bismark to align DNA methylation sequencing data to contigs directly to avoid artifacts potentially caused by joining contigs together in the middle of repeats or genes. We then used MethylExtract to obtain site-specific methylation information and converted the output to bed files using a custom Perl script. Bedtools was used to annotate CG sites as above, and the mean methylation across individual features (for example, repeat, gene, etc.) was calculated by averaging across the fractional methylation at each site with >10 reads of coverage within the feature. Features with ≥5 CGs covered were used to draw Figs. 2d and 3. All statistical analyses and graphics generation were performed in the R environment.
Identification of DNMTs in nematode genomes
We searched the predicted proteins from nematode genomes for cytosine methyltransferase domains using Pfam hmm-search with the cytosine-5-methyltransferase domain. All proteins with matches to this domain were extracted. We then used BLAST to compare these against human DNMT1, DNMT2 and DNMT3 to annotate potential methyltransferases. Any proteins that did not match to DNMT1, DNMT2 or DNMT3 were tested against the Uniprot database; this identified them as bacterial contaminants, and they were removed from further analysis. We verified these annotations by phylogenetic analysis: nematode DNMTs along with selected DNMTs from other animals were aligned using MUSCLE, and these alignments were used to construct a phylogenetic tree according to the workflow in Phylogeny.fr. Domains within the nematode DNMTs were identified using Pfam searches with the seeds for PWWP, BAH and the cytosine DNMT domain. We could not find clear evidence for the CXXC domain in any of the nematode DNMTs, but this could be because of poor assembly of the genome in the N terminus of the protein in R. culicivorax and P. sambesii, as in both of these this region falls near to the boundary of contigs.
Coevolution analysis
We used BLAST on ALKB1–ALKB8 to analyze the conservation of ALKB proteins across the nematodes. The e-value of the best hit was tabulated. To identify coevolving proteins across animals in Ensembl, we downloaded each predicted proteome from Ensembl (release 28). We ran LBASTP using the human proteome as the query sequence and each predicted proteome as a database, retaining the best BLAST hit. Proteins with a BLAST hit log10 (e-value) less than –20 were given a score of 1 and those with a score greater than –20 were given a score of 0 to build a binary conservation matrix. We then used a Fisher’s exact test to identify proteins with a significant tendency to be lost or gained with DNMT3 or DNMT1 using the Benjamini and Hochberg multiple-test correction. Gene ontology information for the entire human Uniprot database was downloaded using BiomaRt, and significantly enriched categories were identified using a Fisher’s exact test following multiple-test correction. To test further for coevolution between presence of ALKB2/3 and DNMTs, we used a modified phylogenetic profiling method. We first used reciprocal BLAST to test for the presence of ALKB2/3 (retaining any hit that was reciprocal to ALKB2 or ALKB3, including examples where the best BLAST hit for ALKB2 was ALKB3 and vice versa), DNMT1 and DNMT3 in all animal, fungal and protozoan genomes downloaded from Ensembl. To ensure we retained data only for phylogenetically independent loss events, we constructed phylogenetic trees for these groups using the references detailed in the supplementary information (Supplementary Note 2). Finally, we mapped loss of ALKB2, DNMT1 and DNMT3 and collated these for each group before testing for co-occurrence of ALKB2 and one or more of DNMT1 or DNMT3 using Fisher’s exact test.
Analysis of DNA methylation levels across arthropods
For all species except S. gregaria, we obtained estimates of mCG/CG genome wide and at coding sequences directly from the relevant references9,17,21. For S. gregaria, the published reference16 did not report a genome-wide mCG/CG estimate as only coding sequences have been sequenced fully in this organism; thus, we used the FastMC algorithm9 to estimate genome-wide mCG/CG directly from raw sequencing data and calculated the coding sequence mCG/CG methylation level directly from the reference. We searched for conservation of ALKB2 in these species using the reciprocal BLAST method described above.
Dot blot analysis of methylation in genomic DNA samples
DNA was extracted using the Qiagen DNA Blood/Tissue isolation kit and redissolved in distilled water. DNA was diluted 50:50 with freshly prepared 0.2 M NaOH and heated for 5 min at 95 °C to denature. 2 μl of DNA was then spotted onto a nitrocellulose membrane and air-dried before cross-linking with a Stratalinker. The membrane was blocked with 5% milk in Tris-buffered saline (TBS). Anti-3meC (Active Motif) used at a 1:5,000 dilution or anti-5meC (clone 33D10, Abcam or Active Motif) used at a 1:2,500 dilution was added for an overnight incubation in 1% milk in TBS with 0.1% Tween-20. The membrane was washed and exposed to appropriate secondary antibody for 2 h at room temperature (20 °C) before developing with ECL.
A positive control for 3meC was prepared by incubating poly(dI:dC) in the presence of 20 mM MMS (Sigma) for 4 h at 37 °C. Excess MMS was quenched by addition of 0.2 M NaOH before dot blot analysis.
Positive and negative controls for 5meC, PCR products from the APC promoter made either with 5meCTP or CTP, were purchased from Active Motif.
LC/MS
N3-methyl-2′-deoxycytidine (3meC) standards were purchased from ChemGenes; 2′-deoxycytidine (dC) and 2′-deoxyguanosine (dG) were purchased from Berry and Associates; and C5-methyl-2′-deoxycytidine (5meC) was purchased from CarboSynth. Genomic DNA or synthetic oligonucleotides were digested to nucleosides for a minimum of 9 h at 37 °C using a digestion enzymatic mix (a kind gift from NEB). All samples and standard curve points were spiked with a similar amount of isotope-labeled synthetic nucleosides: 100 fmol of dC* and dG* purchased from Silantes and 5 fmol of 5meC* obtained from T. Carell (Center for Integrated Protein Science at the Department of Chemistry, Ludwig-Maximilians-Universität München). The nucleosides were separated on an Agilent RRHD Eclipse Plus C18 2.1 × 100 mm, 1.8 m column by using the HPLC 1290 system (Agilent) and analyzed using the Agilent 6490 triple-quadrupole mass spectrometer. Quantification was carried out in multiple-reaction monitoring mode (MRM) by monitoring the specific transition pairs of m/z 250.1/134.1 for dC, 290.1/174.1 for dG, 264.1/148.1 for 5mC and 242.2/95.1 for 3meC. To calculate the concentrations of individual nucleosides (for dC, dG and 5meC), standard curves representing the ratio of the peak response of known amounts of synthetic nucleosides and the peak response of the isotope-labeled nucleosides were generated and used to convert the peak-area values to corresponding concentrations. For 3meC, the concentrations were calculated directly using a standard curve with light nucleosides. The threshold for peak detection was a signal-to-noise ratio (calculated with a peak-to-peak method) above 10, and the limit of quantification (LOQ) was 25 amol for 5mC and 50 amol for 3mC. Final measurements were normalized by dividing by the dG level measured for the same sample. The detectable limit was calculated by dividing the minimum detected value by the dG level for each sample.
DNA methylation in vitro
Unmethylated plasmid was prepared from DAM/DCM– E. coli cells. For mSSSI methylation we used a pUC19 plasmid and, after purification of the plasmid by MaxiPrep (Qiagen), we treated it with mSSSI (NEB) for 1 h at 37 °C. To induce alkylation damage, we exposed unmethylated pUC19 plasmid to 20 mM MMS (Sigma) for 1 h at 25 °C before purification. DNMT3a and mutants thereof were expressed and purified from E. coli cells as described previously35. The reaction mixture was incubated for 2 h at 37 °C, and DNA was purified using phenol-chloroform extraction and analyzed using LC-MS as described above.
Plectus genome sequencing and assembly
We assembled and annotated a genome for P. sambesii using Illumina high-throughput sequencing data and using the methods documented in the supplementary information section (Supplementary Note 1). The final genome had a span of 186 Mb and an N50 of 4,039 bp, comparing well with other nematode genomes used in this study. The genome has been deposited in NCBI (PRJNA390260).
Generation and validation of ALKB2 deletion mutants
We obtained plasmids containing GFP-tagged CRISPR–Cas9 and guide RNAs targeting the first protein-coding exon of ALKB2 from Sigma. We used Lipofectamine transfection to introduce this plasmid into mouse ESCs and, after recovery of cells for 18 h at 37 °C, we used FACS to sort GFP-positive cells into individual wells of a 96-well plate. We screened the resultant clones for ALKB2 using PCR across the targeted exon searching for apparent size shifts. We then used Sanger sequencing of the PCR products to select clones showing indels in both alleles. We confirmed ALKB2 protein reduction using western blot analysis with anti-ALKBH2, a mouse monoclonal antibody (C-9; Santa Cruz, sc515789; dilution 1:1,000), using a rabbit anti-mouse HRP-conjugated secondary antibody (Abcam, ab6728; dilution 1:10,000). To test sensitivity to MMS, cells were treated with 200 mM MMS for 1 h before the MMS was washed out. We then sorted single cells using FACS and counted colonies formed after 5 d, comparing to a control treated with 0 mM MMS for each line.
URLs
Gene Expression Omnibus (GEO), http://www.ncbi.nlm.nih.gov/geo/; Wormbase and Wormbase ParaSite, http://www.sanger.ac.uk/science/tools/wormbase; Ensembl, http://ensemblgenomes.org/; NCBI, https://www.ncbi.nlm.nih.gov/; DNMT annotation (hmmer version 3.1), http://hmmer.org/; BLAST+ (version 2.2.30), https://blast.ncbi.nlm.nih.gov/; phylogenetic tree construction tools, http://www.phylogeny.fr/; Bismark (version 0.14.2), https://www.bioinformatics.babraham.ac.uk/projects/bismark/; Bowtie2 (version 2.1.0), http://bowtie-bio.sourceforge.net/bowtie2/; Methylextract (version 1.9), https://github.com/bioinfoUGR/methylextract?files=1; Bedtools (version 2.19.0), http://bedtools.readthedocs.io/en/latest/; R (version 3.1.0), https://www.r-project.org/.
Life Sciences Reporting Summary
Further information on experimental design is available in the Life Sciences Reporting Summary.
Code availability
Phylogenetic tree construction: MUSCLE v3.8.31 for alignment, Gblocks 0.91b for curation and PhyML 3.1 for maximum-likelihood phylogeny. Data integration was performed using Bedtools. Coevolution analysis was performed using BLAST+ version 2.2.30. All statistical analysis was carried out using R.
Custom Perl scripts (Perl version 5.16) used for intermediate processing of DNA methylation data are available from the authors upon request.
Data availability
Bisulfite sequencing data have been deposited into GEO with accession GSE104339. The P. sambesii genome assembly has been deposited to NCBI (PRJNA390260). Other nematode genomes are available from Wormbase and Wormbase. Animal, fungal and protist genomes are available from Ensembl. The genome of P. hawaiensis is available from NCBI.
References
Law, J. A. & Jacobsen, S. E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 11, 204–220 (2010).
Holliday, R. Epigenetics: a historical overview. Epigenetics 1, 76–80 (2006).
Ponger, L. & Li, W. H. Evolutionary diversification of DNA methyltransferases in eukaryotic genomes. Mol. Biol. Evol. 22, 1119–1128 (2005).
Jurkowski, T. P. & Jeltsch, A. On the evolutionary origin of eukaryotic DNA methyltransferases and Dnmt2. PLoS ONE 6, e28104 (2011).
Zemach, A., McDaniel, I. E., Silva, P. & Zilberman, D. Genome-wide evolutionary analysis of eukaryotic DNA methylation. Science 328, 916–919 (2010).
Feng, S. et al. Conservation and divergence of methylation patterning in plants and animals. Proc. Natl. Acad. Sci. USA 107, 8689–8694 (2010).
Lyko, F. et al. The honey bee epigenomes: differential methylation of brain DNA in queens and workers. PLoS Biol. 8, e1000506 (2010).
Xiang, H. et al. Single base–resolution methylome of the silkworm reveals a sparse epigenomic map. Nat. Biotechnol. 28, 516–520 (2010).
Bewick, A. J., Vogel, K. J., Moore, A. J. & Schmitz, R. J. Evolution of DNA methylation across insects. Mol. Biol. Evol. 34, 654–665 (2017).
Raddatz, G. et al. Dnmt2-dependent methylomes lack defined DNA methylation patterns. Proc. Natl. Acad. Sci. USA 110, 8627–8631 (2013).
Goll, M. G. et al. Methylation of tRNAAsp by the DNA methyltransferase homolog Dnmt2. Science 311, 395–398 (2006).
Gao, F. et al. Differential DNA methylation in discrete developmental stages of the parasitic nematode Trichinella spiralis. Genome Biol. 13, R100 (2012).
Schiffer, P. H. et al. The genome of Romanomermis culicivorax: revealing fundamental changes in the core developmental genetic toolkit in Nematoda. BMC Genomics 14, 923 (2013).
Li, Z. et al. Distinct roles of DNMT1-dependent and DNMT1-independent methylation patterns in the genome of mouse embryonic stem cells. Genome Biol. 16, 115 (2015).
Huff, J. T. & Zilberman, D. Dnmt1-independent CG methylation contributes to nucleosome positioning in diverse eukaryotes. Cell 156, 1286–1297 (2014).
Falckenhayn, C. et al. Characterization of genome methylation patterns in the desert locust Schistocerca gregaria. J. Exp. Biol. 216, 1423–1429 (2013).
Kao, D. et al. The genome of the crustacean Parhyale hawaiensis, a model for animal development, regeneration, immunity and lignocellulose digestion. eLife 5, e20062 (2016).
Sarkies, P. et al. Ancient and novel small RNA pathways compensate for the loss of piRNAs in multiple independent nematode lineages. PLoS Biol. 13, e1002061 (2015).
Ougland, R., Rognes, T., Klungland, A. & Larsen, E. Non-homologous functions of the AlkB homologs. J. Mol. Cell Biol. 7, 494–504 (2015).
Sedgwick, B. Repairing DNA-methylation damage. Nat. Rev. Mol. Cell Biol. 5, 148–157 (2004).
Strepetkaitė, D. et al. Analysis of DNA methylation and hydroxymethylation in the genome of crustacean Daphnia pulex. Genes 7, 1 (2015).
Ringvoll, J. et al. Repair deficient mice reveal mABH2 as the primary oxidative demethylase for repairing 1meA and 3meC lesions in DNA. EMBO J. 25, 2189–2198 (2006).
Nay, S. L., Lee, D.-H., Bates, S. E. & O’Connor, T. R. Alkbh2 protects against lethality and mutation in primary mouse embryonic fibroblasts. DNA Repair 11, 502–510 (2012).
Gowher, H. et al. Mutational analysis of the catalytic domain of the murine Dnmt3a DNA-(cytosine C5)-methyltransferase. J. Mol. Biol. 357, 928–941 (2006).
Klimasauskas, S., Kumar, S., Roberts, R. J. & Cheng, X. HhaI methyltransferase flips its target base out of the DNA helix. Cell 76, 357–369 (1994).
Tsumura, A. et al. Maintenance of self-renewal ability of mouse embryonic stem cells in the absence of DNA methyltransferases Dnmt1, Dnmt3a and Dnmt3b. Genes Cells 11, 805–814 (2006).
Sved, J. & Bird, A. The expected equilibrium of the CpG dinucleotide in vertebrate genomes under a mutation model. Proc. Natl. Acad. Sci. USA 87, 4692–4696 (1990).
Alexandrov, L. B. et al. Clock-like mutational processes in human somatic cells. Nat. Genet. 47, 1402–1407 (2015).
Drabløs, F. et al. Alkylation damage in DNA and RNA—repair mechanisms and medical significance. DNA Repair 3, 1389–1407 (2004).
Furrer, A. & van Loon, B. Handling the 3-methylcytosine lesion by six human DNA polymerases members of the B-, X- and Y-families. Nucleic Acids Res. 42, 553–566 (2014).
Chastain, P. D. II et al. Abasic sites preferentially form at regions undergoing DNA replication. FASEB J. 24, 3674–3680 (2010).
Shrivastav, M., De Haro, L. P. & Nickoloff, J. A. Regulation of DNA double-strand break repair pathway choice. Cell Res. 18, 134–147 (2008).
Blaxter, M. L. et al. A molecular evolutionary framework for the phylum Nematoda. Nature 392, 71–75 (1998).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Emperle, M., Rajavelu, A., Reinhardt, R., Jurkowska, R. Z. & Jeltsch, A. Cooperative DNA binding and protein/DNA fiber formation increases the activity of the Dnmt3a DNA methyltransferase. J. Biol. Chem. 289, 29602–29613 (2014).
Acknowledgements
We thank H. Leitch and M. Borkowska for invaluable help with mouse ESC culture. We would like to thank M. Merkenschlager, L. Aragon, J. Sale and B. Lehner for helpful comments on the manuscript, M. Blaxter for advice on nematode genomics, and M. Berriman for access to the N. brasiliensis draft genome. P.S. is funded by an Imperial College Research Fellowship. Work in the Sarkies and Hajkova laboratories is funded by the Medical Research Council. P.H. is a recipient of the ERC CoG grant “dynamic modifications” and a member of the EMBO Young Investigator Programme. A.J. and M.E. are funded by DFG JE252/10. R.K.G. and A.J.B. are funded by Wellcome Trust grant 083620Z and Centre grant 203128/Z/16/Z. P.H.S. is funded by the ERC in a grant to Max Telford (ERC-2012-AdG 322790).
Author information
Authors and Affiliations
Contributions
P.S. and P.H. conceived the study. P.S., P.H. and A.J. designed the experiments. DNA extraction and bisulfite sequencing were carried out by S.R. and P.S. P.S. performed bioinformatic and computational analyses. 3meC analysis by LC/MS was carried out by R.A., C.E.R., S.L. and P.S. ESC CRISPR deletion and analysis was performed by A.G., J.K.R. and P.S. M.E. and A.J. carried out the in vitro DNMT3a analysis. T.B. and P.H.S. performed genome assembly. S.R., M.E.S., R.K.G. and A.J.B. were responsible for nematode culture. P.S., P.H. and A.J. analyzed the data and prepared the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Extended analysis of the sequence context of DNA methylation in nematodes
a, Overall percentage methylation at CG and non-CG sites respectively for different nematodes. The chi-squared test P value is for a two-tailed test on a 2 by 2 table: meCG/non-meCG|meCX/non-meCX, where X is C, A or T. P = 1 × 10–100 is the minimum P value reported by the software used for calculation. See Supplemental Note 3 for the number of CGs in different contexts across the genome for all genomes analyzed. b,c, Enrichment of methylation for CG sites within different contexts for P. sambesii and T. spiralis respectively (nematodes with the most pronounced differences in CG[X] preferences; see Fig. 2). See Supplemental Note 3 for the number of CGs in different contexts across the genome for all genomes analyzed.
Supplementary Figure 2
Sample genome browser windows for P. sambesii and R. culicivorax showing lack of methylation on genes but high methylation on repeats
Supplementary Figure 3
Sample genome browser windows for T. spiralis and T. muris demonstrating methylation prominence at repeats rather than genes
Supplementary Figure 4 Comparison of DNA methylation in genes and repeats
a–d, Methylation levels across genes for either genes with repeat homology or those without for the different nematode species with DNMT activity. P values shown are from two-tailed Wilcoxon unpaired tests. The box plot shows interquartile range, with a line at the median and whiskers extending to the most extreme point no more than 1.5 times the interquartile range from the box. The notch shows 95% confidence levels on the median (1.57 times the interquartile range/√number of samples). P values are reported to the nearest significant figure. Analysis is based on the number of genes reported in Supplemental Note 3.
Supplementary Figure 5 Cytosine methylation and CG sequence content of genes in nematodes
a–d, Comparison of methylation as a fraction of total cytosine (rather than CG) and CG levels upstream and downstream of the transcriptional start site for the different nematode species with cytosine DNMT activity. e, Variation of CG methylation normalized to CG content with overall CG content for exons (diamonds) or introns (triangles) in different nematodes. The number of CG sites analyzed for all plots is in Supplemental Note 3.
Supplementary Figure 6 Coevolution of DNA repair proteins with DNA methyltransferases
Heat map showing the conservation of different proteins with a GO term for DNA repair that coevolve with cytosine DNA methylation. DNMT1 and DNMT3 are shown for comparison. In addition to alkylation repair, this set of genes includes components of base excision repair, which is also involved in repairing DNA alkylation damage, and double-strand break repair, which would be required in the event of replication fork stalling, for instance at 3meC lesions if not repaired by ALKB2.
Supplementary Figure 7 ALKB2/3 coevolves with DNA methyltransferases across metazoa
Heat map showing the conservation of ALKB and DNMT family members across metazoans. ALKB2/3 are the only ALKB family members that show significant coevolution with DNA methyltransferases.
Supplementary Figure 8
ALKB2/3 and DNMT coevolution in metazoa
Supplementary Figure 9
DNA methylation and ALKB2/3 coevolution in protists
Supplementary Figure 10
DNA methylation and ALKB2/3 coevolution in fungi
Supplementary Figure 12 ALKB2/3 supports higher DNA methylation levels in arthropods
DNA methylation levels across arthropods comparing 4 species with ALKB2/3 and 17 species without ALKB2/3. A coding sequence B across the genome. The only species that has ALKB2/3 yet shows extremely low levels of detectable methylation is T. castaneum. The box plot shows a line at the median, the interquartile range as the box and whiskers extending to the greatest point that is less than 1.5 times the interquartile range. For the full list of arthropod species analyzed, see Supplemental Table 9.
Supplementary Figure 13 Establishment of a method to monitor 3meC by LC/MS
a, Example of a standard curve and LCQ for 3meC. The standard curve was repeated each time measurements were taken (5), with similar results. b,c, 5meC and 3meC levels measured in mSssI methylated and unmethylated plasmid with or without exposure to MMS. Technical variation from two measurements is shown. Note low-level induction of 3meC by mSssI alone in c. The signal-to-noise ratio for 3meC detection was 10 as per the Methods. d,e, Screenshots of LC/MS traces for two further independent replicate in vitro reactions containing the catalytic domain of DNMT3a and an unmethylated plasmid, showing induction of 5meC and 3meC.
Supplementary Figure 14 Analysis of alkylation damage caused by DNMTs
a, Dot blot demonstrating the specificity of 3meC antibody, which recognizes a DNA template alkylated in vitro and not a 5meC-containing template synthesized by PCR, in contrast to the anti-5meC antibody, which only recognizes the 5meC template. The experiment was repeated twice with similar results. b, Dot blot showing 3meC levels in WT ES cells and cells carrying either DNMT3a and DNMT3b deletion (DKO) or DNMT3a, DNMT3b and DNMT1 deletion (TKO). The experiment was repeated four times with similar results. c, Western blot showing comparable protein levels for both WT and mutant (F646A) DNMT3a catalytic domain expressed and purified from E. coli. The experiment was performed twice with similar results.
Supplementary Figure 15 Generation of clones carrying deletions in ALKB2 in mouse ES cells
a, CRISPR strategy to generate disruptions in the first coding exon of mouse ALKBH2, the ortholog of ALKB2. b, Example of sequencing data showing out-of-frame deletion in both alleles, indicating successful disruption of the protein. c, Western blot confirming reduction of ALKB2 protein levels. Non-specific bands are indicated by an asterisk. Tubulin detection from the same membrane is shown as a control. The blot was performed three times for WT clones and two times for TKO clones. d, Sensitivity to 200 mM MMS. The y axis shows percentage surviving colonies relative to an untreated sample from the same line. The mean and range of two biological replicates are shown.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15, Supplementary Tables 1–9 and Supplementary Notes 1–3
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0), which permits use, duplication, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Rošić, S., Amouroux, R., Requena, C.E. et al. Evolutionary analysis indicates that DNA alkylation damage is a byproduct of cytosine DNA methyltransferase activity. Nat Genet 50, 452–459 (2018). https://doi.org/10.1038/s41588-018-0061-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0061-8
This article is cited by
-
Annelid methylomes reveal ancestral developmental and aging-associated epigenetic erosion across Bilateria
Genome Biology (2024)
-
Selfish conflict underlies RNA-mediated parent-of-origin effects
Nature (2024)
-
DNA methylation on C5-Cytosine and N6-Adenine in the Bursaphelenchus xylophilus genome
BMC Genomics (2023)
-
Context-dependent DNA polymerization effects can masquerade as DNA modification signals
BMC Genomics (2022)
-
Small RNA pathways in the nematode Ascaris in the absence of piRNAs
Nature Communications (2022)